Chapter 3 The immune system and disease
Anatomy and principles of the immune system
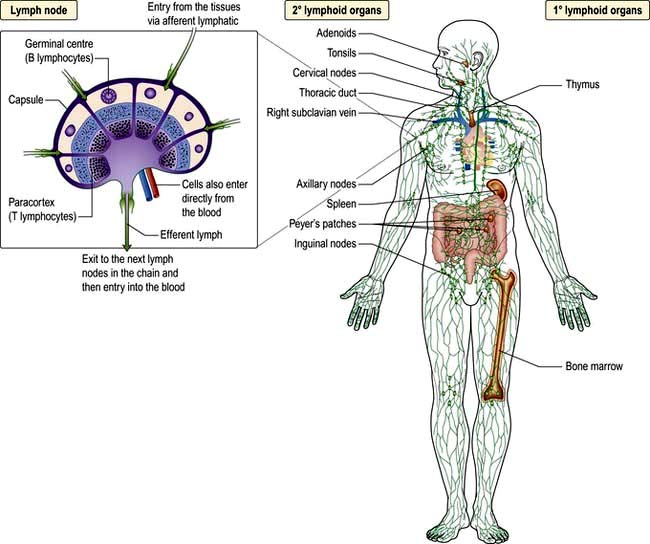
Immunity can be defined as protection from infection, whether it be due to bacteria, viruses, fungi or multicellular parasites. Like other organs involved in human physiology, the immune system is composed of cells and molecules organized into specialized tissues (Fig. 3.1).
Cells involved in immune responses: origin and function
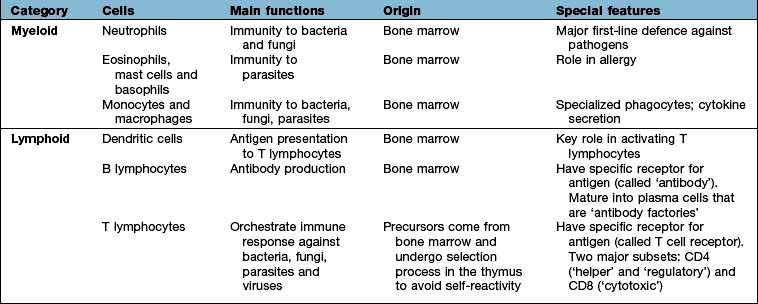
All immune cells have a common source in the pluripotent stem cells generated in the bone marrow (Fig. 3.1). They have diverse functions (Table 3.1). T lymphocytes undergo ‘education’ in the thymus to avoid self-recognition, and populate the peripheral lymphoid tissue, where B lymphocytes also reside. Both sets of lymphocytes undergo activation in the peripheral tissue, to become mature effector cells. B lymphocytes may further differentiate into antibody-secreting plasma cells. Lymphoid tissue is frequently found at mucosal surfaces in non-encapsulated patches, termed mucosa-associated lymphoid tissue (MALT).
The immune system
Cells and molecules involved in immune responses are classified into innate and adaptive systems:
 The innate immune system is inborn and operates throughout life (pp. 51–55)
The innate immune system is inborn and operates throughout life (pp. 51–55)
 The adaptive immune system changes in response to the pathogens it encounters (pp. 58–62).
The adaptive immune system changes in response to the pathogens it encounters (pp. 58–62).
There are also non-immunological barriers that are involved in host protection, and very often it is the lowering of these that allows a pathogen to take a foothold (Table 3.2).
Table 3.2 Non-immunological host defence mechanisms
Normal barriers | Events that may compromise barrier function |
Physical barriers |
|
Skin and mucous membranes | Trauma, burns, i.v. cannulae |
Cough reflex | Suppression, e.g. by opiates, neurological disease |
Mucosal function | Ciliary paralysis (e.g. smoking) |
Increased mucus production (e.g. asthma) | |
Abnormally viscid secretions (e.g. cystic fibrosis) | |
Decreased secretions (e.g. sicca syndrome) | |
Urine flow | Stasis (e.g. prostatic hypertrophy) |
Chemical barriers | Low gastric pH (gastric acid secretion inhibitors) |
Resistance to pathogens provided by commensal skin and gut organisms | Changes in flora (e.g. broad-spectrum antibiotics) |
A major feature of the immune system is the complexity of the surface-bound, intracellular and soluble structures that mediate its functions. In particular, it is necessary to be aware of the CD (clusters of differentiation) classification (Box 3.1) and the functions of cytokines and chemokines.
![]() Box 3.1
Box 3.1
The CD classification
This is the ultimate way of defining a cell
 Immune cells are distinguished by the surface receptors and proteins that they express in order to mediate their particular range of immunological functions, e.g. cell–cell signalling, cell activation
Immune cells are distinguished by the surface receptors and proteins that they express in order to mediate their particular range of immunological functions, e.g. cell–cell signalling, cell activation
 The surfaces are covered with such proteins and indicate the cell lineage or differentiation pathway. The discovery of monoclonal antibodies (proteins tailor-made to bind to a specific target) made this feasible
The surfaces are covered with such proteins and indicate the cell lineage or differentiation pathway. The discovery of monoclonal antibodies (proteins tailor-made to bind to a specific target) made this feasible
 Surface molecules defining the origin and function of selected groups of cells are known as clusters of differentiation (CD). Over 300 CD numbers exist
Surface molecules defining the origin and function of selected groups of cells are known as clusters of differentiation (CD). Over 300 CD numbers exist
 A clinical example is the number of peripheral blood lymphocytes expressing CD4 (’the CD4 count’), which is used to monitor HIV infection (see p. 178)
A clinical example is the number of peripheral blood lymphocytes expressing CD4 (’the CD4 count’), which is used to monitor HIV infection (see p. 178)
An updated listing is available at: http://www.hcdm.org/
Cytokines
These are small polypeptides released by a cell in order to change the function of the same or another cell. These chemical messengers are found in many organ systems, but especially the immune system. Cytokines have become markers in the investigation of disease pathogenesis; therapeutic agents in their own right; and the targets of therapeutic agents (see p. 72). The key features of a cytokine are:
 pleiotropy: different effects on different cells
pleiotropy: different effects on different cells
 autocrine function: modulates the cell secreting it
autocrine function: modulates the cell secreting it
 paracrine function: modulates adjacent cells
paracrine function: modulates adjacent cells
 endocrine effects: modulates cells and organs at remote sites
endocrine effects: modulates cells and organs at remote sites
 synergistic activity: acting in concert with other cytokines to achieve effects greater than the summation of their individual actions.
synergistic activity: acting in concert with other cytokines to achieve effects greater than the summation of their individual actions.
Cells and molecules of the innate immune system
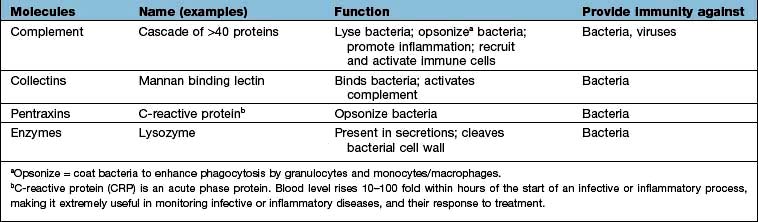
Innate immunity provides immediate, first-line, host defence. The key features of this system are shown in Table 3.3. It is present at birth and remains operative at comparable intensity into old age. Innate immunity is mediated by a variety of cells and molecules (Table 3.4). Activation of innate immune responses is mediated through interaction between the:
 pathogen side comprising a relatively limited array of molecules (pathogen-associated molecular patterns, PAMPs)
pathogen side comprising a relatively limited array of molecules (pathogen-associated molecular patterns, PAMPs)
 host side – a limited portfolio of receptors (pattern recognition receptors, PRRs).
host side – a limited portfolio of receptors (pattern recognition receptors, PRRs).
Table 3.3 Features of the innate and adaptive immune responses
| Innate | Adaptive |
|---|---|
No memory: quality and intensity of response invariant | Memory: response adapts with each exposure |
Recognizes limited number of non-varying, generic molecular patterns on, or made by, pathogens | Recognizes vast array of specific antigensa on, or made by, pathogens |
Pattern recognition mediated by a limited array of receptors | Antigen recognition mediated by a vast array of antigen-specific receptors |
Response immediate on first encounter | Response on first encounter takes 1–2 weeks; on second encounter 3–7 days |
a Antigen is a molecular structure (protein, peptide, lipid, carbohydrate) that generates an immune response.
Activation of certain cells in the innate immune system leads to activation of the adaptive immune response (see p. 58).
Complement
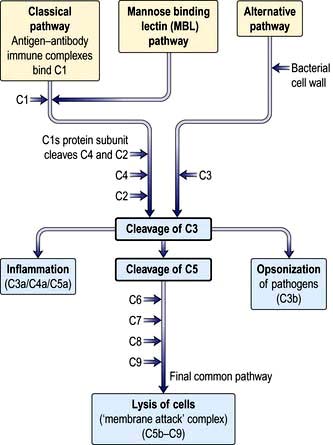
Complement proteins are produced in the liver. Each complement circulates in an inactive form until triggered to become enzymatically active, when it then activates several molecules of the next stage in a series. This complement cascade is initiated via three distinct pathways: alternative, classical and mannan-binding lectin (Fig. 3.2). These pathways are composed of three distinct enzyme cascades that culminate in the cleavage of C3 and C5. Cleavage of C3 has a number of biological consequences; breakdown of C5 achieves the same and, in addition, provides the triggering stimulus to the final common (‘membrane attack’) pathway, which provides most of the biological activity (Fig. 3.2).
The main functions of complement activation are to:
 promote inflammation (e.g. through the actions of the anaphylatoxins C3a, C4a and C5a)
promote inflammation (e.g. through the actions of the anaphylatoxins C3a, C4a and C5a)
 recruit cells (e.g. through chemoattractants)
recruit cells (e.g. through chemoattractants)
 kill targeted cells, such as bacteria
kill targeted cells, such as bacteria
 solubilize and remove from the circulation antigen-antibody (‘immune’) complexes.
solubilize and remove from the circulation antigen-antibody (‘immune’) complexes.
Neutrophils
Neutrophils (see p. 413) phagocytose and kill microorganisms. They are derived from the bone marrow, which can produce between 1011 (healthy state) and 1012 (during infection) new cells per day. In health, neutrophils are rarely seen in the tissues.
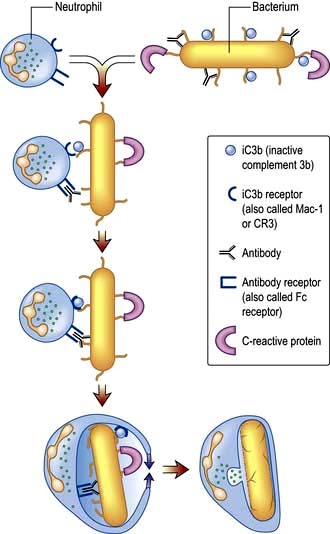
Neutrophil phagocytosis is activated by interaction with bacteria, either directly or after bacteria have been coated (opsonized) to make them more ingestible (Fig. 3.3). The contents of neutrophil granules are released both intracellularly (predominantly azurophilic granules) and extracellularly (specific granules) following fusion with the plasma membrane. Approximately 100 different molecules in neutrophil granules (Table 3.5) kill and digest microorganisms, for example:
 Myeloperoxidase and cytochrome b558 are key components of major oxygen-dependent bactericidal systems.
Myeloperoxidase and cytochrome b558 are key components of major oxygen-dependent bactericidal systems.
 Cathepsins, proteinase-3 and elastase are deadly to Gram-positive and Gram-negative organisms, as well as some Candida species.
Cathepsins, proteinase-3 and elastase are deadly to Gram-positive and Gram-negative organisms, as well as some Candida species.
 Defensins are naturally occurring cysteine-rich antibacterial and antifungal polypeptides (29–35 amino acids).
Defensins are naturally occurring cysteine-rich antibacterial and antifungal polypeptides (29–35 amino acids).
 Collagenase and elastase break down fibrous structures in the extracellular matrix, facilitating progress of the neutrophil through the tissues.
Collagenase and elastase break down fibrous structures in the extracellular matrix, facilitating progress of the neutrophil through the tissues.
Table 3.5 Contents and function of key neutrophil granules
| Function | Primary or azurophilic granules | Secondary or specific granules |
|---|---|---|
Antibacterial | Lysozyme | Respiratory burst components (e.g. cytochrome b558) producing reactive oxygen metabolites, such as hydrogen peroxide, hydroxyl radicals and singlet oxygen |
Cathepsins | Lysozyme | |
Bactericidal/permeability increasing protein (BPI) | Lactoferrin | |
Cell movement |
| Collagenase |
| CD11b/CD18 (adhesion molecule) | |
| N-formyl-methionyl-leucylphenylalanine receptor (FMLP-R) |
Eosinophils
In contrast to neutrophils, several hundred times more eosinophils are present in the tissues than in the blood, particularly at epithelial surfaces where they survive for several weeks. The main role of eosinophils is protection against multicellular parasites such as worms (helminths). This is achieved by the release of pro-inflammatory mediators, which are toxic, cationic proteins. In populations and societies in which such parasites are rare, eosinophils contribute mainly to allergic disease, particularly asthma (see p. 827). Eosinophils have two types of granules:
 Specific granules (95%) contain the cationic proteins, of which there are four main types: major basic protein (MBP), eosinophil cationic protein (ECP), eosinophil neurotoxin, which are all potently and exquisitely toxic to helminths (ECP also has some bactericidal properties), and eosinophil peroxidase, which has similar activity to neutrophil myeloperoxidase.
Specific granules (95%) contain the cationic proteins, of which there are four main types: major basic protein (MBP), eosinophil cationic protein (ECP), eosinophil neurotoxin, which are all potently and exquisitely toxic to helminths (ECP also has some bactericidal properties), and eosinophil peroxidase, which has similar activity to neutrophil myeloperoxidase.
 Primary granules (5%) synthesize and release leukotrienes C4 and D4 and platelet-activating factor (PAF) which alter airway smooth muscle and vasculature (see p. 828).
Primary granules (5%) synthesize and release leukotrienes C4 and D4 and platelet-activating factor (PAF) which alter airway smooth muscle and vasculature (see p. 828).
Mast cells and basophils
Mast cells and basophils share features in common, especially in containing:
 high-affinity receptors for immunoglobulin E (IgE; an antibody type that is involved in allergic disease, see p. 68, 69).
high-affinity receptors for immunoglobulin E (IgE; an antibody type that is involved in allergic disease, see p. 68, 69).
Mast cells are found in tissues (especially skin and mucosae) and basophils in the blood. Both mast cells and basophils release pro-inflammatory mediators which are either pre-formed or synthesized de novo (Table 3.6).
Table 3.6 Mast cell and basophil mediators
| Mediators | Effects |
|---|---|
Pre-formed: |
|
Histamine | Vasodilatation |
Vascular permeability ↑ | |
Smooth muscle contraction in airways | |
Proteases | Digestion of basement membrane causes ↑vascular permeability and aids migration |
Proteoglycans (e.g. heparan) | Anticoagulant activity |
Synthesized de novo: |
|
Platelet-activating factor (PAF) | Vasodilator |
LTB4, LTC4, LTD4 | Neutrophil and eosinophil activators and chemoattractants |
Vascular permeability ↑ | |
Bronchoconstrictors | |
Prostaglandins (mainly PGD2) | Vascular permeability ↑ |
| Bronchoconstrictors |
| Vasodilators |
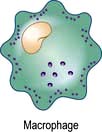
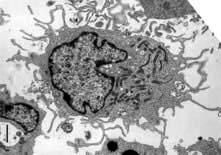
Monocytes and macrophages
Tissue macrophages
Macrophages have pro-inflammatory and microbicidal capabilities similar to those of neutrophils. Under activation conditions, antigen presentation (see p. 57, 58) is enhanced and a range of cytokines secreted, notably TNF-α, IL-1 and IFN-γ. These are necessary for the removal of certain pathogens that live within mononuclear phagocytes (e.g. mycobacteria). Macrophages and related cells may also undergo a process termed autophagy (p. 32, Ch. 2). This self-cannibalization is a critical property of many cell types under starvation conditions, but is used by the immune system to destroy intracellular pathogens such as Mycobacterium tuberculosis, which otherwise persist within cells and block normal antibacterial processes. Autophagy is also a means of enhancing antigen presentation pathways.
Tissue macrophages involved in chronic inflammatory foci may undergo terminal differentiation into multinucleated giant cells, typically found at the site of the granulomata characteristic of tuberculosis and sarcoidosis (see p. 845).
Dendritic cells
 dendritic morphology (Fig. 3.4)
dendritic morphology (Fig. 3.4)
 machinery for sensing pathogens
machinery for sensing pathogens
 the ability to process and present antigens to CD4 and CD8 T lymphocytes, coupled with the ability to activate these T lymphocytes from a naive state
the ability to process and present antigens to CD4 and CD8 T lymphocytes, coupled with the ability to activate these T lymphocytes from a naive state
 the ability to dictate the T lymphocyte’s future function and differentiation.
the ability to dictate the T lymphocyte’s future function and differentiation.
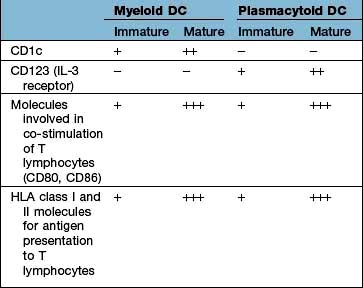
Types of dendritic cell
The major types are the myeloid DC (mDC), the plasmacytoid DC (pDC) and a variety of specialized DCs found in tissues that resemble mDCs (e.g. the Langerhans cell in the skin, see Fig. 24.1). DCs have several distinctive cell surface molecules, some of which have pathogen-sensing activity (e.g. the antigen uptake receptor DEC205 on mDCs) whilst others are involved in interaction with T lymphocytes (Table 3.7). Immature mDCs and pDCs are present in the blood, but at very low levels (<0.5% of lymphocyte/monocyte cells).
 Mannan-binding lectin, which initiates complement activity inducing opsonization (p. 51, 52).
Mannan-binding lectin, which initiates complement activity inducing opsonization (p. 51, 52).
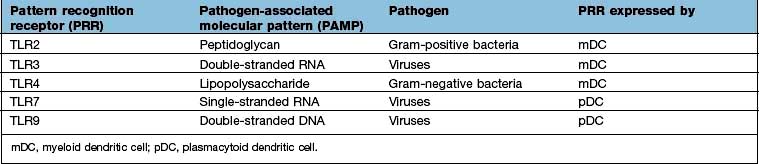
 Signalling receptors such as the PRR known as TLR4 (for toll-like receptor 4), which binds lipopolysaccharide, a molecular pattern found in the cell walls of many Gram-negative bacteria (Table 3.8), whilst others bind double-stranded and single-stranded RNA from viruses. Innate immunity critically depends on toll-like receptor signalling. These receptors act through a critical adaptor molecule, myeloid differentiation factor 88 (MyD88), to regulate the activity of nuclear factor kappa B (NF-κB) pathways.
Signalling receptors such as the PRR known as TLR4 (for toll-like receptor 4), which binds lipopolysaccharide, a molecular pattern found in the cell walls of many Gram-negative bacteria (Table 3.8), whilst others bind double-stranded and single-stranded RNA from viruses. Innate immunity critically depends on toll-like receptor signalling. These receptors act through a critical adaptor molecule, myeloid differentiation factor 88 (MyD88), to regulate the activity of nuclear factor kappa B (NF-κB) pathways.
 Endocytic pattern recognition receptors, which act by enhancing antigen presentation on macrophages, by recognizing microorganisms with mannose-rich carbohydrates on their surface or by binding to bacterial cell walls and scavenging bacteria from the circulation. All lead to phagocytosis.
Endocytic pattern recognition receptors, which act by enhancing antigen presentation on macrophages, by recognizing microorganisms with mannose-rich carbohydrates on their surface or by binding to bacterial cell walls and scavenging bacteria from the circulation. All lead to phagocytosis.
 TREM-1 (triggering receptor expressed on myeloid cells) is a cell surface receptor which, when bound to its ligand, triggers secretion of pro-inflammatory cytokines. It is upregulated by bacterial lipopolysaccharides but not in non-infective disorders.
TREM-1 (triggering receptor expressed on myeloid cells) is a cell surface receptor which, when bound to its ligand, triggers secretion of pro-inflammatory cytokines. It is upregulated by bacterial lipopolysaccharides but not in non-infective disorders.
DCs and T cell activation
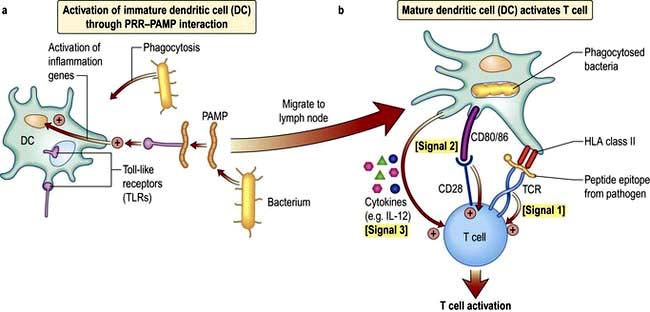
In a sequence of events that spans 1–2 days, immature DCs are activated by PAMPs or DAMPs in the tissues binding to a PRR on DCs. The immature pDC is a small rounded cell that develops dendrites upon activation and secretes enormous quantities of IFN-α, a potent antiviral and pro-inflammatory cytokine. On activation, the DC migrates to the local lymph node with the engulfed pathogen. During migration the DC matures, changing its shape, gene and molecular profile and function within a matter of hours to take on a mature form, with altered functions (Table 3.9, Fig. 3.5), in particular upregulating machinery required to activate T lymphocytes. Once in the lymph node, the mature DC interacts with naive T lymphocytes (antigen presentation), resulting in two key outcomes:
1. Activation of T cells with the ability to recognize peptide fragments (termed epitopes) of the pathogen
2. Polarization of the T cell towards a functional phenotype (see below) that is tailored to the particular pathogen.
Table 3.9 Myeloid dendritic cell (DC) maturation
| Immature mDC | Mature mDC |
|---|---|
Highly pinocytotic | Ceases pinocytosis |
Low level expression of molecules required for T lymphocyte activation | Upregulates CD80, CD86 and HLA molecules |
Low level expression of machinery required to process and present microbial antigens | Begins to process microbial antigens (break down into small peptides) in readiness to present them to T lymphocytes (using HLA molecules) |
Generally localized and sedentary | Begins active migration to local lymph node |
Minimal secretion of cytokines | Active secretion of cytokines in readiness to stimulate T lymphocytes; in particular IL-12 |
mDC, myeloid dendritic cell.
The mature DC provides three major signals to naive T cells Fig. 3.5):
Hla molecules and antigen presentation
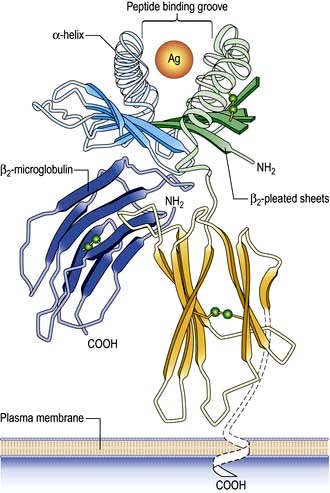
On the short arm of chromosome 6 is a collection of genes termed the major histocompatibility complex (MHC; known as the human leucocyte antigens, or HLA in man), which plays a critical role in immune function. MHC genes code for proteins expressed on the surface of a variety of cell types that are involved in antigen recognition by T lymphocytes. The T lymphocyte receptor for antigen recognizes its ligand as a short antigenic peptide embedded within a physical groove at the extremity of the HLA molecule (Fig. 3.6).
The HLA genes are particularly interesting for clinicians and biologists. First, differences in HLA molecules between individuals are responsible for tissue and organ graft rejection (hence the name ‘histo’(tissue)-compatibility). Second, possession of certain HLA genes is linked to susceptibility to particular diseases (Table 3.10).
Table 3.10 HLA associations with immune-mediated and infectious diseases
| Disease process | Disease | HLA type |
|---|---|---|
Autoimmunity | Type 1 diabetes | Class II: |
| DQA1*0301/DQB1*0302 (susceptibility) | |
DQA1*501/DQB1*201 (susceptibility) | ||
DQA1*0102/DQB1*0602 (protection) | ||
| Class I: | |
| HLA-A24; HLA-B*18; HLA-B*39 | |
Multiple sclerosis | DRB1*1501 (susceptibility) | |
Rheumatoid arthritis | DRB1*0404 (susceptibility) | |
Autoimmune hepatitis | DRB1*03 and DRB1*04 (susceptibility) | |
Goodpasture’s syndrome (anti-glomerular basement membrane disease) | HLA-DRB1*1501 (susceptibility) | |
Pemphigus vulgaris | DRB1*0402; DQB1*0503 (susceptibility) | |
Inflammatory | Coeliac disease | DQA1*0501/DQB1*0201 (susceptibility) |
Ankylosing spondylitis | HLA-B27 (susceptibility) | |
Psoriasis | HLA-Cw*0602 (susceptibility) | |
Infectious | Human immunodeficiency virus infection | HA-B27; HLA-B*51; HLA-B*57 (associated with slow progression of disease) |
HLA-B*35 (associated with rapid progression) |
The human major histocompatibility complex
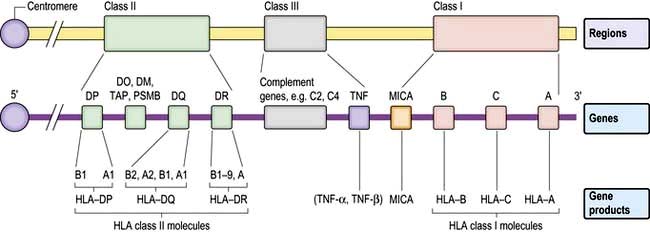
The human MHC comprises three major classes (I, II and III) of genes involved in the immune response (Fig. 3.7).
HLA classes
Classical class I HLA genes (also termed Ia), are designated HLA-A, HLA-B and HLA-C. Each encodes a class I α chain, which combines with a β chain to form the class I HLA molecule (Fig. 3.6). While there are several types of α chains, there is only one type of β chain, β2 microglobulin. The HLA class I molecule has the role of presenting short (8–10 amino acids) antigenic peptides to the T cell receptor on the subset of T lymphocytes that bear the co-receptor CD8. As an example of HLA polymorphism, there are nearly 200 allelic forms at the A gene locus. Class I HLA molecules are expressed on all nucleated cells.
Non-classical HLA class I genes are less polymorphic, have a more restricted expression on specialized cell types, and present a restricted type of peptide or none at all. These are the HLA-E, F and G (Ib genes) and MHC class I-related (MIC, or class Ic) genes, A and B. The products of these genes are predominantly found on epithelial cells, signal cellular stress and interact with lymphoid cells, especially natural killer cells (see p. 61, 62).
HLA genotypes and the range of their protein products
Some general principles apply to the HLA genes and their protein products:
 The presence of multiple genes on each chromosome, and the fact that both maternal and paternal genes are co-dominantly expressed, allows considerable breadth in the number of HLA molecules that an individual expresses.
The presence of multiple genes on each chromosome, and the fact that both maternal and paternal genes are co-dominantly expressed, allows considerable breadth in the number of HLA molecules that an individual expresses.
 The existence of polymorphisms at each locus provides great breadth in the number of HLA molecules expressed at a population level. The polymorphic forms of HLA molecules differ predominantly in the peptide-binding groove (Fig. 3.6).
The existence of polymorphisms at each locus provides great breadth in the number of HLA molecules expressed at a population level. The polymorphic forms of HLA molecules differ predominantly in the peptide-binding groove (Fig. 3.6).
Antigen presentation
 The endogenous route (Fig. 3.8) is a property of all nucleated cells: the internal milieu is sampled to generate peptide–HLA class I complexes for display (‘presentation’) on the cell surface. In a healthy cell, the peptides are derived from self-proteins in the cytoplasm (Fig. 3.8) and are ignored by the immune system. In a virus-infected cell, viral proteins are processed and presented. The resulting viral peptide–HLA class I complex is presented to CD8 T lymphocytes that have cytotoxic (killer) function. In an immune response against a virus infection, CD8 T lymphocytes recognizing viral peptide–HLA complexes on the surface of an infected cell will kill it as a means to limit and eradicate infection.
The endogenous route (Fig. 3.8) is a property of all nucleated cells: the internal milieu is sampled to generate peptide–HLA class I complexes for display (‘presentation’) on the cell surface. In a healthy cell, the peptides are derived from self-proteins in the cytoplasm (Fig. 3.8) and are ignored by the immune system. In a virus-infected cell, viral proteins are processed and presented. The resulting viral peptide–HLA class I complex is presented to CD8 T lymphocytes that have cytotoxic (killer) function. In an immune response against a virus infection, CD8 T lymphocytes recognizing viral peptide–HLA complexes on the surface of an infected cell will kill it as a means to limit and eradicate infection.
 The exogenous route (Fig. 3.9) is a property of APCs: the external milieu is sampled. Antigens are internalized, either in the process of phagocytosis of a pathogen, through pinocytosis, or through specialized surface receptors (e.g. for antigen/antibody/complement complexes). The antigen is broken down by a combination of low pH and proteolytic enzymes for ‘loading’ into HLA class II molecules. At the APC surface the pathogen peptide–HLA class II complex is presented to and able to interact with CD4 T lymphocytes. Presentation by DCs can initiate an adaptive immune response by activating a naive, pathogen-specific CD4 T lymphocyte. Presentation by monocyte/macrophages and B lymphocytes can maintain and enhance this response by activating effector and memory pathogen-specific CD4 T lymphocytes.
The exogenous route (Fig. 3.9) is a property of APCs: the external milieu is sampled. Antigens are internalized, either in the process of phagocytosis of a pathogen, through pinocytosis, or through specialized surface receptors (e.g. for antigen/antibody/complement complexes). The antigen is broken down by a combination of low pH and proteolytic enzymes for ‘loading’ into HLA class II molecules. At the APC surface the pathogen peptide–HLA class II complex is presented to and able to interact with CD4 T lymphocytes. Presentation by DCs can initiate an adaptive immune response by activating a naive, pathogen-specific CD4 T lymphocyte. Presentation by monocyte/macrophages and B lymphocytes can maintain and enhance this response by activating effector and memory pathogen-specific CD4 T lymphocytes.
 Cross-presentation refers to the ability of some APCs (mainly DCs) to internalize exogenous antigens and process them through the endogenous route (Fig. 3.8). This is an essential component in the activation of CD8 cytotoxic T cell responses against a virus.
Cross-presentation refers to the ability of some APCs (mainly DCs) to internalize exogenous antigens and process them through the endogenous route (Fig. 3.8). This is an essential component in the activation of CD8 cytotoxic T cell responses against a virus.
Stay updated, free articles. Join our Telegram channel

Full access? Get Clinical Tree