Paramyxoviridae
Robert A. Lamb
Griffith D. Parks
The Paramyxoviridae include some of the great and ubiquitous disease-causing viruses of humans and animals, including one of the most infectious viruses known (measles virus), some of the most prevalent viruses known (measles virus, parainfluenza viruses [PIVs], mumps virus, respiratory syncytial virus [RSV], and metapneumovirus), a virus that has been targeted by the World Health Organization for eradication (measles virus; however, to date, eradication has failed), a virus that has been eradicated (rinderpest virus), viruses that have a major economic impact on poultry rearing (Newcastle disease virus [NDV]), and many recently identified viruses (pinniped morbilliviruses, Hendra virus, Nipah virus, J virus and Beilong virus), some of which cause deadly diseases (Hendra and Nipah viruses). The Paramyxoviridae are enveloped negative-stranded RNA viruses that have special relationships with two other families of negative-strand RNA viruses, namely the Orthomyxoviridae (for the biological properties of the envelope glycoproteins) and the Rhabdoviridae (for the similarity of organization of the nonsegmented genome and its expression). The Paramyxoviridae are defined by having a protein (F) that causes viral–cell membrane fusion, in most cases at neutral pH. The genomic RNA of all negative-strand RNA viruses has to serve two functions: first as a template for synthesis of messenger RNAs (mRNAs) and second as a template for synthesis of the antigenome positive strand. Negative-strand RNA viruses encode and package their own RNA polymerase (RNAP); however, mRNAs are only synthesized once the virus has been uncoated in the infected cell. Viral replication occurs after synthesis of the mRNAs and requires the continuous synthesis of viral proteins. The newly synthesized antigenome positive strand serves as the template for further copies of the negative-strand genomic RNA.
Classification
The family Paramyxoviridae is classified into two subfamilies: the Paramyxovirinae and the Pneumovirinae. The Paramyxovirinae contains seven genera: Respirovirus, Rubulavirus, Morbillivirus, Henipavirus, Aquaparamyxovirus, Avulavirus, and Ferlavirus. The Pneumovirinae contains two genera Pneumovirus and Metapneumovirus. The classification is based on morphologic criteria, the organization of the genome, the biological activities of the proteins, and the sequence relationship of the encoded proteins now that all of the genome sequences have been obtained. The more recently identified tree shrew (Tupaia) paramyxovirus, J virus, Beilong virus, Salem virus, Menangle virus, Mossman virus, Fer-de-Lance virus, and Tioman virus have yet to be officially classified within the Paramyxovirinae by the International Committee on the Taxonomy of Viruses.
The morphologic distinguishing feature among enveloped viruses for the subfamily Paramyxovirinae is the size and shape of the nucleocapsids (diameter 18 nm, 1 μm in length, a pitch of 5.5 nm), which have a left-handed helical symmetry. The biological criteria are (a) antigenic cross-reactivity between members of a genus and (b) the presence (Respirovirus and Rubulavirus) or absence (Morbillivirus and Henipavirus) of neuraminidase (NA) activity. In addition, the differing coding potentials of the P genes are considered, and there is the presence of an extra gene (SH) in some rubulaviruses as well as J virus and Beilong virus. The pneumoviruses can be distinguished from Paramyxovirinae morphologically, as they contain narrower nucleocapsids. In addition, the Pneumovirinae have major differences in the number of encoded proteins and an attachment protein that is very different from that of Paramyxovirinae. Examples of members of various genera are shown in Table 33.1.
The Structure and Replication Strategy of the Paramyxoviridae
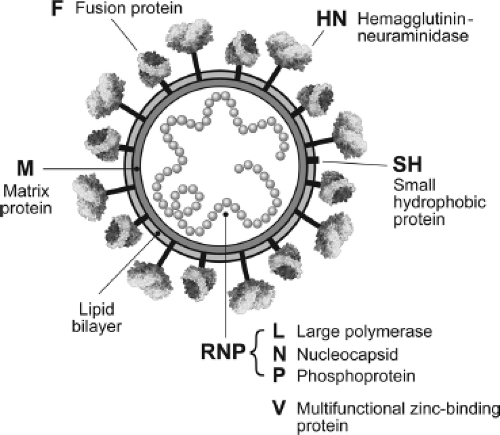
Paramyxoviruses contain nonsegmented single-stranded RNA genomes of negative polarity and replicate entirely in the cytoplasm. Their genomes are 15 to 19 kB in length, and the genomes contain 6 to 10 tandemly linked genes. A lipid envelope containing two surface glycoproteins (F and a second glycoprotein variously referred to as HN, or H or G) surrounds the virions. Inside the envelope lies a helical nucleocapsid core containing the RNA genome and the nucleocapsid (N), phospho- (P), and large (L) proteins, which initiate intracellular virus replication. Residing between the envelope and the core lies the viral matrix (M) protein that is important in virion architecture, and which is released from the core during virus entry. In addition to the genes encoding structural proteins, paramyxoviruses contain “accessory” genes that are found mostly as additional transcriptional units interspersed with the tandemly linked invariant genes. For the Paramyxovirinae, the accessory genes are found mostly as open reading frames (ORFs) that overlap within the P gene transcriptional unit.
Table 33.1 Examples of Members of the Family Paramyxoviridae | |||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
Intracellular replication of paramyxoviruses begins with the viral RNA–dependent RNAP (minimally a homo-tetramer of P and a single L protein) transcribing the N-encapsidated genome RNA (N:RNA) into 5′ capped and 3′ polyadenylated mRNAs. The viral RNA-dependent polymerase (vRNAP)begins RNA synthesis at the 3′ end of the genome and transcribes the genes into mRNAs in a sequential (and polar) manner by terminating and reinitiating at each of the gene junctions. The junctions consist of a gene-end (GE) sequence, at which polyadenylation occurs by the reiterative synthesis of adenylates directed by a template of four to seven uridylates (followed by release of the mRNA), a short nontranscribed intergenic (IG) region, and a gene-start (GS) sequence that specifies mRNA initiation as well as capping. The vRNAP occasionally fails to reinitiate the downstream mRNA at each junction, leading to the loss of transcription of further downstream genes; hence, there is a gradient of mRNA synthesis that is inversely proportional to the distance of the gene from the 3′ end of the genome. After primary transcription and translation, when sufficient amounts of unassembled N protein are present, viral RNA synthesis becomes coupled to the concomitant encapsidation of the nascent [+] RNA chain. Under these conditions, vRNAP ignores all of the junctions (and editing sites) to produce an exact complementary antigenome chain in a fully assembled nucleocapsid.
Virion Structure
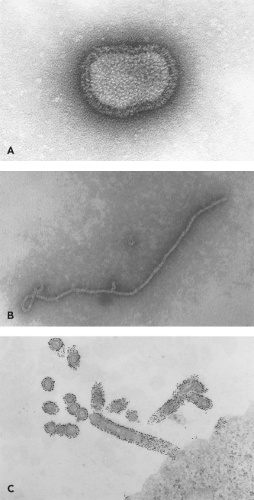
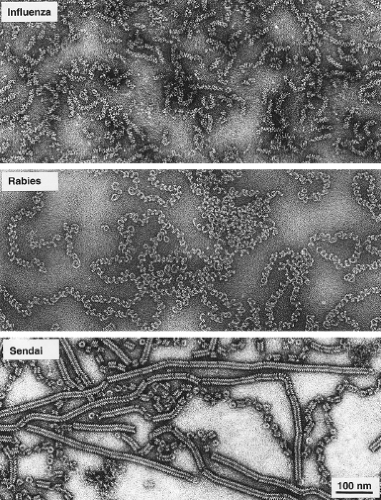
The Paramyxoviridae contain a lipid bilayer envelope that is derived from the plasma membrane of the host cell in which the virus is grown.59 Paramyxoviridae are generally spherical, 150 to 350 nm in diameter, but can be pleomorphic in shape, and filamentous forms can be observed. Inserted into the envelope are glycoprotein spikes that extend approximately 8 to 12 nm from the surface of the membrane, and that can be readily visualized by electron microscopy. Inside the viral membrane is the nucleocapsid core (sometimes referred to as the ribonucleoprotein [RNP] core) that contains the 15,000 to 19,000 nucleotide single-stranded RNA genome. Figure 33.1 shows a highly stylized schematic diagram of the virion. F and HN are trimers and tetramers, respectively. No attempt has been made to represent the real abundance of F, HN, SH, or N subunits in the virion. The pleomorphic nature of virus particles is illustrated in the electron micrograph in Figure 33.2, and a comparison of the RNPs of influenza virus, rabies virus, and Sendai virus is shown in Figure 33.3.
The helical nucleocapsid, rather than the free genome RNA, is the template for all RNA synthesis. For Sendai virus,
each nucleocapsid is composed of approximately 2,600 N, 300 P, and 50 L proteins.214 The N and genome RNA together form a core structure, to which the P and L proteins are attached. This nucleocapsid core is remarkably stable, as it withstands the high salt and gravity forces of cesium chloride (CsCl) density gradient centrifugation. In the electron micrograph of nucleocapsids, the P and L proteins are not observed and have only been visualized with the aid of antibodies.327 Holo-nucleocapsids (N:RNA plus P and L) have the capacity to transcribe mRNAs in vitro, presumably mimicking primary transcription in infected cells, and they are thought to be the minimum unit of infectivity.
each nucleocapsid is composed of approximately 2,600 N, 300 P, and 50 L proteins.214 The N and genome RNA together form a core structure, to which the P and L proteins are attached. This nucleocapsid core is remarkably stable, as it withstands the high salt and gravity forces of cesium chloride (CsCl) density gradient centrifugation. In the electron micrograph of nucleocapsids, the P and L proteins are not observed and have only been visualized with the aid of antibodies.327 Holo-nucleocapsids (N:RNA plus P and L) have the capacity to transcribe mRNAs in vitro, presumably mimicking primary transcription in infected cells, and they are thought to be the minimum unit of infectivity.
When negatively stained preparations of paramyxovirus nucleocapsids are viewed in the electron microscope, the most tightly coiled forms resemble the Tobamovirus tobacco mosaic virus (TMV)—a relatively rigid coiled rod 18 nm in diameter, with a central hollow core of 4 nm and a helical pitch of nearly 5 nm.68,115 Unlike TMV, however, in which the nucleocapsid must disassemble so its positive RNA genome can function as a template, paramyxovirus nucleocapsids function without disassembling their nucleocapsids.
Sendai virus nucleocapsids exist in several distinct morphologic states at normal salt concentration.101,156 The most prevalent form in negatively stained preparations is the most tightly coiled one, with a helical pitch of 5.3 nm. Two other forms—one with a slightly larger pitch of 6.8 nm and another with a much larger pitch of 37.5 nm—have also been noted.
The fact that no structures of intermediate pitch have been found indicates that these are distinct states. It is thought that the template is copied without dissociation of N protein from the nucleocapsid and the uncoiling of the nucleocapsid may be necessary for the polymerase to gain access to the RNA bases. It is possible that the vRNAP traverses the nucleocapsid template by uncoiling the helix in front of it and recoiling it once the polymerase has passed a given position, much the same as cellular RNAP generates its template “bubble” in traversing double-stranded DNA (dsDNA).
The fact that no structures of intermediate pitch have been found indicates that these are distinct states. It is thought that the template is copied without dissociation of N protein from the nucleocapsid and the uncoiling of the nucleocapsid may be necessary for the polymerase to gain access to the RNA bases. It is possible that the vRNAP traverses the nucleocapsid template by uncoiling the helix in front of it and recoiling it once the polymerase has passed a given position, much the same as cellular RNAP generates its template “bubble” in traversing double-stranded DNA (dsDNA).
As expected, the diameter of the nucleocapsid decreases as the pitch increases and the nucleocapsid lengthens; for Sendai virus, the diameter is 3.5 nm less for the 6.8 nm form than for the 5.3 nm pitch form. These latter values are very similar to those of Pneumovirus nucleocapsids, which also have a pitch of 7 nm. As discussed earlier, these differences in nucleocapsid morphology are used to distinguish the different Paramyxoviridae; however, they probably relate mainly to which form predominates in negatively stained preparations.
The Paramyxoviridae Genomes and their Encoded Proteins
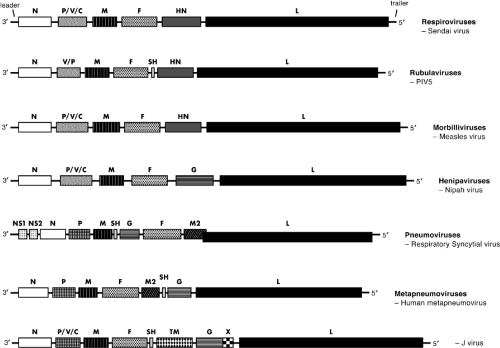
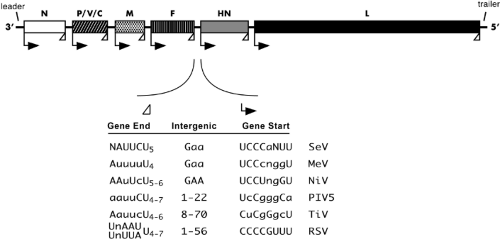
The complete genome sequence for all known members of the Paramyxoviridae has been obtained (available at http://www-ncbi-nlm-nih-gov.easyaccess1.lib.cuhk.edu.hk). The 15,000 to 19,000 nucleotide genomic RNA contain a 3′ extracistronic region of approximately 50 nucleotides known as the leader and a 5′ extracistronic region of 50 to 161 nucleotides known as the trailer (or [−] leader). These control regions are essential for transcription and replication, and flank the six genes (seven for certain rubulaviruses and eight to ten for pneumoviruses). (Note: By the convention used for paramyxoviruses, the term gene refers to the genome sequence encoding a single mRNA, even if that mRNA contains more than one ORF and encodes more than one protein). The coding capacity of the genome of Paramyxovirinae is extended by the use of overlapping ORFs in the P gene. The gene order of a representative member of each subfamily is shown in Figure 33.4. At the beginning and end of each gene are conserved transcriptional control sequences that are copied into mRNA. Between the gene boundaries are intergenic regions (Fig. 33.5). These are precisely three nucleotides long for the respiroviruses and morbilliviruses but are quite variable in length for the rubulaviruses (1–47 nucleotides) and pneumoviruses (1–56 nucleotides) (see Fig. 33.5).
The Nucleocapsid Protein
The nucleocapsid (N) protein is present as the first transcribed gene in the viral genome for all paramyxoviruses except the pneumoviruses and ranges in size from 489 to 553 amino acids (molecular weight ∼53–57 kDa). N is an RNA-binding protein that coats full-length viral negative sense genomic and positive sense antigenomic RNAs to form the helical nucleocapsid template, which is the only biologically active form of these viral RNAs. Electron microscopy and three-dimensional image reconstruction for Sendai virus nucleocapsids reveals that N binds approximately six consecutive nucleotides and 13 N subunits constitute each turn of the nucleocapsid helix.101 In general, these parameters apply to other paramyxovirus nucleocapsids as well, although there can be slight differences in the number of N subunits per helix turn and in the pitch of the helix.15 The binding of N to RNA to form a helical structure is thought to serve several functions, including protection from nuclease digestion, minimizing the annealing of mRNA to complementary genomic RNA, alignment of distal RNA segments to create a functional 3′-end promoter, and most likely providing interaction sites for assembly of progeny nucleocapsids into budding virions.
Expression of paramyxovirus N proteins in the absence of other viral components results in the formation of nucleocapsid-like structures, suggesting that N has inherent self-assembly properties and that N–N interactions drive nucleocapsid assembly.116,277,287 Biochemical and mutational studies have shown that the N protein can be generally divided into two main structural regions: Ncore, an N-terminal domain representing approximately three-fourths of the protein and is conserved in sequence among related viruses, and Ntail, a C-terminal nonconserved acidic domain. Approximately 400 residues of the Sendai virus Ncore are essential for self-assembly, RNA binding, and activity in RNA replication.76
For RSV, the N-terminal 92 residues are sufficient for assembly with RNA.271 A central region of Ncore that is highly conserved for all members of the Paramyxovirinae (residues 258–369 for Sendai virus) contains an F-X4-Y-X3-φ−S-φ-A-M motif (where X is any residue and φ is an aromatic amino acid). This region is essential for self-assembly of N with RNA276 and may be involved in N–N or N–RNA interactions.
For RSV, the N-terminal 92 residues are sufficient for assembly with RNA.271 A central region of Ncore that is highly conserved for all members of the Paramyxovirinae (residues 258–369 for Sendai virus) contains an F-X4-Y-X3-φ−S-φ-A-M motif (where X is any residue and φ is an aromatic amino acid). This region is essential for self-assembly of N with RNA276 and may be involved in N–N or N–RNA interactions.
The C-terminal Ntail region is less well conserved among related paramyxoviruses. Treatment of purified nucleocapsids with trypsin removes a portion of the C-terminal Ntail to yield a more rigid structure,155 suggesting that this domain may confer flexibility in the coiling of the native nucleocapsid. In contrast to the essential role of the N-terminal Ncore in all N functions, the C-terminal Ntail (124 residues for Sendai virus N) is dispensable for binding RNA and for assembly of newly synthesized N-RNA complexes during replication of defective interfering particle RNAs.76 However, nucleocapsids that are assembled with N that lacks this C-terminal Ntail are not functional as templates for the viral RNAP.76 Structural studies have shown that the C-terminal Ntail is intrinsically disordered,26 consistent with a proposed role for this domain in multiple protein–protein interactions. One of these essential interactions with P protein is thought to tether the L-P polymerase complex to the nucleocapsid template.37 For example, the measles virus C-terminal Ntail has been shown to interact with a C-terminal domain of the P protein, undergoing an induced folding in some parts of this N segment.26,186 In the case of measles virus, Ntail has also been shown to bind to the cellular chaperone protein Hsp72455—an interaction that could influence nucleocapsid morphologies and the synthesis of viral RNAs.455
The paramyxovirus nucleocapsid protein is an unusual RNA-binding protein, as it has an overall acidic charge (net charge of –7 to –12, with exception of mumps virus [+2]) and does not contain conventional RNA-binding motifs that are typically found on cellular RNA-binding proteins. The interactions of N with RNA are remarkable stable, and nucleocapsid-associated RNA is protected from nucleases even at very high salt concentrations, or when the hypersensitive C-terminal Ntail is removed by protease digestion.155 N binding to RNA is thought to be independent of nucleotide sequence and, through interactions with the phosphodiester backbone,181 a mechanism that would leave the nucleoside bases accessible to the viral RNAP during RNA synthesis. The Sendai virus nucleocapsid-associated RNA shows hyperreactivity to chemical treatment at cytidine residues predicted to be at positions one and six of a hexamer of nucleotides.181 Together with the finding that the Sendai virus N protein binds six nucleotides,101 these results have led to the proposal that the accessibility of the viral RNAP to bases within the nucleocapsid-associated genomic RNA may be controlled by their position within a hexamer of N-bound nucleotides.181,203
N protein exists in at least two forms in infected cells: one stably associated with RNA in a nucleocapsid structure and a second unassembled soluble form termed N0. This latter form of N has been found to be associated with P in several viruses, including Sendai virus,169 PIV5 (formerly known as simian virus 5 [SV5]),331 measles virus,379 and RSV.121 N0 is thought to be the functional form of N that encapsidates the nascent RNA strand during genome and antigenome replication.78,169 N-terminal regions of Ncore are important for formation of the N0-P complex,166 and these domains are distinct from those involved in binding of P to N in the assembled nucleocapsid.
Table 33.2 Examples of Identified P Gene Open Reading Frames | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
The P Gene and Its Encoded Proteins
The Paramyxovirinae P gene is a remarkable example of exploiting the coding capacity within a viral gene. The Sendai virus P/V/C gene is the most diverse of the paramyxovirus P genes,
directing the expression of at least seven polypeptides, including the P, V, W, C′, C, Y1, and Y2 proteins. Whereas other paramyxoviruses express fewer proteins from the P/V/C gene than Sendai virus, the P gene always produces more than one polypeptide species (Table 33.2). Expression of P/V/C proteins can involve two main mechanisms, with members of a paramyxovirus genus having a characteristic combination of these expression strategies. The first expression mechanism, which produces the P, V, and W/I/D proteins, has been termed RNA editing or pseudotemplated addition of nucleotides.307,408,419 This mechanism involves the production of mRNAs whose ORFs are altered by insertion of G residues at a specific position in the mRNA. As described later, the second expression mechanism involves ribosome initiation at alternative translation codons and produces the family of C proteins.
directing the expression of at least seven polypeptides, including the P, V, W, C′, C, Y1, and Y2 proteins. Whereas other paramyxoviruses express fewer proteins from the P/V/C gene than Sendai virus, the P gene always produces more than one polypeptide species (Table 33.2). Expression of P/V/C proteins can involve two main mechanisms, with members of a paramyxovirus genus having a characteristic combination of these expression strategies. The first expression mechanism, which produces the P, V, and W/I/D proteins, has been termed RNA editing or pseudotemplated addition of nucleotides.307,408,419 This mechanism involves the production of mRNAs whose ORFs are altered by insertion of G residues at a specific position in the mRNA. As described later, the second expression mechanism involves ribosome initiation at alternative translation codons and produces the family of C proteins.
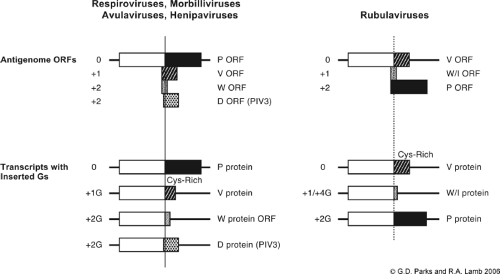
The P and V proteins, as well as virus-specific proteins variously referred to as W, I, and D, are produced as a co–N-terminal nested set of proteins. These polypeptides are translation products from distinct mRNAs that differ only by inserted G nucleotides that shift the translational reading frame at the site of insertion. As shown in Figure 33.6, the P gene of the respiro-, morbilli-, and henipaviruses codes for a long N-terminal ORF shared by all three proteins and three shorter ORFs starting at approximately base 400 in the mRNA. During transcription of the nucleocapsid template, the viral RNAP is directed to make an accurate copy of the P gene template or to insert one or two G residues at a precise site in the nascent mRNA. The result is that the accurate transcription product encodes the full-length P ORF, whereas the mRNAs with insertions of +1G and +2G have a shift in the translational ORF such that the 5′ end P ORF is fused at the site of insertion in the mRNA coding sequence to a more 3′ ORF encoding V (+1G) or W (+2G). Thus, the P, V, and W/I/D proteins that are produced as a result of RNA editing share a common N-terminal region but differ in their C-terminal regions starting at the site of G insertion.
All viruses of the Paramyxovirinae (with the exception of human parainfluenza virus type 1 [HPIV1]) 250 encode a characteristic editing site in the P gene, and the number of inserted G residues, as well as the frequency of inserting G nucleotides, is determined by sequences surrounding and within the editing site. For example, Sendai virus encodes the P protein as the translation product from the unedited mRNA (+0 G; see Fig. 33.6). The
V protein is produced from a transcript containing a single G residue at the insertion site (+1 G), which fuses the common N-terminal ORF to the V-specific ORF. The Sendai virus transcript with two inserted G nucleotides codes for the W protein (+2 G). As shown in Figure 33.6, rubulaviruses differ from other paramyxoviruses in that V protein is produced by translation of the unedited mRNA (+0 G), and P is produced by translation of an mRNA containing a two-G insertion (+2 G).
V protein is produced from a transcript containing a single G residue at the insertion site (+1 G), which fuses the common N-terminal ORF to the V-specific ORF. The Sendai virus transcript with two inserted G nucleotides codes for the W protein (+2 G). As shown in Figure 33.6, rubulaviruses differ from other paramyxoviruses in that V protein is produced by translation of the unedited mRNA (+0 G), and P is produced by translation of an mRNA containing a two-G insertion (+2 G).
Insertion of G residues into P gene mRNA transcripts is a co-transcriptional event catalyzed by the vRNAP419 and is usually limited to insertions of between 0 and 2 nucleotides, depending on the virus. Human and bovine parainfluenza virus type 3 (HPIV3 and BPIV3) are exceptions to this general rule, and mRNAs with one to six inserted G residues are almost equally abundant.120
The Phosphoprotein
The P protein is the only P/V/C gene product that is essential for viral RNA synthesis.78 P is generally 400 to 600 amino acids long and is heavily phosphorylated at serine and threonine residues, predominantly within the N-terminal region. P protein contains regions of high intrinsic disorder,25 consistent with the requirement for interacting with multiple partners during the viral growth cycle. P is an essential component of both the vRNAP enzyme141 and the N0 nascent chain assembly complex that functions to encapsidate RNA during replication.169 Extensive mutational analyses have identified distinct modular C- and N-terminal domains within the P protein that play essential roles as a polymerase cofactor and in nascent chain assembly, respectively.
The C-terminal polymerase cofactor module is relatively well conserved in predicted secondary structure for all viruses of the Paramyxovirinae, and all P proteins carry this essential module as the C-terminal segment of fusion protein with the shared P/V domain; this module is never naturally expressed by itself. The P protein C-terminal region contains domains for P-P multimerization, for interactions with L protein, and for binding to the N:RNA template. P protein functions as a multimer, and structural analysis suggests that the Sendai virus P protein is a tetramer.402 For Sendai virus, the P-carboxy region is sufficient for catalyzing viral RNA synthesis, as this protein fragment by itself (residues 325–568) can substitute for intact P protein in all aspects of mRNA transcription.79 Although the L protein is thought to contain all vRNAP catalytic activities, L binds to the nucleocapsid template via the P protein.169 This P–L interaction requires a domain in P that maps to the C-terminal end of the coiled-coil P-P multimerization region.32,80 At the end of the C-terminal domain, P also contains a region that binds to the N:RNA template,32,80 providing the bridge to link L with the N:RNA template. In the case of Sendai virus, structural data indicate that this C-terminal region of P binds through weak hydrophobic interactions to the C-terminal tail of N, inducing folding of the intrinsically disordered Ntail.232 The ability of P and Ntail to form transient weak interactions between intrinsically disordered domains may be important for the dynamic functions of P during movement of the viral RNAP across the N:RNA template or in the flexibility of the nucleocapsid template.17,25,200
In contrast to viral transcription, genome replication requires an N-terminal region of P (defined by deletion of Sendai virus residues 33–41). A short segment of the P protein N-terminal domain is thought to facilitate interactions with unassembled N0 to prevent N aggregation and to ensure specificity in assembly.78,169 The rest of the N-terminal domain of P protein is apparently dispensable for genome RNA synthesis and assembly, as a P protein in which residues 78 through 324 have been deleted is still active for minigenome replication in transfected cells.74
The V Protein
The V protein is an approximately 25- to 30-kDa polypeptide that shares an N-terminal domain with the P protein but has a distinct C-terminal domain as a result of RNA editing.48,303,408,419 The C-terminal V-specific domain is highly conserved among related paramyxoviruses (Fig. 33.7), with invariantly
spaced histidine and cysteine residues forming a novel domain that binds two zinc molecules per V protein.118,225,231,304 Despite the high level of intracellular synthesis, paramyxovirus particles typically contain little V protein,304 and the degree of incorporation of V into virions varies among paramyxoviruses.75,431
spaced histidine and cysteine residues forming a novel domain that binds two zinc molecules per V protein.118,225,231,304 Despite the high level of intracellular synthesis, paramyxovirus particles typically contain little V protein,304 and the degree of incorporation of V into virions varies among paramyxoviruses.75,431
V protein plays several important roles in the virus replication cycle, as evidenced by recombinant viruses that have been engineered to disrupt expression of the V protein Cys-rich domain.12,153,191,412 In many cases, these mutant viruses display an elevated RNA synthesis phenotype, although they generally grow well in many tissue culture cell lines.82,192 However, in many cases, these viruses are severely attenuated for growth in vivo or are cleared more rapidly than wild-type viruses from lungs of infected animals.94,192,418 These results suggest that V is an accessory protein that plays a role in viral pathogenesis, perhaps involving a counteracting of host cell antiviral responses that occur early after infection and that can lead to enhanced clearance of virus.
V protein has also been shown to inhibit viral RNA synthesis in transfection experiments involving model RNA genomes.78,168,229 Recombinant viruses that are engineered with V protein mutations often show increased viral RNA synthesis.81,191,360,412,430 This has led to the proposal that V protein serves as a negative regulator of viral RNA synthesis. V protein shares the amino-terminal domain of P protein that can interact with N0 to form the assembly competent P-N0. Thus, the mechanism of V inhibition may involve interactions with N that result in a form of a V-N0 complex that is not competent to function during the RNA encapsidation step of replication. This V–N0 interaction has been detected in the case of PIV5, Sendai virus, and measles virus,168,331,412 and a model whereby V and P compete for soluble N0 has been proposed.78,168 The V protein is also capable of binding RNA,226 and it has been proposed that this function is involved in inhibiting RNA synthesis for the Sendai virus V protein.300
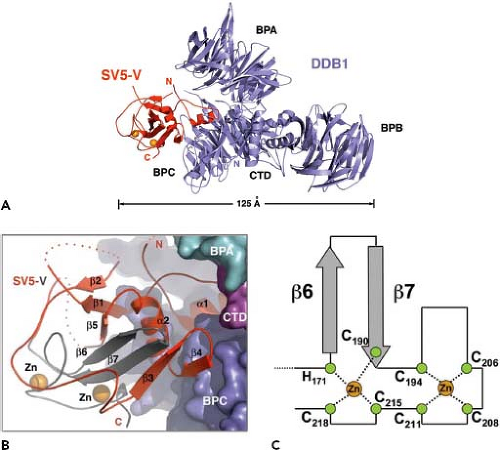
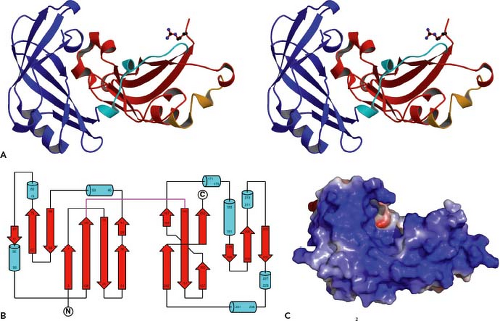
In addition to binding viral components, V protein also has been detected in interactions with cellular proteins. For several paramyxoviruses, V protein interacts in the cytoplasm with the cellular damage-specific DNA-binding protein 1 (DDB1).4,227 In the case of the PIV5 V protein, interaction with DDB1 is important for the function of blocking signaling through the type I interferon pathway (see later discussion). Interaction of V with DDB1 and the ability of V to inhibit host cell antiviral responses depends on the C-terminal Cys-rich domain but can also be disrupted by alterations to the common N-terminal P/V region.4,227 The structural analysis of PIV5 V protein complexed with DDB1 shows that V protein has a bi-partite structure,225 with a core domain built around a central seven-stranded β sheet, which is in turn sandwiched between one alpha helix and two long loops (see Fig. 33.8). The unique C-terminal domain forms the middle two β sheets and part of the central core, and this structure is anchored through the Cys-rich zinc-binding region. Thus, despite sharing a 164 amino acid N-terminal domain, the PIV5 P and V proteins can adopt very different structures owing to the unique properties of the C-terminal Cys-rich region. V protein from several paramyxoviruses has been shown to interact through the Cys-rich domain with the cellular protein MDA-5, an IFN-inducible host cell DExD/H box helicase that is involved in signaling to initiate host cell antiviral responses.
The W/D/I Proteins
The W and D ORFs of respiro-, morbilli- and henipaviruses are expressed from mRNAs with two inserted G residues (see Fig. 33.6). For most of these viruses, the insertion of two G residues into the mRNA is relatively rare, and the ORF is closed by a stop codon shortly after the editing site, resulting in the ORF for the W protein. Thus, the W protein is essentially a truncated P protein, containing the N-terminal N0 assembly module of the P protein alone. W protein is abundantly expressed in Sendai virus–infected cells73 and has been found to interact with unassembled N0, suggesting an inhibitory role in viral RNA synthesis.168 In the case of BPIV3 and HPIV3, the +2 ORF extends for 131 residues from the editing site, and the protein that links the amino-terminal P domain to this ORF is called D protein.120 The Rubulavirus I protein is generated when the upstream N-terminal P region is fused to a downstream ORF by the insertion of either one or four G residues during RNA editing.303,408 The role that the W/D/I proteins play in the viral growth cycle has not been established.
The C Proteins
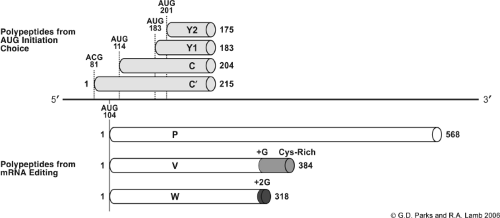
In addition to RNA editing, some paramyxoviruses use a second mechanism to express P gene polypeptides that involves the use of alternative translation initiation codons to yield the C proteins (Fig. 33.9). The Sendai virus C′, C, Y1, and Y2 proteins comprise a nested set of carboxy–co-terminal polypeptides that range in size from 175 to 215 residues. These proteins are expressed independently from a P/V mRNA through the use of alternative start codons (Fig. 33.9), with the C protein ORF being in the +1 reading frame relative to the P ORF. The C′ and C proteins are translated by a leaky scanning mechanism, being initiated at an unconventional ACG triplet at base 81 and AUG at base 114, respectively.77 By contrast, translation of the Y1 and Y2 proteins occurs through a scanning-independent ribosome shunting mechanism that is directed by a 5′ noncoding RNA segment, resulting in ribosomes initiating at AUG codon bases 183 and 201, respectively. Translation of each of the C′, C, Y1, and Y2 ORFs is initiated at a different site, although translation is terminated at the same downstream stop codon; thus, these proteins share a common C-terminal region. The C protein is abundantly expressed in infected cells at levels higher than C′, Y1, and Y2; however, virions contain only very low levels of these polypeptides.217 Morbilliviruses express one C protein,13 as do the henipaviruses,427 whereas the respiroviruses such as Sendai virus and HPIV1 express all four C′, C, Y1, and Y2 polypeptides.128 Rubula- and avulaviruses do not express C proteins (see Table 33.2).
C proteins are small basic polypeptides that play multiple functions in the viral growth cycle, being involved in the control of viral RNA synthesis, counteracting host cell antiviral pathways, and facilitating release of virus from infected cells. Although nonessential for infectivity, Sendai virus mutants engineered to express only a subset of C proteins or lacking expression of all four proteins show defects in virus growth.209 The C proteins have been shown to inhibit mRNA transcription and suppress RNA replication in a promoter-specific manner.193,238,399 Consistent with this, viral mutants that are engineered to lack C protein expression show elevated synthesis of viral mRNA and protein.147 The inhibition of RNA synthesis by C proteins correlates with the ability to bind to the
L subunit of the viral polymerase,170 and in the case of Sendai virus, naturally occurring variant C proteins can have differential effects on inhibition of virus RNA synthesis.10
L subunit of the viral polymerase,170 and in the case of Sendai virus, naturally occurring variant C proteins can have differential effects on inhibition of virus RNA synthesis.10
The role of paramyxovirus C proteins in pathogenesis and in counteracting host cell IFN responses is best understood in the cases of Sendai virus and measles virus. For Sendai virus, the C′, C, Y1, and Y2 proteins can antagonize IFN signaling when assayed in stably transfected HeLa cells193; however, there may be more subtle differences in the functions of each polypeptide in the context of viral mutants.124 In the case of measles virus, recombinant viruses defective for C protein expression grow well in certain culture cells but are defective for growth in peripheral blood mononuclear cells104 and are less virulent in vivo.310 Changes in pathogenesis of C-mutant viruses may be related to the ability of the C proteins to inhibit type I IFN responses.368 This proposal is further supported by a naturally occurring mutation in the Sendai virus C protein (phenylalanine 170 to serine) that eliminates the ability of C protein to block IFN signaling,124 and a mutant Sendai virus harboring this altered C protein is attenuated for growth in mice. The mechanism by which C proteins attenuate IFN signaling has not been elicited but may involve binding of C to STAT1124 or altering STAT1 phosphorylation patterns.205
An additional role for C proteins in virus release became evident with the analysis of mutant Sendai virus that cannot express any of the four C proteins.209 Whereas viral RNA and protein synthesis was high for this mutant virus, production of infectious virions was low, and heterogeneous noninfectious particles were produced.147 C protein expression enhances release of virus-like particles (VLPs), possibly through interactions with AIP1/Alix—a cellular protein involved in apoptosis and endosomal trafficking.347
The Large Protein
The large (L) protein is an essential subunit of the paramyxovirus RNAP. Consistent with a catalytic role in viral RNA synthesis, the L protein is invariably encoded as the most promoter-distal gene in the paramyxovirus genome (see Fig. 33.4). L protein is generally found in only very low amounts in infected cells or associated with nucleocapsids and virions.214 A paramyxovirus particle typically contains only about 50 copies of L,214 where it is found on the nucleocapsid in clusters that co-localize with P protein.327 L is thought to possess all of the enzymatic activities needed for synthesis of functional viral mRNA, including nucleotide polymerization as well as 5′-end capping and methylation and 3′-end polyadenylation of mRNAs.137,158,289 Polyadenylation of viral mRNAs occurs co-transcriptionally, where L is thought to add poly A tails to nascent viral mRNAs through a mechanism that involves stuttering at a stretch of template U residues at the end of each viral gene (see Fig. 33.5). L protein is also responsible for the replication of viral genomic and antigenomic RNA; however, this form of RNA synthesis differs from mRNA transcription by having a strict requirement for soluble N0 to allow encapsidation of the nascent genomic RNA.139,169
The paramyxovirus L protein is generally approximately 2,200 amino acids in length (∼250 kDa). Although the N- and C-terminal regions of the L proteins are diverse, sequence comparisons among L proteins have identified six highly conserved domains (I–VI) near the middle of the polypeptide. It was originally proposed that these domains may be individually responsible for each of the multiple L functions.319 Domain II is proposed to be an RNA-binding domain owing to the high net positive charge. In domain III, mutational analyses are consistent with the proposal of a conserved GDNQ motif as the active site for nucleotide polymerization.237 Based on sequence homologies, domain VI of the rhabdovirus L protein has been implicated in playing a major role in 5′ cap formation, perhaps as a methyltransferase domain.114,319 The precise roles of the remaining domains I, IV, and V in individual steps of RNAP activity are not clear; however, for Sendai virus, mutations in some of these domains result in L proteins that can transcribe viral mRNA but are defective in RNA replication.56,113 In the case of the L proteins of Sendai virus and rinderpest virus, sequence alignment has identified nonconserved hinge regions that can be modified by insertions of green fluorescent protein (GFP), and remarkably, viable recombinant viruses encoding these L-GFP hybrid proteins have been isolated and used to identify sites of L localization during infection.35,92
L protein activity in RNA synthesis highly depends on protein–protein interactions, involving self-assembly as well as binding to other viral and cellular proteins. Biochemical evidence and genetic complementation studies indicate that the Sendai and measles virus L proteins function as homo-multimers that interact through an N-terminal self-assembly domain.54,376 L also binds to the viral P protein—an interaction that is essential for formation of the active enzyme complex and can lead to enhanced stability of L.141,169 L–P interaction domains generally map to an N-terminal domain of L that is distinct from the L-L assembly domain.165 Within the L-P complex, P protein serves as the bridge to link the L polymerase to the nucleocapsid template.80,169 In addition to L–L and L–P interactions, L protein also interacts with host cell proteins.269,375 In the case of measles virus and Sendai virus, L interactions with tubulin are thought to promote L activity.269 Other cellular proteins have also been shown to promote viral RNA synthesis (e.g., β-catenin for HPIV3),22 although the precise role that these proteins play in viral RNA synthesis has not been determined.
Whereas interactions of L with P are generally thought to promote activity, L protein can also interact with other viral components that inhibit the vRNAP. For both rinderpest virus and Sendai virus, L protein has been found to bind the viral C proteins.170,390 The Sendai virus L–C interactions are through a domain of L that maps to the first 895 residues (domains I–III),170 and this binding correlates with inhibition of RNA synthesis.138 Other proteins encoded in the viral P/V/C gene (C′, Y1, and Y2) also interact with L and inhibit defective interfering RNA synthesis in vitro138 and in vivo.193
The Matrix Protein
The paramyxovirus matrix (M) protein is the most abundant protein in the virion. The M proteins contain 341 to 375 residues (Mr ∼38,500–41,500), are quite basic proteins (net charge at neutral pH of +14 to +17), and are somewhat hydrophobic, although there are no domains of sufficient length to span a lipid bilayer. In electron micrographs of virions, an electron-dense layer is observed underlying the viral lipid bilayer, and this is thought to represent the location of this protein. Fractionation studies of virions indicate that the M protein is peripherally associated with membranes and is not an intrinsic membrane protein. Reconstitution studies of purified
M protein and fractionation studies of infected cells indicate that the M protein can associate with membranes.105,212,280
M protein and fractionation studies of infected cells indicate that the M protein can associate with membranes.105,212,280
As a purified protein, the Sendai virus M protein can self-associate and form two-dimensional paracrystalline assays (sheets and tubes) in low salt conditions.7,157 There is a paracrystalline array of identical periodicity at the inner surface of the plasma membrane of infected cells when examined by freeze-fracture techniques in the electron microscopy.7 In addition, the M protein is associated with nucleocapsids.386 As of January 2011 the only atomic structure of M to be obtained is that of RSV M.264 It shows that the protein has extensive β-sheets and a continuously charged region covering approximately 600 Å, which probably interacts with a negatively charged surface on the RNP (Fig. 33.10). Genetically engineered recombinant measles virus and PIV5 that lack glycoprotein cytoplasmic tails show a subcellular redistribution of the matrix protein,46,357 which implies that there is an interaction of the F and HN cytoplasmic tails with the M protein. Thus, the M protein is considered to be the central organizer of viral morphogenesis interacting with the cytoplasmic tails of the integral membrane proteins, the lipid bilayer, and the nucleocapsids. The self-association of M and its contact with the nucleocapsid may be the driving force in forming a budding virus particle.311 The relative abundance of basic residues in the M protein may reflect their importance in ionic interactions with the acidic N proteins.
For several enveloped viruses, it has been shown that budding occurs by using components of the endosomal sorting complexes required for transport (ESCRTs)—proteins involved in multivesiculate body formation. Protein–protein interaction domains called late domains have been identified in the matrix proteins of several viruses; for the paramyxoviruses, a late domain has been identified in PIV5 M protein.358 This topic is discussed further in the Assembly of the Envelope section.
Consistent with its central role in virus budding, M is often inactivated in persistent paramyxovirus infections where budding fails to occur. For example, in subacute sclerosing panencephalitis (SSPE)—a rare, progressive, and invariably fatal persistent measles virus infection of the brain—the M protein is either absent for various reasons47 or, when present, is not associated with budding structures in vivo and is unable to bind to viral nucleocapsids in vitro.162 Although a genetically engineered recombinant measles virus that lacks a matrix
protein has been obtained,45 it produces approximately 4 logs lower titer of released infectious particles than wild-type virus and remains mostly cell associated. Therefore, it is reasonable to conclude that the M protein does play a very important function in virus assembly. Moreover, in model systems of persistent Sendai virus infection in culture, the normally lytic infection is converted to a persistent one using defective interfering particles. This change correlates mainly with M protein instability and an absence of budding structures.341
protein has been obtained,45 it produces approximately 4 logs lower titer of released infectious particles than wild-type virus and remains mostly cell associated. Therefore, it is reasonable to conclude that the M protein does play a very important function in virus assembly. Moreover, in model systems of persistent Sendai virus infection in culture, the normally lytic infection is converted to a persistent one using defective interfering particles. This change correlates mainly with M protein instability and an absence of budding structures.341
The M protein of several paramyxoviruses is phosphorylated. For Sendai virus, a large proportion of the M protein is phosphorylated, whereas the M protein found in virions is not phosphorylated.212 However, a Sendai virus could be rescued from an infectious complementary DNA (cDNA) in which the single phosphorylation site in Sendai virus M protein had been eliminated.348 This M protein phosphorylation-minus mutant did not show an altered phenotype from wild-type virus in either cultured cells or mice.
Envelope Glycoproteins
All Paramyxoviridae possess two integral membrane proteins, and some rubulaviruses and all pneumoviruses encode a third integral membrane protein (Fig. 33.11). One glycoprotein (HN, H, or G) is involved in cell attachment and the other glycoprotein (F) in mediating pH-independent fusion of the viral envelope with the plasma membrane of the host cell. The Rubulavirus and Pneumovirus third integral membrane protein is referred to as SH; for PIV5, this 44 amino acid integral membrane protein is thought to block virus-induced apoptosis. The assignment of specific biological activities of F and HN was originally made on the basis of purification and reconstitution studies, mainly for the Sendai virus and PIV5 proteins.353,354 The attachment proteins (HN, H, or G) are all type II integral membrane proteins, and bioinformatics and structural predictions indicate that the proteins will all exhibit a related propeller-like fold despite having different receptors and the presence or absence of NA activity.
For the respiroviruses and rubulaviruses, the attachment glycoprotein binds to cellular sialic acid–containing receptors, and these can be glycoproteins or glycolipids. The binding is probably of fairly low affinity but of sufficiently high avidity that these viruses agglutinate erythrocytes (hemagglutination). The attachment proteins of respiroviruses and rubulaviruses also have NA activity (receptor-destroying activity), and the proteins have been designated hemagglutinin-neuraminidase (HN). However, a possible role of a specific protein–protein involvement in infection of host cells has not been ruled out.
The restricted host range of measles virus for primate cells and the lack of NA or esterase activity make it unlikely that sialic acid is the primary receptor for measles virus. Nonetheless, the Morbillivirus attachment protein (H) can cause agglutination of primate erythrocytes, most likely owing to receptor binding: the designation of measles and CDV glycoprotein as H is thus a misnomer. In 1993, human CD46 was identified as a cellular receptor for Edmonston and Halle strains of measles virus.91,282 Edmonston and Vero cell-isolated strains of measles virus are capable of infecting any CD46+ primate cell. However, viruses isolated from B- and T-cell lines do not grow in CD46+ cells. A second receptor was identified—human CD150 (SLAM), a membrane glycoprotein involved in lymphocyte activation.404,442 It is now thought that CD150 is the principle receptor for unadapted isolates of lymphotropic measles virus.291
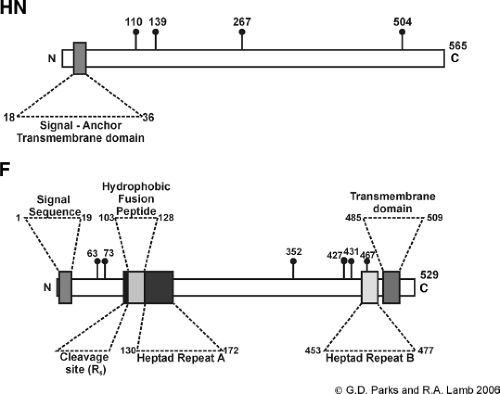 Figure 33.11. Schematic diagram showing the orientation and domains of paramyxovirus integral membrane proteins. A: Hemagglutinin-neuraminidase (HN) attachment protein (based on the predicted sequence of the parainfluenza virus type 5 [PIV5] HN gene.159 The signal anchor transmembrane domain and the sites used for addition of N-linked carbohydrate (lollipops)285 are indicated. B: Fusion protein (based on the predicted sequence of the PIV5 F gene). The position of the signal sequence, the transmembrane domain, the cleavage site, the hydrophobic fusion peptide, and the heptad repeats A and B are indicated. The sites used for addition of N-linked carbohydrate (lollipops)8 are indicated. R5 indicates the five arginine residues site for cleavage activation. |
Very recently, a third receptor for measles virus has been identified, known as poliovirus receptor-like (PVRL4; Nectin 4) or adherens junction protein nectin 4. It is proposed that this new receptor is the epithelial receptor for measles virus that
is used to transfer virus from the basolateral surface of epithelial cells to the lumenal side of the airway.270,288
is used to transfer virus from the basolateral surface of epithelial cells to the lumenal side of the airway.270,288
The receptor for Hendra virus and Nipah virus G glycoprotein has been shown to be ephrin-B2 or ephrin-B3. In one approach, direct binding of Nipah G to receptor was obtained and the identity of the receptor determined by protein sequencing and bioinformatics.284 In another approach, microarray analysis was used to identify mRNAs that were expressed in henipavirus-susceptible cells and not in cells refractory to henipavirus infection.20 Ephrin-B2 and -B3 are members of a family of cell surface glycoprotein ligands that bind to ephrin (Eph) receptors—a large family of tyrosine kinases. The identification of ephrin-B2/B3 as the cellular receptor for both Hendra virus and Nipah viruses and the widespread occurrence of ephrin-B2/3 in vertebrates, particularly in arterial endothelial cells and in neurons, provides an explanation for the wide host range of henipaviruses and their systemic infection.98
The Pneumovirus RSV does not cause detectable hemagglutination, and the cellular receptor for RSV is not completely understood but involves interactions with heparan sulfate—a glycosaminoglycan that is part of the extracellular matrix. Interestingly, the G protein of RSV and human metapneumovirus (HMPV) can be deleted from the viral genome,189 and Sendai virus–like particles devoid of HN can infect cells via the asialoglycoprotein receptor.224 Both of these cases suggest that some paramyxovirus F proteins may have a binding activity. After attachment of a Paramyxoviridae particle to the host cell receptor, the viral envelope fuses with the host cell plasma membrane, and the major viral protein involved in this process is the F glycoprotein.
Paramyxovirus Attachment Protein
The Respirovirus and Rubulavirus surface glycoprotein HN is a multifunctional protein and the major antigenic determinant of the paramyxoviruses. HN has three activities: (a) receptor binding to sialic acid; (b) cleavage of sialic acid from complex carbohydrate chains (NA activity); and (c) fusion promotion—that is, co-expression of HN and F is required for cell–cell fusion (see later discussion). By analogy to the role of influenza virus NA, it seems likely that the role of this NA activity is to prevent self-aggregation of viral particles during budding at the plasma membrane. These dual activities of HN can be modulated by pH.258 Whereas the pH of the extracellular environment is optimal for hemagglutination, paramyxovirus NAs have an acidic pH optima (pH 4.8–5.5), suggesting that NA acts in the acidic trans-Golgi network to remove sialic acid from the HN carbohydrate chains and from the F protein carbohydrate chains.
The HN polypeptide chain ranges from 565 to 582 residues. For some strains of NDV, HN is synthesized as a biologically inactive precursor (HNo), and 44 residues from the C-terminus are removed to activate the molecule.278,279 HN is a type II integral membrane protein that spans the membrane once and contains an N-terminal cytoplasmic tail, a single N-terminal transmembrane (TM) domain, a membrane-proximal stalk domain, and a large C-terminal globular head domain.159 The globular head domain contains the receptor-binding and enzymatic activity.301,353,410 HN is glycosylated and contains from four to six potential sites for the addition of N-linked carbohydrate chains. For PIV5 and NDV HN, it is known that four sites are used.253,285 HN is noncovalently associated to form a dimer of dimers, based on biochemical, cross-linking, electron microscopy, and structural studies that, depending on the paramyxovirus, can be composed of two disulfide-linked dimers.72,148,218,252,285,286,409,450,451 The covalent linkage occurs through a cysteine residue at the C-terminal end of the stalk domain, just prior to the beginning of the head domain. The stalk domain appears to play an essential role in the formation of the tetramer,451 and head domains when expressed without the stalk are often monomeric.72,218,451
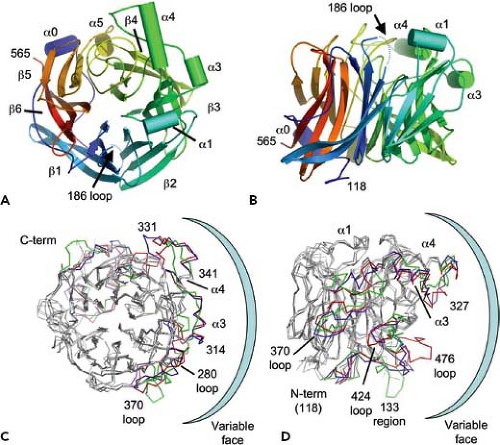
The structure of the enzymatically active head domain of HN is similar to other NAs or sialidases, such as influenza NA,103 with the globular head composed of identical subunits. Each NA domain exhibits the six-blade propeller fold typical of other NA/sialidase structures from viral, protozoan, or bacterial origin.41,216,405 Atomic structures of soluble head domains of NDV, HPIV3, PIV5 (liganded and bound to a receptor/substrate sialyllactose), measles virus H (unliganded and bound to its receptor CD150/SLAM), and Hendra and Nipah virus G (unliganded and bound to its receptor ephrin-B2 or ephrin-B3) have been obtained,30,31,60,72,148,149,218,441,451,453 and it shows the typical sialidase fold consisting of six antiparallel β-strands organized as a super barrel with a centrally located active site (Fig. 33.12). The seven highly conserved active site residues found in NA and sialidases are found in the paramyxovirus HN structures. However, these key active site residues are mutated in the measles H and Nipah/Hendra G proteins, rendering these proteins enzymatically dead. Superimposition of the NDV, HPIV3, and PIV5 HN monomer structures indicates a high degree of conservation on one face of the molecule, with the other face containing more variability and additional protein loops.451
It has long been debated whether the hemagglutinin and NA activities of HN involve one or two separate sialic acid binding sites.72,329,453 The disparate theories of one site with dual function or of two distinct sites that are intimately related are both consistent with the observation that sialic acid–derived NA inhibitors interfere with receptor binding.177,275,355 A single site can provide both hemagglutinin and NA activities by binding sialic acid tightly and hydrolyzing the molecules slowly.355 For NDV HN, two sialic acid sites have been observed in the x-ray structures: one is the active site, and a second site is located at the dimer interface.453 Strong biological evidence supports the notion of a second sialic acid binding site in NDV.29,323 Mutagenesis of a key residue involved in the dimer interface sialic acid binding site abolishes sialic acid binding to the second site.29 However, virus containing this key residue mutation is only marginally affected in growth properties.29 For HPIV3, the second sialic acid binding site is blocked by a carbohydrate chain that prevents its function,261 although mutagenesis to ablate the carbohydrate chain allows the HPIV3 second site to bind sialic acid. The growth curve of HPIV3 with or without the carbohydrate chain that shields the second sialic acid binding site is very similar, suggesting no major biological importance.
For PIV5 HN, not only was a second sialic acid binding site not observed, the molecule could not form the second sialic acid binding site between two monomers owing to changes in sequence and conformation.218,451 Thus, the biological importance of the second sialic acid binding sites in NDV HN and the one created in HPIV3 by removal of the carbohydrate addition site are a conundrum.261,322,325
From the structural studies of NDV HN, it was also suggested that the NA domain could form two distinct dimeric
assemblies that were ligand dependent.72 One of the dimers, observed after co-crystallization with ligand, formed an extensive buried interface, whereas the second dimer, crystallized in the absence of ligand and at low pH, formed a much smaller interface. Conformational changes were observed in the active site of the HN protein upon ligand binding that were correlated with changes in the dimer interface, suggesting a possible mechanism for coupling ligand recognition to changes in the oligomeric assembly of the HN protein. However, engineered disulfide bonds block dimer dissociation and do not affect fusion, rendering major HN rearrangements unlikely.234 Structural studies of HPIV3 and PIV5 HN also do not support the notion that there are ligand-dependent conformational changes within the monomeric protein structure.218,451 The dimer of HN that is observed in the HPIV3 and PIV5 structures occurs in the absence of ligand binding, and there is no crystallographic evidence that monomeric ligand binding influences the oligomeric structure of these HN proteins.
assemblies that were ligand dependent.72 One of the dimers, observed after co-crystallization with ligand, formed an extensive buried interface, whereas the second dimer, crystallized in the absence of ligand and at low pH, formed a much smaller interface. Conformational changes were observed in the active site of the HN protein upon ligand binding that were correlated with changes in the dimer interface, suggesting a possible mechanism for coupling ligand recognition to changes in the oligomeric assembly of the HN protein. However, engineered disulfide bonds block dimer dissociation and do not affect fusion, rendering major HN rearrangements unlikely.234 Structural studies of HPIV3 and PIV5 HN also do not support the notion that there are ligand-dependent conformational changes within the monomeric protein structure.218,451 The dimer of HN that is observed in the HPIV3 and PIV5 structures occurs in the absence of ligand binding, and there is no crystallographic evidence that monomeric ligand binding influences the oligomeric structure of these HN proteins.
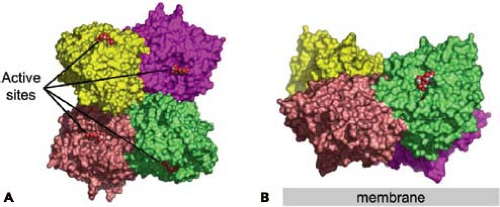
The HN tetrameric arrangement451,453 is unusual, because rather than having fourfold rotational symmetry as might be anticipated, it is arranged with two twofold symmetry axes (Fig. 33.13) that are orientated at approximately 90 degrees
to each other and in the crystal lattice, allowing neighboring dimers and tetramers to associate in infinitely long oligomers. The dimer places the two HN active sites at nearly 90 degrees to each other. The calculated buried surface area for each monomer in the PIV5 HN dimer is 1,818 A2. In contrast to the dimer interaction, the dimer-of-dimers interface is much smaller, involving only 10 residues and burying only 657 A2. The small surface of interaction suggests that the arrangement is not very strong and that the dimers may dissociate.
to each other and in the crystal lattice, allowing neighboring dimers and tetramers to associate in infinitely long oligomers. The dimer places the two HN active sites at nearly 90 degrees to each other. The calculated buried surface area for each monomer in the PIV5 HN dimer is 1,818 A2. In contrast to the dimer interaction, the dimer-of-dimers interface is much smaller, involving only 10 residues and burying only 657 A2. The small surface of interaction suggests that the arrangement is not very strong and that the dimers may dissociate.
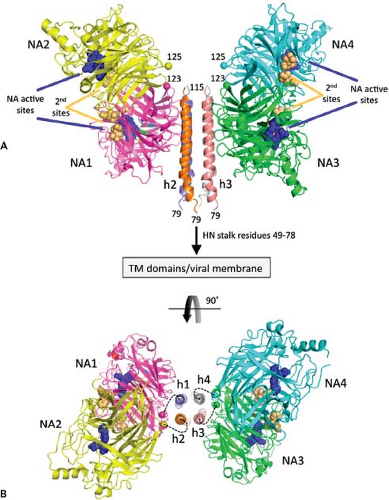
Recently, the atomic structure of the NDV head domain with a tetrameric stalk has been obtained450 (Fig. 33.14). The stalk forms a four-helix bundle, and on either side are dimers of head domains. One head domain of each dimer makes extensive interactions with the stalk. This structure, as compared with the head-only tetramer,451 suggests plasticity in the stalk/head-connecting region.
The structure of the Pneumovirus attachment protein (G) is very different from the attachment protein of the Paramyxovirinae. The RSV G protein has neither hemagglutinating nor NA activity. The nucleotide sequence of the RSV G gene predicts that the protein is of 289 to 299 amino acids (Mr)32,587 and is a type II integral membrane protein with a single N-terminal hydrophobic signal/anchor domain.352,435 The G protein is found in virus-infected cells in both membrane-bound and proteolytically cleaved soluble forms. The distinguishing feature of the RSV G protein is the extent of its carbohydrate modification. On SDS-PAGE, the protein migrates with an apparent Mr of approximately 84,000 to 90,000, and the dramatic increase in molecular weight over that predicted for the polypeptide chain is because 8 to 12 kDa is owing to addition of N-linked carbohydrate (four potential addition sites) and 40 to 50 kDa is owing to the addition of O-linked glycosylation (77 potential acceptor serine or threonine residues; 30% of total residues) (61 and references therein). Quite remarkably, it appears that the RSV G protein is not essential for virus assembly or growth in tissue culture or animals, although it does confer a growth advantage. A virus that had been extensively passaged in cells was found to contain a spontaneous deletion of the G and SH genes,189 yet the virus replicated in Vero cells. In addition, the G gene has been deleted from recombinant virus recovered from
an infectious cDNA clone (see Chapter 38). These findings suggest that RSV has an alternate mechanism for attachment to cells that does not involve G protein, and evidence has been obtained that RSV lacking G protein can bind to heparan sulfate and possibly other molecules.111,140,406 Similar observations have been made for HMPV.57
an infectious cDNA clone (see Chapter 38). These findings suggest that RSV has an alternate mechanism for attachment to cells that does not involve G protein, and evidence has been obtained that RSV lacking G protein can bind to heparan sulfate and possibly other molecules.111,140,406 Similar observations have been made for HMPV.57
Paramyxovirus Fusion Protein
The paramyxovirus fusion (F) proteins mediate viral penetration by fusion between the virion envelope and the host cell plasma membrane, and this fusion event occurs at neutral pH for all family members except a few isolates of HMPV, where low pH appears to have some role in fusion activation.249,362 The consequence of the fusion reaction is that the nucleocapsid is delivered to the cytoplasm. Later in infection, the F proteins expressed at the plasma membrane of infected cells can mediate fusion with neighboring cells to form syncytia (giant cell formation), which is a cytopathic effect that can lead to tissue necrosis in vivo and might also be a mechanism of virus spread.
The F proteins are homotrimers58,346,444,445 that are synthesized as inactive precursors (F0). To be biologically active, they have to be cleaved by a host cell protease at the cleavage activation site. Cleavage releases the new N-terminus of F1, thus forming the biologically active protein consisting of the disulfide-linked chains F1 and F2.167,354 The paramyxovirus F genes encode 540 to 580 residues (see Fig. 33.11). The F proteins are type I integral membrane proteins that span the membrane once and contain at their N-terminus a cleavable signal sequence that targets the nascent polypeptide chain synthesis to the membrane of the endoplasmic reticulum. At their C-termini, a hydrophobic stop-transfer domain (TM domain) anchors the protein in the membrane, leaving a short cytoplasmic tail (∼20–40 residues). Sequences adjacent to the fusion peptide and the TM anchor domain typically reveal a 4–3 (heptad) pattern of hydrophobic repeats and are designated HRA and HRB, respectively. Approximately 250 residues separate HRA and HRB (Fig. 33.15A).
Evidence has been presented that there is a second polytopic form of the NDV F protein that is 10% to 50% of the total F protein.254 The proposed second polytopic form of F has not been found for other paramyxovirus F proteins, and it is unclear why NDV F protein would be different from other F proteins. Because the second form of NDV F is only partially membrane translocated, it would have a very different protein fold from prefusion F (see Fig. 33.15), and it is unclear why NDV would uniquely require this form of F for the viral replication cycle.
The F protein is thought to drive membrane fusion by coupling irreversible protein refolding to membrane juxtaposition, initially folding into a metastable form that subsequently undergoes discrete/stepwise conformational changes to a lower energy state.183,211 The F protein found on virions is considered to be in a prefusion form; after membrane fusion has occurred, the F protein is considered to be in a postfusion form. Cleavage of F0 primes the protein for membrane fusion. The varying nature of the residues found at the cleavage site, the enzymes involved in cleavage, and the role of cleavage in pathogenesis will be discussed later.
Comparison of the amino acid sequences of paramyxovirus F proteins (reviewed in 267) does not show overall major regions of sequence identity, with the exception of the fusion peptide, which has a conserved sequence (up to 90% identity). However, the overall placement of cysteine, glycine, and proline residues suggests a similar structure for all F proteins. The Respirovirus and Rubulavirus F2 and F1 subunits are glycosylated, and there are a total of 3 to 6 potential sites for the addition of N-linked carbohydrate. For PIV5 F protein, it is known that all four potential sites for addition of N-linked carbohydrate are used.8 The measles virus F protein contains three sites in the F2 subunit for N-linked carbohydrate addition, and all three sites are used; there are no sites in F1 for N-linked carbohydrate addition.1
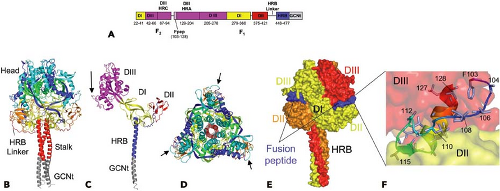
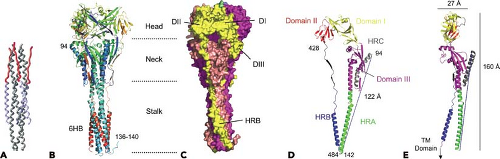 Figure 33.16. The F protein postfusion structure. A: The complete parainfluenza virus type 5 (PIV5) F1 core trimer is shown with the N1 helix colored gray and the C1 peptide colored blue except for the extended-chain N-terminal residues of C1 that are colored red. B: Ribbon diagram of the HPIV3 solF0 trimer. The three chains are colored similarly from blue (N-terminus) to red (C-terminus). Residues 95 through 135 are disordered in all chains. Residue 94 is labeled in one chain, and residues 136 through 140 at the base of the stalk are ordered in one chain owing to crystal packing interactions. C: Surface representation of the solF0 trimer. Each chain is a different color, and domains I through III and HRB for one chain (yellow) are indicated by the DI, DII, DIII, and HRB labels. One radial channel is readily apparent below domain I and II of the yellow chain and above domain III of the red chain. D: Ribbon diagram of the solF0 protein monomer colored by domain. The direct distance within one monomer between residue 94 at the end of HRC and residue 142 at the base of the stalk region is 122 Å. E: Ribbon diagram of the monomer rotated by 90 degrees, indicating the width and height of the solF0 monomer. An arrow at the C-terminus of the HRB segment points toward the likely position of the transmembrane anchor domain that would be present in the full length protein. (Adapted from Yin H-S, Paterson RG, Wen X, et al. Structure of the uncleaved ectodomain of the paramyxovirus [HPIV3] fusion protein. Proc Natl Acad Sci U S A 2005;102:9288–9293.)
Stay updated, free articles. Join our Telegram channel
Full access? Get Clinical Tree
 Get Clinical Tree app for offline access
Get Clinical Tree app for offline access

|