Chapter 2 Molecular cell biology and human genetics
Cell biology
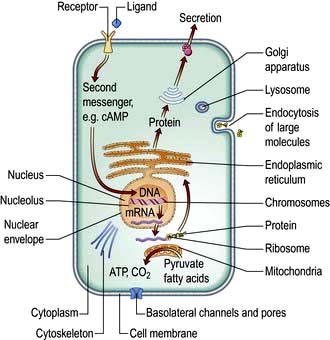
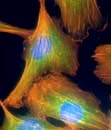
Cells consist of cytoplasm enclosed within a lipid sheath (the plasma membrane). The cytoplasm contains a variety of organelles (sub-cellular compartments enclosed within their own membranes) in a mixture of salts and organic compounds (the cytosol). These are held within an adaptive internal scaffold (the cytoskeleton) that radiates from the nucleus outwards to the cell surface (Fig. 2.1). Many cells have special functions and their size, shape and behaviour adapt to meet their physiological roles. Cells can be organized into tissues and organs in which the individual component cells are in contact and able to send and receive messages, both directly and indirectly. Coordinated cellular responses can be achieved through systemic signalling, e.g. via hormones.
Cell structure
Cellular membranes
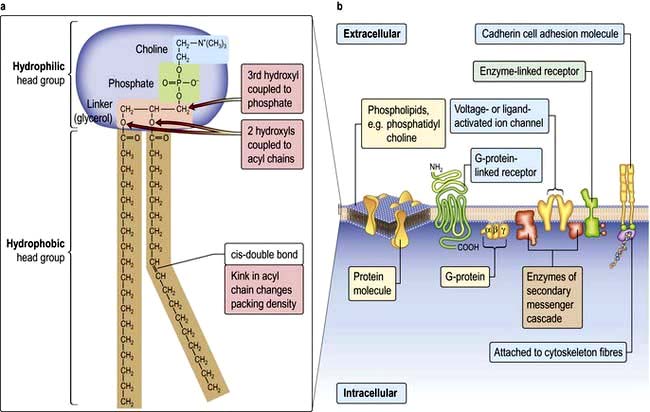
 Lipid bilayers separate the cell contents from the external environment and compartmentalize distinct cellular activities into organelles. These consist of a large variety of glycerophospholipids and sphingolipids. Membrane lipids usually have two hydrophobic acyl chains linked via glycerol or serine, to polar hydrophilic head groups (Fig. 2.2). This amphiphilic nature, with a ‘water-loving’ head and a ‘water-hating’ tail, means that in aqueous solution membrane lipids self-associate into a tail-to-tail bilayer with their hydrophobic chains separated from the aqueous phase by their polar head groups.
Lipid bilayers separate the cell contents from the external environment and compartmentalize distinct cellular activities into organelles. These consist of a large variety of glycerophospholipids and sphingolipids. Membrane lipids usually have two hydrophobic acyl chains linked via glycerol or serine, to polar hydrophilic head groups (Fig. 2.2). This amphiphilic nature, with a ‘water-loving’ head and a ‘water-hating’ tail, means that in aqueous solution membrane lipids self-associate into a tail-to-tail bilayer with their hydrophobic chains separated from the aqueous phase by their polar head groups.
 Liposomes are spheres enclosed within a lipid bilayer. This is the most energetically favourable form for membrane lipids in solution. These have been used clinically to deliver more hydrophilic cargo, such as drugs or DNA, to cells.
Liposomes are spheres enclosed within a lipid bilayer. This is the most energetically favourable form for membrane lipids in solution. These have been used clinically to deliver more hydrophilic cargo, such as drugs or DNA, to cells.
 Plasma membranes are more complicated than liposomes. Their lipids are organized asymmetrically in the bilayer. For example, the outer leaflet of the plasma membrane is enriched in phosphatidyl-choline (PC) and the sphingolipids, whereas the inner leaflet is enriched in phosphatidyl-serine (PS) and phosphatidyl-ethanolamine (PE). This arrangement is necessary in normal physiology and in disease, not just for barrier function. For example, PC is extracted from the outer-leaflet of the canalicular membrane of hepatocytes to form the lipid/bile-salt micelles of bile. One of the sphingolipids, GM1-ganglioside, is the receptor for cholera toxin. The appearance of PS in the outer leaflet of the membrane is an early step in the apoptotic pathway and signals to macrophages to clear the dying cell, while PE, once cleaved by phospholipase, produces two signalling molecules as second messengers (see p. 25). Cholesterol is also an essential component of the plasma membrane and cannot be substituted by plant sterols, which have a subtly different shape. For this reason, the liver secretes plant sterols back into the gut.
Plasma membranes are more complicated than liposomes. Their lipids are organized asymmetrically in the bilayer. For example, the outer leaflet of the plasma membrane is enriched in phosphatidyl-choline (PC) and the sphingolipids, whereas the inner leaflet is enriched in phosphatidyl-serine (PS) and phosphatidyl-ethanolamine (PE). This arrangement is necessary in normal physiology and in disease, not just for barrier function. For example, PC is extracted from the outer-leaflet of the canalicular membrane of hepatocytes to form the lipid/bile-salt micelles of bile. One of the sphingolipids, GM1-ganglioside, is the receptor for cholera toxin. The appearance of PS in the outer leaflet of the membrane is an early step in the apoptotic pathway and signals to macrophages to clear the dying cell, while PE, once cleaved by phospholipase, produces two signalling molecules as second messengers (see p. 25). Cholesterol is also an essential component of the plasma membrane and cannot be substituted by plant sterols, which have a subtly different shape. For this reason, the liver secretes plant sterols back into the gut.
Membrane proteins
Cells can absorb gases or small hydrophobic compounds directly across the plasma membrane by passive diffusion, but membrane proteins are required to take-up hydrophilic nutrients or secrete hydrophilic products, to mediate cell–cell communication and to respond to endocrine signals. Membrane proteins can be integral to the membrane (i.e. their protein chain traverses the membrane one or multiple times) or they can be anchored to the membrane by an acyl chain (Fig. 2.2).
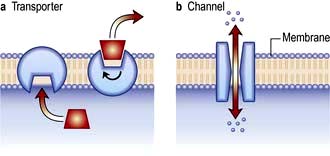
 Membrane channel proteins (Fig. 2.3): membrane proteins that form solute channels through the membrane can only work downhill and only to equilibrium. Solute actually moves down its electrochemical gradient, which is the combined force of the electric potential and the solute concentration gradient across the membrane. The bulk flow can be very high, the opening and closing of the channel can be regulated, and they can be selective for specific solutes. For example, the cystic fibrosis transmembrane regulator (CFTR; Fig. 2.22), the protein whose malfunction causes cystic fibrosis, is a chloride channel found on the apical surface of epithelial cells. CFTR functions to regulate the fluidity of the extra-epithelial mucous layer. When the channel opens, millions of negatively-charged chloride ions flow out of the cell down their electrochemical gradient. This induces positively-charged sodium ions to flow between the cells of the epithelium (via a paracellular pathway) to balance the electrical charge. Water follows the efflux of sodium chloride by osmosis, thus maintaining the fluidity of the mucus.
Membrane channel proteins (Fig. 2.3): membrane proteins that form solute channels through the membrane can only work downhill and only to equilibrium. Solute actually moves down its electrochemical gradient, which is the combined force of the electric potential and the solute concentration gradient across the membrane. The bulk flow can be very high, the opening and closing of the channel can be regulated, and they can be selective for specific solutes. For example, the cystic fibrosis transmembrane regulator (CFTR; Fig. 2.22), the protein whose malfunction causes cystic fibrosis, is a chloride channel found on the apical surface of epithelial cells. CFTR functions to regulate the fluidity of the extra-epithelial mucous layer. When the channel opens, millions of negatively-charged chloride ions flow out of the cell down their electrochemical gradient. This induces positively-charged sodium ions to flow between the cells of the epithelium (via a paracellular pathway) to balance the electrical charge. Water follows the efflux of sodium chloride by osmosis, thus maintaining the fluidity of the mucus.
 Transporters (Fig. 2.3): in contrast to channels, transporters have a low capacity and work by binding solute on one side of the membrane which induces a conformational change that exposes the solute binding site on the other side of the membrane for release.
Transporters (Fig. 2.3): in contrast to channels, transporters have a low capacity and work by binding solute on one side of the membrane which induces a conformational change that exposes the solute binding site on the other side of the membrane for release.
 Receptors: there are three major receptor categories: receptors that mediate endocytosis, anchorage receptors (e.g. integrins, see p. 23) and signalling receptors (see cell signalling p. 24). There are two forms of receptor-mediated endocytosis:
Receptors: there are three major receptor categories: receptors that mediate endocytosis, anchorage receptors (e.g. integrins, see p. 23) and signalling receptors (see cell signalling p. 24). There are two forms of receptor-mediated endocytosis:
Organelles
Cytoplasmic organelles
 Endoplasmic reticulum (ER) is an array of interconnecting tubules or flattened sacs (cisternae) that is contiguous with the outer nuclear membrane (Fig. 2.1). There are three types of ER:
Endoplasmic reticulum (ER) is an array of interconnecting tubules or flattened sacs (cisternae) that is contiguous with the outer nuclear membrane (Fig. 2.1). There are three types of ER:
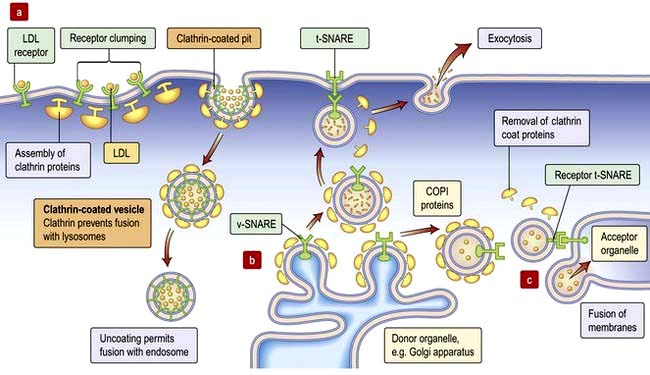
 Golgi apparatus has flattened cisternae similar to those of the ER but arranged in a stack (Fig. 2.1). Vesicles that bud from the ER with cargo destined for secretion, for the plasma membrane or for other organelles, fuse with the Golgi stack. The proteins, lipids and sterols synthesized in the ER are exported to the Golgi apparatus to complete maturation (e.g. the final stages of membrane protein glycosylation occurs here). The mature products are then sorted into vesicles that bud from the Golgi for transport to their final destination (Fig. 2.4b,c). Mutation in the Golgin protein GMAP-210, with a probable role in tethering of the Golgi cisternae, causes achondrogenesis type 1A, where Golgi architecture is disrupted, particularly in bone cells.
Golgi apparatus has flattened cisternae similar to those of the ER but arranged in a stack (Fig. 2.1). Vesicles that bud from the ER with cargo destined for secretion, for the plasma membrane or for other organelles, fuse with the Golgi stack. The proteins, lipids and sterols synthesized in the ER are exported to the Golgi apparatus to complete maturation (e.g. the final stages of membrane protein glycosylation occurs here). The mature products are then sorted into vesicles that bud from the Golgi for transport to their final destination (Fig. 2.4b,c). Mutation in the Golgin protein GMAP-210, with a probable role in tethering of the Golgi cisternae, causes achondrogenesis type 1A, where Golgi architecture is disrupted, particularly in bone cells.
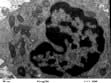
 Lysosomes mature from vesicles (endosomes) that bud from the Golgi. They contain digestive enzymes such as lipases, proteases, nucleases and amylases that work in an acidic environment. The membrane of the lysosome therefore includes a proton ATPase pump to acidify the lumen of the organelle. Lysosomes fuse with phagocytotic vesicles to digest their contents. This is crucial to the function of macrophages and polymorphs (neutrophils and eosinophils) in killing and digesting infective agents, in tissue remodelling during development, and osteoclast remodelling of bone. Not surprisingly, many metabolic disorders result from impaired lysosomal function (p. 1040).
Lysosomes mature from vesicles (endosomes) that bud from the Golgi. They contain digestive enzymes such as lipases, proteases, nucleases and amylases that work in an acidic environment. The membrane of the lysosome therefore includes a proton ATPase pump to acidify the lumen of the organelle. Lysosomes fuse with phagocytotic vesicles to digest their contents. This is crucial to the function of macrophages and polymorphs (neutrophils and eosinophils) in killing and digesting infective agents, in tissue remodelling during development, and osteoclast remodelling of bone. Not surprisingly, many metabolic disorders result from impaired lysosomal function (p. 1040).
 Peroxisomes contain enzymes for the catabolism of long-chain fatty acids and other organic substrates like bile acids and D-amino acids. Hydrogen peroxide (H2O2), a by-product of these reactions, is a highly reactive oxidizing agent, so peroxisomes also contain catalase to detoxify the peroxide. Catalase can reduce H2O2 to water while oxidizing harmful phenols and alcohols thus beginning their detoxification. Peroxisome dysfunction can lead to rare metabolic disorders such as leukodystrophies and rhizomelic dwarfism.
Peroxisomes contain enzymes for the catabolism of long-chain fatty acids and other organic substrates like bile acids and D-amino acids. Hydrogen peroxide (H2O2), a by-product of these reactions, is a highly reactive oxidizing agent, so peroxisomes also contain catalase to detoxify the peroxide. Catalase can reduce H2O2 to water while oxidizing harmful phenols and alcohols thus beginning their detoxification. Peroxisome dysfunction can lead to rare metabolic disorders such as leukodystrophies and rhizomelic dwarfism.
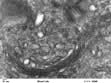
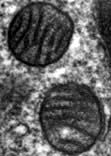
 Mitochondria are the engines of the cell, providing energy in the form of ATP. Mitochondria can be small, discrete and few in number in cells with low energy demand, or large and abundant in cells with a high energy demand like hepatocytes or muscle cells. The mitochondrion has its own genome encoding 13 proteins. The other proteins (~1000) required for mitochondrial function are encoded by the nuclear genome and imported into the mitochondrion. The mitochondrion has a double membrane surrounding a central matrix. The central matrix contains the enzymes for the Krebs cycle, which accepts the products of sugar and fatty acid catabolism and uses it to produce cofactors that donate their electrons into the electron transport chain of the inner membrane (see pp. 20, 31). The inner membrane is highly folded into cristae to increase its effective surface area. The protein complexes of the electron transport chain accept and donate electrons in redox reactions, releasing energy to efflux protons (H+) into the inter-membrane space. ATP synthase, another integral membrane protein, uses this H+ electrochemical gradient to drive formation of ATP. Mitochondria have many additional functions, including roles in apoptosis (see p. 32) and supply of substrates for biosynthesis. Mitochondria are also necessary for the synthesis of porphyrin, deficiency of which causes a range of diseases collectively called porphyrias (p. 1043).
Mitochondria are the engines of the cell, providing energy in the form of ATP. Mitochondria can be small, discrete and few in number in cells with low energy demand, or large and abundant in cells with a high energy demand like hepatocytes or muscle cells. The mitochondrion has its own genome encoding 13 proteins. The other proteins (~1000) required for mitochondrial function are encoded by the nuclear genome and imported into the mitochondrion. The mitochondrion has a double membrane surrounding a central matrix. The central matrix contains the enzymes for the Krebs cycle, which accepts the products of sugar and fatty acid catabolism and uses it to produce cofactors that donate their electrons into the electron transport chain of the inner membrane (see pp. 20, 31). The inner membrane is highly folded into cristae to increase its effective surface area. The protein complexes of the electron transport chain accept and donate electrons in redox reactions, releasing energy to efflux protons (H+) into the inter-membrane space. ATP synthase, another integral membrane protein, uses this H+ electrochemical gradient to drive formation of ATP. Mitochondria have many additional functions, including roles in apoptosis (see p. 32) and supply of substrates for biosynthesis. Mitochondria are also necessary for the synthesis of porphyrin, deficiency of which causes a range of diseases collectively called porphyrias (p. 1043).
The cytoskeleton
 Microtubules (20–25 nm diameter) are polymers of α and β tubulin. These tubular structures resist bending and stretching, and are polar with plus and minus ends. They emanate from the microtubule organizing centre (MTOC), a complex of centrioles, γ-tubulin and other proteins, with their plus ends extending into the cell. At their plus ends repeated cycles of assembly and disassembly permit rapid changes in length. Microtubules form a ‘highway’, transporting organelles and vesicles through the cytoplasm. The two major microtubule-associated motor proteins (kinesin and dynein) allow movement of cargo to the plus and minus ends, respectively. During cell division the MTOC forms the mitotic spindle (see p. 28). Drugs that disrupt microtubule assembly (e.g. colchicine and vinca alkaloids) or stabilize microtubules (taxanes) preferentially kill dividing cells by preventing mitosis.
Microtubules (20–25 nm diameter) are polymers of α and β tubulin. These tubular structures resist bending and stretching, and are polar with plus and minus ends. They emanate from the microtubule organizing centre (MTOC), a complex of centrioles, γ-tubulin and other proteins, with their plus ends extending into the cell. At their plus ends repeated cycles of assembly and disassembly permit rapid changes in length. Microtubules form a ‘highway’, transporting organelles and vesicles through the cytoplasm. The two major microtubule-associated motor proteins (kinesin and dynein) allow movement of cargo to the plus and minus ends, respectively. During cell division the MTOC forms the mitotic spindle (see p. 28). Drugs that disrupt microtubule assembly (e.g. colchicine and vinca alkaloids) or stabilize microtubules (taxanes) preferentially kill dividing cells by preventing mitosis.
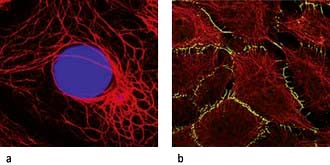
 Intermediate filaments (~10 nm) form a network around the nucleus extending to the periphery of the cell. They make cell-to-cell contacts with adjacent cells via desmosomes, and with basement matrix via hemidesmosomes (Fig. 2.5; see also Fig. 24.27). Their function appears to be structural integrity; they are prominent in cellular tissues under stress and their disruption in genetic disease can cause structural defects or cell collapse. More than 40 different types of proteins polymerize to form intermediate filaments specific to particular cell types. For example keratin intermediate fibres are only found in epithelial cells whilst vimentin is in mesothelial (fibroblastic) cells. However, lamin intermediate filaments form the nuclear membrane skeleton in most cells.
Intermediate filaments (~10 nm) form a network around the nucleus extending to the periphery of the cell. They make cell-to-cell contacts with adjacent cells via desmosomes, and with basement matrix via hemidesmosomes (Fig. 2.5; see also Fig. 24.27). Their function appears to be structural integrity; they are prominent in cellular tissues under stress and their disruption in genetic disease can cause structural defects or cell collapse. More than 40 different types of proteins polymerize to form intermediate filaments specific to particular cell types. For example keratin intermediate fibres are only found in epithelial cells whilst vimentin is in mesothelial (fibroblastic) cells. However, lamin intermediate filaments form the nuclear membrane skeleton in most cells.
 Microfilaments (3–6 nm) are polymers of actin, one of the most abundant proteins in all cells. The actin microfilament network controls cell shape, prevents cellular deformation, is involved in cell–cell and cell–matrix adhesion, in cell movements such as crawling and cytokinesis (cell division), and in intracellular vesicle transport. Bundles of actin filaments form the structural core of cellular protrusions such as microvilli, lamellipodia and filopodia (see below). Actin microfilament bundles within the cell can associate with myosin II to form contractile stress fibres, similar to muscle sarcomeres. Stress fibres are often found as circumferential belts around the apical surfaces of epithelial cells where cells associate with adjacent cells via adherens junctions, permitting reaction to external stresses as a cellular sheet. Stress fibres also form where actin interacts via accessory proteins with the extracellular matrix at sites of focal adhesion (see Fig. 2.8c). This occurs during cell movements during inflammation, wound healing and metastasis. During cytokinesis actin-myosin II bundles form the contractile ring separating dividing cells. Like microtubules, microfilaments are polar, so can be used to transport secretory vesicles, endosomes and mitochondria, powered by motor proteins, including myosin I and V.
Microfilaments (3–6 nm) are polymers of actin, one of the most abundant proteins in all cells. The actin microfilament network controls cell shape, prevents cellular deformation, is involved in cell–cell and cell–matrix adhesion, in cell movements such as crawling and cytokinesis (cell division), and in intracellular vesicle transport. Bundles of actin filaments form the structural core of cellular protrusions such as microvilli, lamellipodia and filopodia (see below). Actin microfilament bundles within the cell can associate with myosin II to form contractile stress fibres, similar to muscle sarcomeres. Stress fibres are often found as circumferential belts around the apical surfaces of epithelial cells where cells associate with adjacent cells via adherens junctions, permitting reaction to external stresses as a cellular sheet. Stress fibres also form where actin interacts via accessory proteins with the extracellular matrix at sites of focal adhesion (see Fig. 2.8c). This occurs during cell movements during inflammation, wound healing and metastasis. During cytokinesis actin-myosin II bundles form the contractile ring separating dividing cells. Like microtubules, microfilaments are polar, so can be used to transport secretory vesicles, endosomes and mitochondria, powered by motor proteins, including myosin I and V.
Cell shape and motility
The cytoskeleton determines cell shape and surface structures.
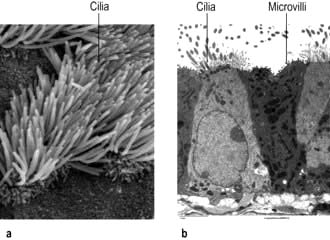
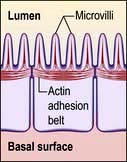
Microvilli. The apical surface of some epithelial cells is covered in tiny microvilli (~1 µm long) forming a brush border of thousands of small finger-like projections of the plasma membrane that increase the surface area for uptake or efflux (Fig. 2.6). At their core are 20–30 cross-linked actin microfilaments.
Motile cilia are also fine, finger-like protrusions but these are longer (~10–20 µm long) (Fig. 2.6). At their core is an axoneme, a bundle of nine cross-linked tubulin microtubule doublets surrounding a central pair. The action of the motor domain dynein serves to bend the cilium. Neighbouring cilia tend to beat in unison generating waves of motion that move fluid over the cell surface in the gut and airways (see Fig. 15.9), and also in the fallopian tubes.
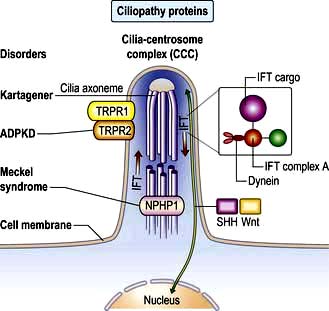
Non-motile or primary cilia. Most cells also have a single primary cilium. These cilia have a variant axoneme with no central pair of microtubules and while they have dynein they are non-motile (the dynein is used to traffic cargo along the axoneme). Primary cilia are used for signalling during development and in the adult. Other related non-motile cilia are found in specialized cells, e.g. in the photoreceptors of the retina, the sensory neurones of the olfactory system, and in the sensory hair cells of the cochlea. A range of human ciliopathies (Fig. 2.7) have been described with pleiotropic symptoms depending on which cilia are affected. These include polycystic kidney disease, Bardet–Biedl syndrome (p. 1007), Joubert’s syndrome and Ellis–van Creveld syndrome.
 Filopodia: if remodelled essentially in one dimension into a long actin filament, the leading edge of the plasma membrane is pushed forward as spikes, similar to long thin villi.
Filopodia: if remodelled essentially in one dimension into a long actin filament, the leading edge of the plasma membrane is pushed forward as spikes, similar to long thin villi.
 Lamellipodia: if remodelled in two dimensions to form a network of cross-linked actin microfilaments, a broad flat skirt or lamellipodium is formed.
Lamellipodia: if remodelled in two dimensions to form a network of cross-linked actin microfilaments, a broad flat skirt or lamellipodium is formed.
 Pseudopodia: are more three-dimensional projections as the actin cytoskeleton is remodelled into a gel-like lattice.
Pseudopodia: are more three-dimensional projections as the actin cytoskeleton is remodelled into a gel-like lattice.
Movement. A similar mechanism involving the coordinated remodelling of the cytoskeleton and the formation and release of cell adhesions underlies all three modes of migration. Essentially, actin is polymerized at the leading edge extending the plasma membrane forward. New adhesions are formed with the substratum (cells and/or extracellular matrix) at the leading edge to provide purchase. Release of attachments and depolymerization of the actin filaments at the trailing edge then allows the cell to move forward. Myosin and myosin motor proteins may also be involved at the trailing edge providing the tractive force to pull the cell body forward. The complex coordination of these processes is controlled via signalling pathways involving members of the Rho protein family of GTPases (see p. 21). Key signalling targets are the WASp family of proteins which stimulate actin polymerization. The significance of cell motility in humans is illustrated by mutation of the WASp expressed in blood cell lineages, which causes Wiskott–Aldrich syndrome (p. 66), and is characterized by severe immunodeficiency and thrombocytopenia (platelet deficiency).
The cell and its environment
 Epithelial tissues comprise layers of cells held tightly together by intercellular junctions and are usually separated from underlying tissue by specialized ECM called basal lamina. Epithelia cover surfaces (e.g. epidermis, tongue surface) and line passageways (airways, digestive tract, blood vessels), providing protection and regulating absorption and secretion.
Epithelial tissues comprise layers of cells held tightly together by intercellular junctions and are usually separated from underlying tissue by specialized ECM called basal lamina. Epithelia cover surfaces (e.g. epidermis, tongue surface) and line passageways (airways, digestive tract, blood vessels), providing protection and regulating absorption and secretion.
 Connective tissues provide support to other tissues and give organs shape. They comprise cells (fibroblasts) embedded within ECM such as the matrix of bone, dermis of skin and the fluid matrix of blood.
Connective tissues provide support to other tissues and give organs shape. They comprise cells (fibroblasts) embedded within ECM such as the matrix of bone, dermis of skin and the fluid matrix of blood.
Extracellular matrix
The gel or ground substance of the ECM is made from polysaccharides (glycosaminoglycans or GAGs), usually bound to proteins to form proteoglycans (p. 494). These are a diverse group of molecules conferring different matrix properties in different tissues. They form hydrated gels which can resist compression yet permit diffusion of metabolites and signalling molecules.
 Hyaluronan, a very large hydrated GAG, is secreted into the joint space in synovial joints (p. 493), where it aids lubrication and helps reduce compressive forces.
Hyaluronan, a very large hydrated GAG, is secreted into the joint space in synovial joints (p. 493), where it aids lubrication and helps reduce compressive forces.
 Aggrecan, a very large proteoglycan, forms part of the articular cartilage of joints (p. 494) also contributing to compression resistance.
Aggrecan, a very large proteoglycan, forms part of the articular cartilage of joints (p. 494) also contributing to compression resistance.
 Decorin is a much smaller proteoglycan from loose connective tissue of skin with both structural and signalling function (through binding and regulating growth factor activity).
Decorin is a much smaller proteoglycan from loose connective tissue of skin with both structural and signalling function (through binding and regulating growth factor activity).
Fibrous proteins of ECM (p. 495) include collagens and tropoelastin, which polymerize into collagen and elastin fibres, and fibronectin which is insoluble in many tissues but soluble in plasma. Collagen provides tensile strength, elastin confers elasticity, while the widely distributed fibronectin adheres to both cells and ECM, and thus positions cells within the ECM. Collagens, the most abundant proteins in the body, are widely distributed and play a structural role in skin and bone, where collagen defects and disorders often manifest. Elastin fibres are abundant in arteries, lung and skin. Elastic fibres have a fibrillin sheath and fibrillin mutations underlie Marfan’s syndrome (p. 760). The ECM can be degraded and remodelled by proteins of the matrix metalloproteinase (MMP) family. These are needed for angiogenesis and morphogenesis and are also involved in the pathophysiology of cancer, cirrhosis and arthritis.
Basal lamina or basement membrane (lamina propria) is a specialized form of ECM, which separates cells from underlying tissue and provides a supportive, anchoring and protective role. Basal lamina can also act as molecular filters (e.g. glomerular filtration barrier, p. 636) and mediate signalling between adjacent tissues (e.g. epidermal-dermal signalling in skin). Type IV collagen, heparan sulphate proteoglycan, laminin and nidogen are key basal lamina proteins. Inherited abnormalities in these proteins cause skin blistering diseases (see Fig. 24.27). Breach of the basal lamina by invading cancer cells is a key stage in progression of epithelial carcinoma in situ to a malignant carcinoma.
Cell–cell adhesion
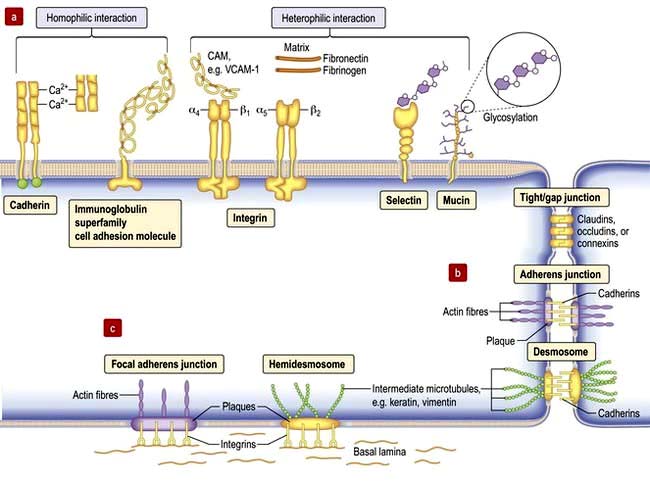
Cell–cell adhesion proteins (Fig. 2.8a)
 Immunoglobulin-like cell adhesion molecules (iCAMs or CAMs) (Fig. 2.8a) are structurally related to antibodies. The neural cell adhesion molecule (N-CAM) is found predominantly in the nervous system. It mediates a homophilic (like-like) adhesion. When bound to an identical molecule on another cell, N-CAM can also associate laterally with a fibroblast growth factor receptor and stimulate its tyrosine kinase activity to induce neurite growth thus triggering cellular responses by indirect activation of the recipient.
Immunoglobulin-like cell adhesion molecules (iCAMs or CAMs) (Fig. 2.8a) are structurally related to antibodies. The neural cell adhesion molecule (N-CAM) is found predominantly in the nervous system. It mediates a homophilic (like-like) adhesion. When bound to an identical molecule on another cell, N-CAM can also associate laterally with a fibroblast growth factor receptor and stimulate its tyrosine kinase activity to induce neurite growth thus triggering cellular responses by indirect activation of the recipient.
 Selectins. Unlike most adhesion molecules which bind to other proteins, the selectins interact with carbohydrate ligands or mucin complexes on leucocytes and endothelial cells (vascular and haematological systems). Leucocyte-selectin (CD62L) mediates the homing of lymphocytes to lymph nodes. Endothelial-selectin (CD62E) is expressed after activation by inflammatory cytokines; the small basal amount of E-selectin in many vascular beds appears to be necessary for the migration of leucocytes. Platelet-selectin (CD62P) is stored in the alpha granules of platelets and the Weibel–Palade bodies of endothelial cells, but it moves rapidly to the plasma membrane upon stimulation of these cells. All three selectins play a part in leucocyte rolling (p. 63).
Selectins. Unlike most adhesion molecules which bind to other proteins, the selectins interact with carbohydrate ligands or mucin complexes on leucocytes and endothelial cells (vascular and haematological systems). Leucocyte-selectin (CD62L) mediates the homing of lymphocytes to lymph nodes. Endothelial-selectin (CD62E) is expressed after activation by inflammatory cytokines; the small basal amount of E-selectin in many vascular beds appears to be necessary for the migration of leucocytes. Platelet-selectin (CD62P) is stored in the alpha granules of platelets and the Weibel–Palade bodies of endothelial cells, but it moves rapidly to the plasma membrane upon stimulation of these cells. All three selectins play a part in leucocyte rolling (p. 63).
 Integrins are membrane glycoproteins with α and β subunits which exist as active and inactive forms. The amino acid sequence arginine–glycine–aspartic acid (RGD) is a potent recognition system for integrin binding
Integrins are membrane glycoproteins with α and β subunits which exist as active and inactive forms. The amino acid sequence arginine–glycine–aspartic acid (RGD) is a potent recognition system for integrin binding
Tight junctions (zonula occludens)
These are mediated by the integral membrane proteins, claudins and occludens; they hold cells together. They form at the top (apical) side of epithelial cells including intestinal, skin and kidney cells, and endothelial cells of blood vessels (Fig. 2.8) to provide a regulated barrier to the movement of ions and solutes through the epithelia or endothelia but also between cells (paracellular transport). Tight junctions also confer polarity to cells by acting as a gate between the apical and the baso-lateral membranes, preventing diffusion of membrane lipids and proteins. Twenty-four claudins (the protein in the junction) are differentially expressed in different cell types to regulate paracellular transport. For example, changes in claudin expression in the kidney nephron correlate with permeability changes. Mutations in claudins 16 (previously named parcellin-1) and 19, expressed in the thick ascending limb in the loop of Henle in the kidney, cause an inherited renal disorder, familial hypomagnesaemia with hypercalciuria and nephrocalcinosis (FHHNC; p. 657).
Gap junctions
Gap junctions (Fig. 2.8) allow low molecular weight substances to pass directly between cells, permitting metabolic and electric coupling (e.g. in cardiomyocytes). Protein channels made of six connexin proteins (as well as claudins and occludens) are aligned between adjacent cells and allow the passage of solutes up to 1000 kDa (e.g. amino acids, sugars, ions, chemical messengers). The channels are regulated by many factors such as intracellular Ca2+, pH, voltage. Gap junctions form in almost all interacting cells, but connexin family members are differentially expressed. Mutant connexins cause many inherited disorders, such as the X-linked form of Charcot–Marie–Tooth disease (GJB1; p. 1147) and are also a major cause of genetic hearing loss (GJB2).
Adherens junctions
Adherens junctions are multiprotein intercellular adhesive structures, prominent in epithelial tissues (Fig. 2.8b). They attach principally to actin microfilaments inside the cell with the aid of multiple additional proteins, and also attach and stabilize microtubules. At the apical sides of epithelial cells a prominent type of adherens junction, the zonula adherens, attaches to the circumferential actin stress fibres. The fascia adherens in cardiac muscle is also an adherens junction. Transmembrane proteins of the cadherin family provide the adhesion through interaction of their extracellular domains. Downregulation of cadherins is a feature of cancer progression in many cells.
Desmosomes (macula adherens)
Desmosomes provide strong attachment between cells and are prominent in tissues subject to stress such as skin and cardiac muscle (see Fig. 2.5, Fig. 2.8b and Fig. 24.1). Like adherens junctions, they are multiprotein complexes, where adhesion is provided by transmembrane cadherin proteins, desmogleins and desmocollins. However, within the cell desmosomes interact principally with intermediate filaments rather than microfilaments and microtubules. Germline mutations in genes encoding desmosomes are a cause of cardiomyopathy with/without cutaneous features and in pemphigus vulgaris and pemphigus foliaceus (p. 1222).
Basement membrane adhesion
Cells adhere (Fig. 2.8c) to non-basal lamina ECM via secreted proteins such as fibronectin and collagen, and to basal lamina proteins via focal adhesion and hemidesmosome multiprotein complexes (e.g. keratin or vimentin). Here, integrins replace cadherins as surface adhesion molecules as the key adhesive proteins. Integrins are transmembrane sensors or receptors, which change shape upon binding to ECM, a process called ‘outside-in’ signalling. Inside the cell, integrins interact with the cytoskeleton and a complex array of over 150 proteins that influence intracellular signalling pathways affecting proliferation, survival, shape, mobility and gene expression.
 Outside-in signalling: forms the basis for anoikis or apoptotic death, such as occurs in cancer cells that inappropriately lose cell-substratum adhesion.
Outside-in signalling: forms the basis for anoikis or apoptotic death, such as occurs in cancer cells that inappropriately lose cell-substratum adhesion.
 Inside-out signalling: intracellular changes can also be communicated extracellularly via integrins whereby intracellular changes cause integrins to change from an inactive to an actively adhesive conformation. This ‘inside-out’ signalling occurs when platelet integrins glycoprotein IIb-IIIa (GPIIb-IIa) are activated to bind fibrinogen at sites of vessel injury, resulting in platelet aggregation (p. 415 and Fig. 8.41).
Inside-out signalling: intracellular changes can also be communicated extracellularly via integrins whereby intracellular changes cause integrins to change from an inactive to an actively adhesive conformation. This ‘inside-out’ signalling occurs when platelet integrins glycoprotein IIb-IIIa (GPIIb-IIa) are activated to bind fibrinogen at sites of vessel injury, resulting in platelet aggregation (p. 415 and Fig. 8.41).
Defective integrins are associated with many immunological and clotting disorders such as Bernard–Soulier syndrome and Glanzmann’s thrombasthenia (p. 420).
FURTHER READING
De Matteis MA, Luini A. Mendelian disorders of membrane trafficking. N Engl J Med 2011; 365:927–928.
Jean C, Gravelle P, Fournie JJ et al. Influence of stress on extracellular matrix and integrin biology. Oncogene 2011; 30:2697–2706.
Thomason HA, Scothern A, McHarg S et al. Desmosomes: adhesive strength and signalling in health and disease. Biochem J 2010; 429:419–433.
Cellular mechanisms
Cell signalling
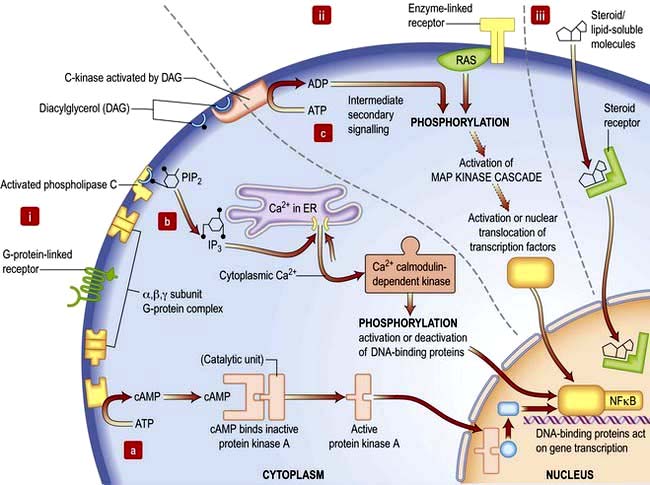
Signalling or communication between cells is often via extracellular molecules or ligands which can be proteins (e.g. hormones, growth factors), small molecules (e.g. lipid-soluble steroid hormones such as oestrogen and testosterone) or dissolved gases such as nitric oxide. The signal is usually received by membrane protein receptors, although some signals such as steroid hormones, enter the target cell where they interact with intracellular receptors (Fig. 2.9). Some signalling, especially in the immune system, relies on cell–cell contact, where the signalling molecule (ligand) and receptor are on adjacent cells.
Receptors transduce signals across the membrane to an intracellular pathway or second messengers to change cell behaviour, often ultimately affecting gene expression (Figs 2.9, 2.10). The membrane-bound receptors fall into three main groups based on downstream signalling pathways:
 Ion channel linked receptors (voltage or ligand activated ion channels; see Fig. 2.3). At synaptic junctions between neurones (Fig. 22.1), these receptors open in response to neurotransmitters such as glutamate, epinephrine (adrenaline) or acetylcholine to cause a rapid depolarization of the membrane.
Ion channel linked receptors (voltage or ligand activated ion channels; see Fig. 2.3). At synaptic junctions between neurones (Fig. 22.1), these receptors open in response to neurotransmitters such as glutamate, epinephrine (adrenaline) or acetylcholine to cause a rapid depolarization of the membrane.
 G-protein-linked receptors such as the odorant and light (opsin) family of receptors belong to a large family of seven-pass transmembrane proteins (see Figs 2.2 and 2.9). On activation by ligand G-protein-linked receptors bind a GTP-binding protein (G-protein), which activates adjacent enzyme complexes or ion channels (Figs 2.9 and 22.1). The adjacent enzymes can be adenylcyclase (see below).
G-protein-linked receptors such as the odorant and light (opsin) family of receptors belong to a large family of seven-pass transmembrane proteins (see Figs 2.2 and 2.9). On activation by ligand G-protein-linked receptors bind a GTP-binding protein (G-protein), which activates adjacent enzyme complexes or ion channels (Figs 2.9 and 22.1). The adjacent enzymes can be adenylcyclase (see below).
 Enzyme-linked receptors (Figs 2.2 and 2.9) typically have an extracellular ligand-binding domain, a single transmembrane-spanning region, and a cytoplasmic domain that has intrinsic enzyme activity or which will bind and activate other membrane-bound or cytoplasmic enzyme complexes. This group of receptors is highly variable but many have kinase activity or associate with kinases, which act by phosphorylating substrate proteins usually on a tyrosine (e.g. the platelet-derived growth factor (PDGF) receptor) or a serine/threonine (e.g. the transforming growth factor-beta (TGF-β) receptor).
Enzyme-linked receptors (Figs 2.2 and 2.9) typically have an extracellular ligand-binding domain, a single transmembrane-spanning region, and a cytoplasmic domain that has intrinsic enzyme activity or which will bind and activate other membrane-bound or cytoplasmic enzyme complexes. This group of receptors is highly variable but many have kinase activity or associate with kinases, which act by phosphorylating substrate proteins usually on a tyrosine (e.g. the platelet-derived growth factor (PDGF) receptor) or a serine/threonine (e.g. the transforming growth factor-beta (TGF-β) receptor).
Signal transduction
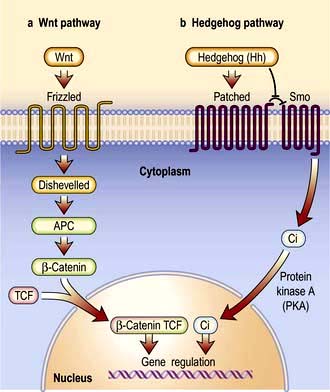
Signal transduction from the receptor to the site of action in the cell is mediated by small signalling molecules called second messengers, or by signalling proteins (Fig. 2.9). Changes to activity of signalling proteins by acquired mutation occur in cancer, and many anti-cancer drugs target signalling pathways. For example, the Hedgehog pathway is involved in human development, tissue repair and cancer (Fig. 2.10). Inhibitors of this pathway are being developed for therapeutic interventions. The Wnt pathway is also involved in bone formation (p. 550).
 Second messengers include cAMP and lipid-derived inositol triphosphate (IP3) and diacylglycerol (Fig. 2.9). These molecules diffuse from the receptor to bind and change the activity of downstream proteins propagating the signal. cAMP triggers a protein signalling cascade by activating a cAMP-dependent protein kinase. Diacylglycerol activates protein kinase C while IP3 mobilizes calcium from intracellular stores (e.g. from the ER; Fig. 14.9).
Second messengers include cAMP and lipid-derived inositol triphosphate (IP3) and diacylglycerol (Fig. 2.9). These molecules diffuse from the receptor to bind and change the activity of downstream proteins propagating the signal. cAMP triggers a protein signalling cascade by activating a cAMP-dependent protein kinase. Diacylglycerol activates protein kinase C while IP3 mobilizes calcium from intracellular stores (e.g. from the ER; Fig. 14.9).
 G-proteins or GTP-binding proteins are signalling proteins which switch between an active state when GTP is bound and an inactive state when bound to GDP. The most well-known members are the Ras superfamily, comprising Ras, Rho, Rab, Arf and Ran families. Activation of Ras members by somatic mutation is found in ~33% of human cancers. Ras members are often bound downstream of tyrosine kinase receptors, where they transmit signals by activating a cascade of downstream protein kinase activity (Fig. 2.9). Ras signalling molecules have roles in many cellular activities, including regulation of cell cycle, intracellular transport, and apoptosis.
G-proteins or GTP-binding proteins are signalling proteins which switch between an active state when GTP is bound and an inactive state when bound to GDP. The most well-known members are the Ras superfamily, comprising Ras, Rho, Rab, Arf and Ran families. Activation of Ras members by somatic mutation is found in ~33% of human cancers. Ras members are often bound downstream of tyrosine kinase receptors, where they transmit signals by activating a cascade of downstream protein kinase activity (Fig. 2.9). Ras signalling molecules have roles in many cellular activities, including regulation of cell cycle, intracellular transport, and apoptosis.
 Kinase and phosphatase signalling proteins are enzymes that phosphorylate or dephosphorylate residues on downstream proteins to alter their activity. Chains of kinase activity (phosphorylation cascades) consisting of sequential phosphorylation of proteins can transduce signals from the membrane receptor to the site of action in the cell. The tyrosine kinase receptors phosphorylate each other when ligand binding brings the intracellular receptor components into close proximity (see Fig. 2.9). The inner membrane and cytoplasmic targets of these activated receptor complexes are ras, protein kinase C and ultimately the MAP (mitogen-activated protein) kinase, Janus-Stat pathways or phosphorylation of IκB causing it to release its DNA-binding protein, nuclear factor kappa B (NFκB). For example, activated Ras binds and activates the kinase Raf, the first of a set of three mitogen-activated protein (MAP) kinases, which transmit signals by successive phosphorylation of target proteins which can ultimately effect transcription (Fig. 2.9). Kinases and phosphatases are frequently mutated in cancers. Somatic mutations in one Raf member, B-Raf, occur in ~60% of malignant melanomas (usually the mutation V600E) and are common in other cancers (p. 1225).
Kinase and phosphatase signalling proteins are enzymes that phosphorylate or dephosphorylate residues on downstream proteins to alter their activity. Chains of kinase activity (phosphorylation cascades) consisting of sequential phosphorylation of proteins can transduce signals from the membrane receptor to the site of action in the cell. The tyrosine kinase receptors phosphorylate each other when ligand binding brings the intracellular receptor components into close proximity (see Fig. 2.9). The inner membrane and cytoplasmic targets of these activated receptor complexes are ras, protein kinase C and ultimately the MAP (mitogen-activated protein) kinase, Janus-Stat pathways or phosphorylation of IκB causing it to release its DNA-binding protein, nuclear factor kappa B (NFκB). For example, activated Ras binds and activates the kinase Raf, the first of a set of three mitogen-activated protein (MAP) kinases, which transmit signals by successive phosphorylation of target proteins which can ultimately effect transcription (Fig. 2.9). Kinases and phosphatases are frequently mutated in cancers. Somatic mutations in one Raf member, B-Raf, occur in ~60% of malignant melanomas (usually the mutation V600E) and are common in other cancers (p. 1225).
Nuclear control
DNA and RNA structure
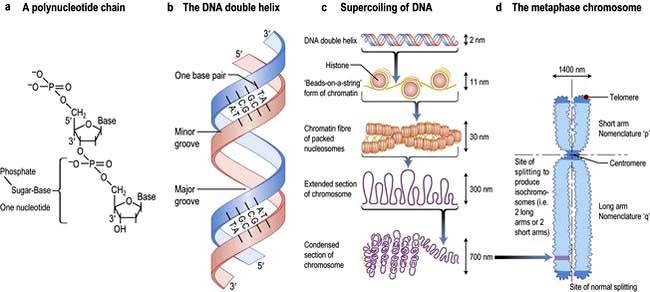
Hereditary information is contained in the sequence of the building blocks of double-stranded deoxyribonucleic acid (DNA) (Fig. 2.11). Each strand of DNA is made up of a deoxyribose-phosphate backbone and a series of purine (adenine (A) and guanine (G)) and pyrimidine (thymine (T) and cytosine (C)) bases, and because of the way the sugar phosphate backbone is chemically coupled, each strand has a polarity with a phosphate at one end (the 5′ end) and a hydroxyl at the other (the 3′ end). The two strands of DNA are held together by hydrogen bonds between the bases. A can only pair with T, and G can only pair with C, therefore each strand is the antiparallel complement of the other (Fig. 2.11b). This is key to DNA replication because each strand can be used as a template to synthesize the other.
The two strands twist to form a double helix with a major and a minor groove, and the large stretches of helical DNA are coiled around histone proteins to form nucleosomes (Fig. 2.11c). They can be condensed further into the chromosomes that can be visualized by light microscopy at metaphase (see below; Fig. 2.11, Fig. 2.19).
To express the information in the genome, cells first transcribe the code into the single strand ribonucleic acid (RNA). RNA is similar to DNA in that it comprises four bases A, G and C but with uracil (U) instead of T, and a sugar phosphate backbone with ribose instead of deoxyribose. Several types of RNA are made by the cell. Messenger RNA (mRNA) codes for proteins that are translated on ribosomes. Ribosomal RNA (rRNA) is a key catalytic component of the ribosome and amino acids are delivered to the nascent peptide chain on transfer RNA (tRNA) molecules. There are also a variety of RNAs that regulate gene expression or RNA processing. These include microRNA (miRNA) and small interfering RNA (siRNA) (see p. 27) that typically bind to a subset of mRNAs and inhibit their translation, or initiate their degradation, respectively. Other non-coding RNAs are involved in X-inactivation and telomere maintenance or RNA splicing and maturation.
DNA transcription
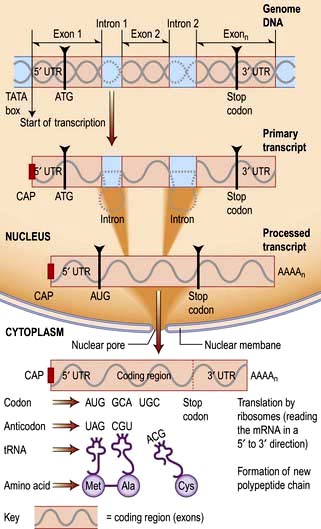
RNA is transcribed from the DNA template by an enzyme complex of more than one hundred proteins including RNA polymerase, transcription factors and enhancers. Promoter regions upstream of the gene dictate the start point and direction of transcription. The complex binds to the promoter region, the nucleosomes are remodelled to allow access, and a DNA helicase unwinds the double helix. RNA, like DNA, is synthesized in the 5′ to 3′ direction as ribonucleotides are added to the growing 3′ end of a nascent transcript. RNA polymerase does this by base-pairing the ribonucleotides to the DNA template strand running in the 3′ to 5′ direction. Messenger RNA is modified as it is synthesized (Fig. 2.12). It is capped at the 5′ end with a modified guanine that is required for efficient processing of the mRNA and efficient translation, and introns are spliced from the nascent chain. Finally, the 3′ of the mRNA is modified with up to 200 A nucleotides by the enzyme poly-A polymerase. This 3′ poly-A tail is essential for nuclear export (through the nuclear pores), stability and efficient translation into protein by the ribosome.
Human protein coding sequences (exons) are interrupted by intervening sequences that are non-coding (introns) at multiple positions (Fig. 2.12). These have to be spliced from the nascent message in the nucleus by an RNA/protein complex called a spliceosome. Differential splicing describes the process by which two or more introns and their intervening exons are spliced from the mRNA. This contributes significantly to the complexity of the human transcriptome as proteins translated from these messages lack particular domains. This exon skipping can produce different protein activities.
Control of gene expression
The genome of all cells in the body encodes the same genetic information, yet different cell types express a very different subset of proteins and respond to external signals to switch on a new set of genes or to switch off a pathway. Gene expression can be controlled at many steps from transcription to protein degradation. However, for many genes transcription is the key point of regulation. This is controlled primarily by proteins which bind to short sequences within the promoter regions that either repress or activate transcription, or to more distant sequences where proteins bind to enhance expression. These transcription factors and enhancers are often the end points of signalling pathways that transduce extracellular signals to changes in gene expression (Fig. 2.9).
Often this involves the translocation of an activated factor from the cytoplasm to the nucleus. In the nucleus the DNA binding proteins recognize the shape and position of hydrogen bond acceptor and donor groups within the major and minor grooves of the double helix (i.e. the double helix does not need to be unwound). There are several classes of DNA binding protein that differ in the protein structural motif that allows them to interact with the double helix. These primarily include helix-turn-helix, zinc finger and leucine zipper motifs, although protein loops and β-sheets are used by some proteins. More permanent control of gene expression patterns can be achieved epigenetically. These are modifications (typically methylation and/or acetylation) of the DNA, or the histones of the nucleosome, that silence genes. Epigenetic modification is also heritable meaning that a dividing liver cell, for example, can give rise to two daughter cells with the same epigenetic signals such that they express the appropriate transcriptome for a liver cell. Epigenetic change forms the basis of genetic imprinting (see p. 42).
Most of the genome is transcribed but only a minority of transcripts encode proteins (see Human Genetics, p. 34). The non-coding RNAs (ncRNAs) include a group that regulate gene expression (see DNA and RNA structure). miRNAs and siRNAs are short ncRNAs (19–29 bp) that are known to regulate expression of approximately 30% of genes by degradation of transcripts or repression of protein synthesis. With further annotation of the genome a growing range of additional regulatory ncRNA classes are being identified, many of which control gene expression by epigenetic mechanisms.