Flaviviridae
Brett D. Lindenbach
Catherine L. Murray
Heinz-Jürgen Thiel
Charles M. Rice
Introduction
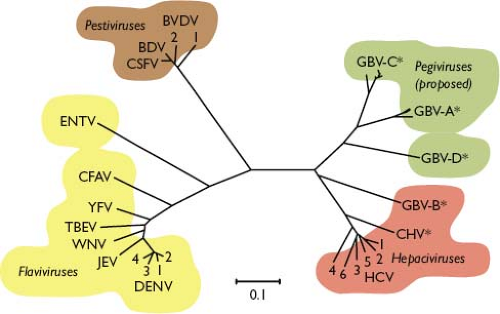
The first human virus was discovered over one century ago when Walter Reed demonstrated that yellow fever could be experimentally transferred via the filtered serum of an infected individual, and that this infectious agent was transmitted to humans by mosquitoes.817 It is now appreciated that yellow fever virus (YFV) is but one representative of a large family of related positive-strand RNA viruses, the Flaviviridae (from the Latin flavus, “yellow”). This family currently consists of three genera: Flavivirus, Pestivirus (from the Latin pestis, “plague”), and Hepacivirus (from the Greek hepar, hepatos, “liver”)851 (Table 25.1). A fourth genus, Pegivirus (persistent GB virus), has recently been proposed to encompass the previously unclassified GB virus A (GBV-A), GBV-C, and GBV-D.805 As detailed later, the Flaviviridae share similarities in virion morphology, genome organization, and replication strategy but exhibit diverse biological properties and lack serologic cross-reactivity. The phylogenetic relationships of the Flaviviridae are shown in Figure 25.1. The increasing significance of Flaviviridae as human and animal pathogens emphasizes that their study remains no less pertinent than in Reed’s time.
Family Classification
Positive-stranded RNA viruses are classified into three superfamilies based on the evolutionary relatedness of their RNA-dependent RNA polymerases (RdRPs). The Flaviviridae are members of superfamily 2, bearing distant similarity to coliphages and the plant-infecting carmo-, tombus-, diantho-, and subgroup I luteoviruses.418 Before the era of molecular biology, some members of the family Flaviviridae were classified as Togaviridae.
Family Characteristics and Replication Cycle
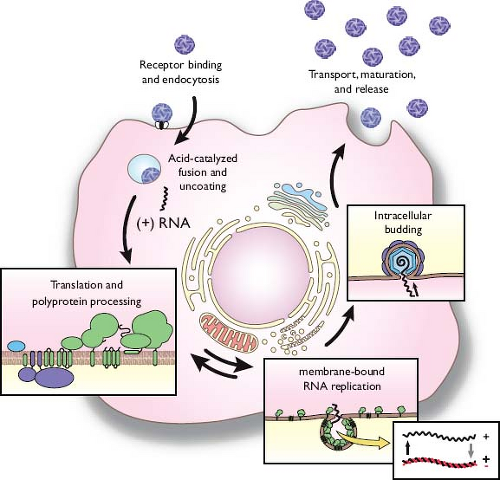
This chapter is organized around common features of the family Flaviviridae life cycle (Fig. 25.2). The enveloped virions are composed of a lipid bilayer with two or more species of envelope (E) glycoprotein surrounding a nucleocapsid, which consists of a single-stranded, positive-sense RNA genome complexed with multiple copies of a small, basic capsid (C) protein. Binding and uptake are believed to involve receptor-mediated endocytosis. The low pH of the endosome induces fusion of the virion envelope with cellular membranes. Following uncoating of the nucleocapsid, the RNA genome is released into the cytoplasm. The genome serves three discrete roles within the life cycle: as the messenger RNA (mRNA) for translation of all viral proteins, a template during RNA replication, and the genetic material packaged within new virus particles. The organization
of the genome is similar for all genera. Viral proteins are produced as part of a single polyprotein that is cleaved by a combination of host and viral proteases. The structural proteins are located in the N-terminal portion of the polyprotein with the nonstructural (NS) proteins in the remainder. Sequence motifs characteristic of a serine protease, RNA helicase, and an RdRP are found in similar locations in the polyproteins of all three genera.585 RNA replication occurs entirely in the cytoplasm in close association with intracellular membranes; the synthesis of a genome-length minus-strand RNA provides the intermediate. Progeny virions assemble by budding into an intracellular membrane compartment, most likely the endoplasmic reticulum (ER), then transit through the host secretory pathway and are released at the cell surface.
of the genome is similar for all genera. Viral proteins are produced as part of a single polyprotein that is cleaved by a combination of host and viral proteases. The structural proteins are located in the N-terminal portion of the polyprotein with the nonstructural (NS) proteins in the remainder. Sequence motifs characteristic of a serine protease, RNA helicase, and an RdRP are found in similar locations in the polyproteins of all three genera.585 RNA replication occurs entirely in the cytoplasm in close association with intracellular membranes; the synthesis of a genome-length minus-strand RNA provides the intermediate. Progeny virions assemble by budding into an intracellular membrane compartment, most likely the endoplasmic reticulum (ER), then transit through the host secretory pathway and are released at the cell surface.
Table 25.1 Members of the Flaviviridae | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||
Flaviviruses
Background and Classification
The Flavivirus genus consists of more than 50 species, many of which are arthropod-borne human pathogens. Flaviviruses cause a variety of diseases, including fever, encephalitis, and hemorrhagic fevers. Entities of major global concern include dengue virus (DENV)—with its associated dengue hemorrhagic fever (DHF) and dengue shock syndrome (DSS)—Japanese encephalitis virus (JEV), West Nile virus (WNV), and YFV (reviewed in537). Other flaviviruses with regional or endemic distribution include Murray Valley encephalitis virus (MVEV) and St. Louis encephalitis virus (SLEV). Tick-borne encephalitis virus (TBEV) is a name commonly applied to either central European encephalitis virus or Far Eastern encephalitis virus, although these are clearly distinct species.208 Decreases in mosquito control efforts during the latter part of the 20th century, coupled with societal factors (e.g., increased transportation and dense urbanization), have contributed to the re-emergence of flaviviruses such as DENV in South and Central America. Following an outbreak in New York City in 1999, WNV has spread throughout much of North America and Central America.
Flavivirus species are further categorized into antigenic complexes and subcomplexes based on serologic criteria or into clusters, clades, and species, according to molecular phylogenetics.117 Mosquito-borne and tick-borne flaviviruses, although
distinct, appear to have evolved via a common ancestral line that diverged from viruses with no known arthropod vector. DENV circulates as four distinct serotypes, which show significant sequence diversity (reviewed in337). Some reports have documented intertypic recombination among DENV isolates, although the taxonomic status of these isolates is currently unclear.
distinct, appear to have evolved via a common ancestral line that diverged from viruses with no known arthropod vector. DENV circulates as four distinct serotypes, which show significant sequence diversity (reviewed in337). Some reports have documented intertypic recombination among DENV isolates, although the taxonomic status of these isolates is currently unclear.
The development of the first live-attenuated flavivirus vaccine, YFV strain 17D,849 led to Max Theiler’s recognition by the Nobel Prize committee in 1951. Only a limited number of flavivirus vaccines are available, including inactivated TBEV and JEV for use in humans and inactivated WNV for use in animals.694 Development of effective DENV vaccines that exhibit cross-protection between serotypes is proving to be particularly challenging. The ability to genetically manipulate flaviviruses has led to novel approaches, including live attenuated chimeric vaccines based on the YFV-17D backbone.
Structure and Physical Properties of the Virion
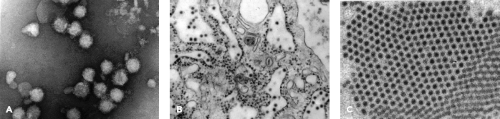
Infectious flavivirus particles are roughly spherical, approximately 50 nm in diameter, and surrounded by a lipid envelope615 (Fig. 25.3). Viruses sediment between 170 and 210S and have buoyant densities of 1.19 to 1.23 g/cm3 depending on the lipid composition, which can vary by host.745 The outer shell of the particle is made up of two viral proteins, envelope (E) and membrane (M). The E glycoprotein is the major antigenic determinant of the virion and mediates binding and fusion during virus entry. The M protein is a small proteolytic fragment of the precursor (pr)M protein and is produced during viral maturation within the secretory pathway. Removal of the lipid envelope with nonionic detergents reveals discrete nucleocapsids (120 to 140S; 1.30 to 1.31 g/cm3), which consist of capsid (C) protein and genomic RNA (reviewed in745). Isolated nucleocapsids become unstable under high salt conditions, disassembling into C protein dimers.399
Cryoelectron microscopy and image reconstruction have provided a wealth of information on flavivirus structure. Mature infectious particles of DENV430 and WNV610 display a relatively smooth outer surface. Fitting the E protein crystal structure722 into the electron density maps showed that glycoprotein dimers lie flat across the surface of the virion. Interestingly, the 180 copies of E are tightly packed in an unusual herringbone array that completely covers the lipid bilayer. Beneath the protein shell, the M protein associates closely with the membrane. Notably, the nucleocapsid lacks discernible symmetry, and neither E nor M sequences extend through the membrane to make contacts with the nucleocapsid.956
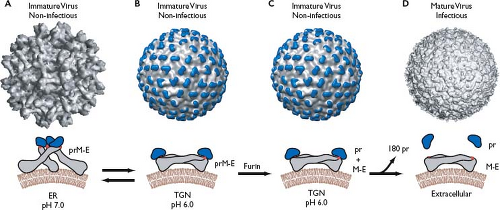
Immature flavivirus particles adopt various appearances as they egress through the secretory pathway.667 Soon after they are formed, immature virions are larger (60 nm in diameter) than mature virions and display 60 prominent spikes on their surface.957 Each protrusion is composed of three E-prM heterodimers, with the prM molecule capping the E fusion peptide. As immature particles pass through the low pH environment of the trans-Golgi network, a dramatic rearrangement of the glycoproteins occurs. The immature virions now adopt a smooth appearance almost indistinguishable from mature particles, with the exception that prM remains attached.946 This conformational change is followed by cleavage of prM by the host cell enzyme furin802 and the release of the protective fragment upon exit from the cell, revealing the mature virions.946 This process is not always efficient, and immature or partially mature particles can be released in significant quantities. Although immature particles are deemed noninfectious because they cannot undergo fusion,303 they have recently been shown to initiate infection when internalized in complex with anti-prM antibodies by cells bearing the Fc receptor.735 The mechanism of this is not well understood, but it may invoke prM cleavage in the endosome and could be especially relevant during secondary infection when antibody levels are high. In addition, partially matured particles that retain only some prM have been visualized in flavivirus populations and can undergo attachment and fusion to initiate infection similar to fully mature particles.137
Small, noninfectious subviral particles (SVPs) are the final class of particles released from flavivirus-infected cells. SVPs
contain E and M proteins but lack C and RNA.797 They complete the same maturation process as whole virions and can undergo fusion with a target cell760; due to lack of a genome, however, they are not infectious. Recombinant subviral particles (RSPs) form in cells experimentally transfected with only prM and E, indicating that interactions between these envelope proteins are sufficient to drive budding.14,512,760 RSPs are generally about 30 nm in diameter and slightly less dense than infectious virus (1.14 g/cm3),760 although virion-sized particles have also been observed in these expression systems.16,512 Cryoelectron microscopy and image reconstruction of TBEV RSPs suggests a markedly different arrangement of the E proteins compared to infectious virions. Thirty E dimers lie flat against the surface in a T = 1 icosahedral shell232 rather than a herringbone array.430 It is hypothesized that this arrangement may resemble a fusion intermediate that is adopted as E dimers rearrange to trimers upon virus entry.611
contain E and M proteins but lack C and RNA.797 They complete the same maturation process as whole virions and can undergo fusion with a target cell760; due to lack of a genome, however, they are not infectious. Recombinant subviral particles (RSPs) form in cells experimentally transfected with only prM and E, indicating that interactions between these envelope proteins are sufficient to drive budding.14,512,760 RSPs are generally about 30 nm in diameter and slightly less dense than infectious virus (1.14 g/cm3),760 although virion-sized particles have also been observed in these expression systems.16,512 Cryoelectron microscopy and image reconstruction of TBEV RSPs suggests a markedly different arrangement of the E proteins compared to infectious virions. Thirty E dimers lie flat against the surface in a T = 1 icosahedral shell232 rather than a herringbone array.430 It is hypothesized that this arrangement may resemble a fusion intermediate that is adopted as E dimers rearrange to trimers upon virus entry.611
Binding and Entry (see Video in e-Book)
Flaviviruses infect a variety of target cells through receptor-mediated endocytosis, followed by intracellular membrane fusion. Flavivirus receptors are not well characterized, perhaps because these viruses use a range of entry factors for different cell types and employ more than one host molecule to enter a target cell. Highly sulphated glycosaminoglycans, such as heparin sulphate, are ubiquitously expressed molecules used as initial attachment factors by many viruses. These factors have also been shown to play a role in binding and entry of flaviviruses, such as DENV,132 YFV,269 TBE,426 and JEV.130 However, high affinity for glycosaminoglycans in tissue culture–adapted strains is associated with in vivo attenuation.458 C-type lectins are cellular proteins that bind mannose-rich glycans and are involved in flavivirus infection of dendritic cells (DCs). Intradermal DCs are often the primary targets encountered by the arthropod-delivered pathogens and can transport the virus to draining lymph nodes where a second round of replication initiates viremia. C-type lectin Dendritic cell-specific intercellular adhesion molecule 3-grabbing nonintegrin (DC-SIGN) is thought to function as an attachment receptor for DENV infection of DCs.516,624,833 WNV preferentially uses DC-SIGN-related (DC-SIGNR),182 while YFV-17D, which lacks glycan modifications on E, can infect DC cells in a lectin-independent manner.39 The mannose receptor is a C-type lectin that is constitutively internalized by clathrin-mediated endocytosis and has been suggested to play a role in endocytosis of DENV, JEV, and TBEV.584 Interestingly, C-type lectin domain family 5, member A (CLEC5) interacts with DENV but does not mediate its entry; instead, CLEC5 binding triggers the release of inflammatory cytokines, leading to DHF/DSS-like symptoms in mice.131
Additional receptors that have been implicated in flavivirus entry include the glycosphingolipid neolactotetraosylceramide,26 low-density lipoprotein receptor (LDL-R),138 laminin receptor 1,850,857 αvβ3 integrins,144 and a CD14-containing complex.134 Heat shock proteins have also been suggested as entry factors. GRP78 (BIP) plays a role in liver cell uptake of DENV,112,368 and Hsp90/Hsp70 acts in DENV and JEV entry into human monocytes/macrophages, neuroblastoma cell lines, and mosquito cells.180,720,723 Finally, virus particles opsonized with subneutralizing concentrations of immunoglobulins show enhanced binding and infection of cells expressing Fc receptors.661,764 It is widely speculated that antibody-enhanced infection is relevant to the pathogenesis of DSS and DHF, which occur more frequently in people previously exposed to other DENV serotypes.
After capture by the appropriate receptor(s), flaviviruses are internalized by endocytosis. Single particle tracking of fluorescently labeled DENV particles has shown that virions diffuse across the surface of the cell until they encounter a preformed clathrin-coated pit.871 Following internalization, DENV particles are delivered to early or intermediate endosomes, which then mature into late endosomes.871 Fusion of the viral and host membranes occurs during endosomal trafficking, although the exact compartment that triggers this event seems to differ between strains and flavivirus species, perhaps indicating an optimal pH.871 In the acidic environment, E protein dimers dissociate and undergo an irreversible conformational change to become fusogenic trimers.13,810 The fusion peptide, previously buried at the E homodimer interface, is exposed and inserts into the endosomal membrane (Video 25.1). The efficiency of fusion is influenced by the lipid composition of target membranes: cholesterol, oleic acid, and anionic lipids such as bis(monoacylglycero)phosphate and phosphatidylserine enhance fusion, whereas lyophosphatidylcholine inhibits the process.812,813,950 Lipid composition can also influence the pH threshold of fusion.420 Following fusion, viral genomes are immediately accessible for translation.420
Genome Structure
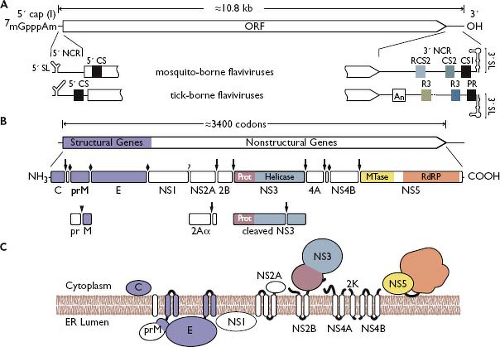
As for other positive-strand RNA viruses, the genomes of flaviviruses are infectious.662 Full-length infectious complementary DNA (cDNA) clones have been constructed for several species, allowing flavivirus biology to be dissected by reverse genetics.491,725,742 Flavivirus genomes consist of a single, positive-strand RNA of approximately 11 kilobases (kb) in length (sedimentation, 42S) with a 5′ type 1 cap, m7GpppAmN151,901 (Fig. 25.4). The cap structure serves to stabilize the viral RNA, initiate translation, and subvert innate antiviral defenses.178,253 Unlike cellular mRNAs, flavivirus genomes lack a 3′ polyadenylate tail.901 Genomes encode a single long open reading frame (ORF, ∼3,400 codons) flanked by 5′ and 3′ noncoding regions (NCRs) of ∼100 nucleotides (nt) and 400 to 700 nt, respectively544 (Fig. 25.4).
The sequence of the 5′ NCR is not well conserved between flaviviruses, although common secondary structural elements have been identified, including a bifurcating 5′ stem-loop (5′ SL). These structures influence viral genome translation, as antisense oligonucleotides complementary to the 5′ SL abolish DENV translation and replication, and second-site mutations in this region compensate for replication defects caused by reduced viral cap methylation. In addition, 5′ SL likely acts as a promoter to initiate RNA replication by binding the viral NS5 polymerase/methyltransferase protein.196,236,237,506 Consistent with this, deletions within the 5′ NCR cause severe defects in DENV-4 replication, but not viral translation.113 Interestingly, one of the viable mutants exhibited a limited host-range growth phenotype, suggesting that host-specific factors interact with either the 5′ NCR or the complementary 3′ end of the negative strand. Indeed, several human proteins, including La and TIAR, can bind to the 3′ end of negative-strand RNA.472,778,938 WNV replication is inhibited in a TIAR-knockout cell line,472 and mutagenesis of the TIA-1/TIAR binding sites suggests a role in initiating positive-strand RNA synthesis.217
The organization of the 3′ NCR differs between mosquito-borne viruses, tick-borne viruses, and viruses with no known vector. Nevertheless, conserved regions, sequence duplications, and predicted RNA secondary structures are shared among the groups (Fig. 25.4). The greatest structural similarity is a long (90 to 120 nt) 3′ stem-loop (3′ SL) that differs in primary sequence between mosquito-borne and tick-borne flaviviruses.296 Mutational analysis of DENV-2 and WNV revealed essential virus-specific and host-specific functional regions within the 3′ SL.216,856,947,953 The 3′ SL enhances translation of reporter mRNAs containing the DENV 3′ NCR,140,335 while DENV-2 translation and replication were inhibited by a corresponding antisense oligonucleotide.336 The 3′ SL also interacts with several important proteins, including the viral NS2A, NS3, and NS5 proteins129,175,535 and translation elongation factor 1A (EF1A).75,183,185 These results are intriguing, because EF1A, and its prokaryotic homolog EF-Tu, contribute to the replication of other positive-strand RNA viruses.81,314,379,951 In addition, the human La autoantigen,185,262,875 polypyrimidine tract binding (PTB) protein,185 and murine Mov34 protein822 were found to bind 3′ SL of DENV-4 and JEV, although the functional relevance of these interactions is presently unknown.
Upstream of the 3′ SL lie conserved sequence repeats (CS1, CS2, CS3, RCS2, and RCS3), secondary structures, and putative pseudoknots.296 Some of these structures confer resistance to the cellular 5′-3′ exoribonuclease Xrn-1,252,785 suggesting that this region of the genome forms a compact structure.
Flavivirus genomes can be circularized through long-distance base pairing between elements located near the 5′ and 3′ ends. In mosquito-borne flaviviruses, these interactions are mediated by the 5′ UAR (upstream of AUG region), 5′ DAR (downstream of AUG region), and 5′ CS (conserved sequence), which base-pair with the 3′ UAR, 3′ DAR, and 3′ CS1 regions, respectively, located more than 10 kb downstream at the base of the 3′ SL.354 A distinct set of long-distance interactions circularizes the genomes of tick-borne flaviviruses.394,410 These long-range base pairs are important for RNA replication, presumably by bringing the 5′ SL–bound NS5 protein in proximity to the 3′ site of minus-strand initiation.18,96,167,196,236,237,248,354,394,410,940 It should be noted that genome circularization requires the melting of local secondary structures within the 5′ and 3′ ends and leads to the occlusion of the translation start site. Thus, large-scale conformation changes within the flavivirus genome may regulate the switch from translation to RNA replication.
Translation and Proteolytic Processing
The efficiency of genome translation is a primary determinant of flavivirus infectivity.209 The viruses therefore use several mechanisms to ensure translational competence, including specialized structures within the 5′ and 3′ NCRs. Translation is cap dependent, and 2′-O methylation of the 5′ cap helps to overcome innate antiviral defenses that down-regulate translation in infected cells.178 While translation initiates via ribosomal scanning, many mosquito-borne flaviviruses lack a canonical Kozak initiation motif and contain several AUG codons near the correct start site. To help ensure proper AUG selection, DENV uses a small RNA stem-loop embedded within the C gene to induce ribosomal pausing over the authentic initiation codon.155
Translation of the single, long ORF produces a large polyprotein that is co- and posttranslationally cleaved into at least 10 proteins (Fig. 25.4B). The N-terminal region of the polyprotein encodes the structural proteins (C-prM-E), which are followed by the NS proteins (NS1-NS2A-NS2B-NS3-NS4A-2K-NS4B-NS5).119,120,726 Host signal peptidase is responsible for cleavages between C/prM, prM/E, E/NS1, and 2K-NS4B. A virus-encoded serine protease, NS2B-3, processes at the NS2A/NS2B, NS2B/NS3, NS3/NS4A, NS4A/2K, and NS4B/NS5 junctions. The enzyme responsible for NS1–2A cleavage is presently unknown. The expected topology of the flavivirus polyprotein is depicted in Figure 25.4C.
Features of the Structural Proteins
C Protein
Capsid (C) protein is a highly basic protein of ∼11 kD. The nascent protein contains a C-terminal hydrophobic tail that serves as a signal peptide for ER translocation of prM. This anchor is cleaved in two steps, first by the viral NS2B-3 protease and then by signal peptidase.503 Mature C protein folds into a compact dimer with each monomer containing four α-helices193,372,529 (Fig. 25.5). The N-terminal region of the protein remains unstructured and, along with charged residues at the C-terminus, is thought to be involved in RNA binding.398 An internal hydrophobic region mediates membrane association of C.529
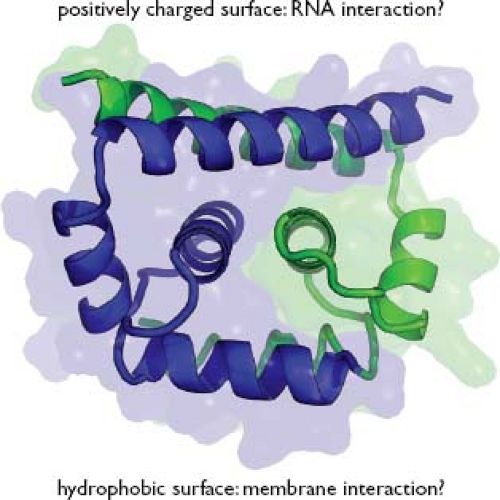 Figure 25.5. Flavivirus C protein structure. The WNV-KUN C protein is shown as a ribbon diagram with the protein surface rendered transparent, from PDB accession number 1SFK.193 One monomer of the dimer is colored blue, the other green. |
Overall, flavivirus C proteins demonstrate remarkable functional flexibility, with tolerance for large deletions. YFV C retains its ability to package RNA even after deletion of nearly 40 residues from the N-terminus or 27 residues of the C-terminus; internal deletions of the hydrophobic sequence are less tolerated.654 The TBEV C protein can accept deletions of up to 16 amino acids from the central hydrophobic helix, albeit with increased production of empty particles.409 Mutants containing larger deletions are not viable but can be rescued by second-site changes that increase the hydrophobicity of downstream sequences.411 WNV tolerates small deletions in hydrophobic helix α2 to various degrees. Remarkably, infectivity of the deleted genomes was improved by even larger deletions—up to one-third of the C protein sequence—encompassing all of helix α3.765 This results in the loss of a hydrophilic stretch, again suggesting the importance of hydrophobicity. It is not yet clear how C protein dimers are organized within the apparently disordered nucleocapsids, but interaction with RNA or DNA can induce isolated C protein dimers to assemble into nucleocapsid-like particles in vitro.399
Membrane Glycoprotein prM
The glycoprotein precursor of M, prM (∼26 kD), is translocated into the ER via a signal sequence provided by the hydrophobic tail of C. Signal peptidase cleavage is delayed, however, until the viral serine protease cleaves on the cytosolic side of the membrane to generate the mature form of C.19,503,924 This delay seems to result from the combination of a fairly short (14 to 22 amino acids) signal sequence, suboptimal residues at the signalase cleavage site, and residues in downstream regions of prM504,815 and E protein.511 Interestingly, uncoupling signal peptidase cleavage from NS2B-3 processing leads to increased production of empty virus particles.459,504,505 Coordinated cleavage therefore serves to delay structural protein processing until the viral serine protease has accumulated and replication is under way, which may limit the release of immunogenic but noninfectious SVPs early in infection.
The N-terminal region of prM contains one to three N-linked glycosylation sites122 and six conserved cysteine residues, all of which are disulfide linked.633 The prM protein folds rapidly and assists in the proper folding of E.415,511 The C-terminal transmembrane (TM) domains of prM and E act as ER retention signals and may assist in their heterodimerization.486,641,643 A major function of prM is to prevent E from undergoing acid-catalyzed rearrangement and fusion during transit of the virions through the secretory pathway.302,322 The crystal structure of DENV prM in complex with E has recently been solved and demonstrates how this function is performed.471 The pr domain is a unique fold consisting of seven β strands, with the previously identified disulfide bonds stabilizing the structure. In immature particles, the pr region sits at the tip of the E protein, forming the pr-E spike and shielding the fusion peptide from the cellular environment (Fig. 25.6A). prM is not accessible to furin cleavage in these particles due to steric hindrance.471 The acidity of the trans-Golgi compartment induces a global rearrangement that exposes the furin cleavage site.946 After cleavage, the pr peptide does not immediately
disassociate from the virus particle.802 Exposure to the neutral pH of the extracellular space is required to release pr and reveal the fusion-competent mature virion. This delay prevents the cleaved particles from undergoing premature membrane fusion within the Golgi.
disassociate from the virus particle.802 Exposure to the neutral pH of the extracellular space is required to release pr and reveal the fusion-competent mature virion. This delay prevents the cleaved particles from undergoing premature membrane fusion within the Golgi.
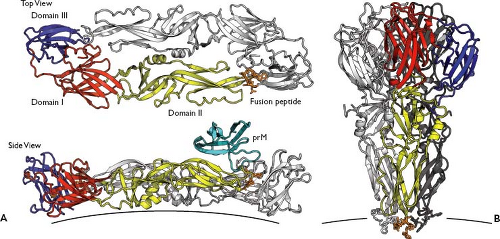 Figure 25.6. Flavivirus glycoprotein structures. A: The structure of a dengue virus 2 (DENV-2) E glycoprotein dimer is represented in this ribbon diagram, as viewed perpendicular (top) or laterally (bottom) with respect to the lipid bilayer. One E monomer is colored red (domain I), yellow (domain II), and blue gray (domain III). The amino acid side chains of the fusion peptide are shown (orange). Rendered from PDB 1OAN.591 In the bottom panel, the low pH conformation of the DENV-2 pr protein from PDB 3C5X471 was modeled onto the structure of 1OAN. B: tick-borne encephalitis virus (TBEV) E protein trimers in their postfusion form, colored as in A. Rendered from PDB 1URZ.98 |
Envelope Glycoprotein
E protein (∼53 kD) is the major protein on the surface of flavivirus virions. E is synthesized as a type I membrane protein containing 12 conserved cysteines that form disulfide bonds634; in some viral species E is N-glycosylated.123,909 Proper folding, stabilization in low pH, and secretion of E depends on co-expression with prM.415,511 E is a class II fusion protein that mediates both receptor binding and membrane fusion.
Atomic resolution structures of E proteins from several flaviviruses have been solved in pre- and postfusion conformations.98,382,591,592,593,625,635,722,958 In its prefusion form, E folds into an elongated structure rich in β-sheets and forming head-to-tail homodimers that lie parallel to the virus envelope.591,722 Each E protein is composed of three domains: DI, which forms an eight-stranded β-barrel; DII, a long, finger-like domain that projects along the virus surface; and DIII, which maintains an immunoglobulin-like fold (Fig. 25.6A). The fusion peptide12 is located at the tip of DII and remains covered by the pr peptide or buried in a hydrophobic pocket formed by DI and DIII of the partner monomer until triggered to insert into the target cell membrane.722 DIII projects slightly from the virion surface and is thought to be involved in receptor binding; it is a major target of neutralizing antibodies.145 Between the ectodomain of E and the membrane is a short but functionally important stem region composed of two α-helices that lie parallel to the plane of the membrane.15,956
On exposure to low pH, E protein dimers dissociate into monomeric subunits, which then form fusogenic trimers.13,810,811 Interestingly, the WNV E protein crystal structure shows an array of perpendicular monomers, suggesting a mechanism for E protein rotation without exposing the fusion loop.635,814 Crystal structures of postfusion E show the protein folded back onto itself, bringing the N-terminal fusion peptide, with its associated cellular membrane, into proximity with C-terminal TM domain, which is still integrated in the viral membrane98,592 (Fig. 25.6B). To accomplish this, DIII must rotate and fold back more than 30 Å in relation to DI. Indeed, neutralizing antibodies against DIII can inhibit a postattachment step of viral entry,636 and a soluble, recombinant form of DIII is a potent dominant-negative inhibitor of fusion.478 In addition, DII rotates relative to DI,98,592 with similar displacement of DII seen in crystals of native E protein grown in the presence of the detergent β-octylglucoside.591,958 Residues that influence the pH threshold for membrane fusion surround the DI/DII pocket.591 Protonation of conserved histidines at the interface of DI and DIII also contribute to E domain rearrangements in TBEV RSPs.249 Mutagenesis studies of WNV, however, failed to identify histidine residues that entirely control the switch.629
Features of the Nonstructural Proteins
NS1 Glycoprotein
The NS1 glycoprotein (∼46 kD) is translocated into the ER during synthesis and processed at its N-terminus by host signal
peptidase. The C-terminus of NS1 arises through NS1–2A cleavage by an unknown ER-resident host enzyme, which requires the eight C-terminal residues of NS1 and greater than 140 amino acids of NS2A.227,228,340 In addition, JEV expresses an elongated form of NS1, termed NS1′, which arises through a ribosomal −1 frameshifting event.80,238,547,565
peptidase. The C-terminus of NS1 arises through NS1–2A cleavage by an unknown ER-resident host enzyme, which requires the eight C-terminal residues of NS1 and greater than 140 amino acids of NS2A.227,228,340 In addition, JEV expresses an elongated form of NS1, termed NS1′, which arises through a ribosomal −1 frameshifting event.80,238,547,565
NS1 contains two or three N-linked glycosylation sites and 12 conserved cysteines that form disulfide bonds.79,461,547,797,884 Around 30 minutes after synthesis, NS1 simultaneously forms highly stable homodimers and acquires an affinity for membranes.910,911 As NS1 lacks a known membrane interaction domain, the nature of its membrane association remains unclear. One possibility is that dimerization creates a hydrophobic surface for peripheral membrane binding. Alternatively, it has been reported that DENV-2 NS1 exhibits properties of a glycosylphosphatidylinositol (GPI)-anchored protein, although this mechanism seems inconsistent with the C-terminal peptide sequence of this protein.902
NS1 is retained within a secretory-derived compartment, expressed on the surface of infected cells, and efficiently secreted from mammalian, but not insect, cells.491,816 The relative distribution of NS1 within these compartments is regulated through an unknown mechanism involving a short, N-terminal region of the protein.942 The secreted form of NS1 accumulates to high levels in human sera and tissues and can be used to diagnose flavivirus infections at an early stage.8,147,531 Secreted NS1 forms soluble, hexameric lipoprotein particles of ∼10 nm that appear as three dimers held together in a barrel configuration.172,239,304 The secreted form of NS1 can bind to uninfected cells by interaction with sulfated glycosaminoglycans33 and can be internalized and trafficked to late endosomes, where it accumulates.9 The function of endocytosed NS1 is not yet clear, but it may enhance subsequent infection with the homologous virus.9
The intracellular form of NS1 localizes to sites of viral RNA synthesis and plays an essential role in genome replication.533,904 Mutations in NS1 can lead to dramatic defects in RNA replication and infectious virus production.170,619,620 trans-Complementation studies revealed that NS1 functions at an early stage in RNA replication through a genetic interaction with NS4A.395,396,490,492
The extracellular forms of NS1 are highly antigenic and induce a strong humoral response. Secreted NS1 was originally characterized as a soluble, complement-fixing antigen present in the serum and tissues of DENV-infected animals. Antibodies that recognize cell surface–bound NS1 can direct complement-mediated lysis of infected cells and protect animals from lethal disease; other NS1-specific antibodies are protective in a complement-independent manner.148,149 Antibody-mediated cross-linking of cell surface NS1 can also induce signaling cascades in DENV-2–infected cells, and it has been proposed that NS1 may contribute to pathogenesis by inducing antibodies that cross-react with human proteins. Despite the strong link between NS1-specific humoral responses and complement fixation, recent evidence indicates that WNV NS1 can inhibit the alternative pathway of complement activation by binding to and inhibiting the serum protein factor H. Furthermore, DENV, WNV, and YFV NS1 inhibit the classical pathway of complement fixation by binding to and increasing the turnover of complement factor C4. NS1 clearly plays an important role in flavivirus-specific humoral responses.
NS2A and NS2B Proteins
NS2A is a relatively small (∼22 kD) hydrophobic protein. Its N-terminus is generated by an unidentified ER-resident host enzyme,228 whereas the C-terminus is generated by NS2B-3 cleavage in the cytoplasm. Thus, NS2A is membrane spanning, although the precise topology of the protein is unknown. In addition, the YFV serine protease can cleave at an internal site in NS2A to generate a C-terminally truncated form, NS2Aα.123,630 Interestingly, mutations at the YFV NS2Aα cleavage site block virus particle production and can be suppressed by a second mutation on the surface of the NS3 helicase domain.434 Mutations in KUNV NS2A similarly block virus assembly, while the protein also localizes to subcellular sites of RNA replication and interacts with replicase components NS3, NS5, and the 3′ NCR of genome RNA. The involvement of NS proteins, in particular the NS2–3 region, in replication and infectious virus assembly appears to be an emerging theme for all three genera of the family Flaviviridae.
DENV-2 and WNV NS2A can also inhibit interferon (IFN) signaling, as evidenced by specific mutations in the protein that diminish this inhibitory activity and attenuate virulence in mice.498,500,613 Interestingly, these mutations are cell culture adaptive and enhance the ability of KUNV replicons to establish persistence in IFN-competent cell lines. Remarkably, NS2A of the tick-borne flavivirus Langat does not share this property, which appears to be carried out instead by NS5.66
NS2B is also a small (∼14 kD) membrane-associated protein.154 NS2B forms a stable complex with NS3, serves to anchor this complex to cellular membranes, and acts as an essential co-factor for the NS2B-3 serine protease.229 The co-factor activity lies in a central peptide that intercalates within the fold of the serine protease domain.31,125,221
NS3 Protein
NS3 is a large (∼70 kD) multifunctional protein, encoding enzymatic activities required for polyprotein processing and RNA replication. The N-terminal third of the protein is the catalytic domain of the NS2B-3 serine protease,50,124,279 which has specificity for substrates containing adjacent basic residues at the NS2A/NS2B, NS2B/NS3, NS3/NS4A, and NS4B/NS5 junctions.122 In addition, this protease generates the C-termini of mature capsid protein19,924 and NS4A481 and can cleave at internal sites within NS2A and NS3.
Soluble, recombinant forms of the NS2B-3 serine protease domain have been purified and crystallized for x-ray diffraction.125,221,733 These structures show that the co-factor region of NS2B contributes a β-strand to complete the chymotrypsin-like fold of the protease (Fig. 25.7A), similar to hepatitis C virus (HCV) NS3-4A. The C-terminal region of the NS2B co-factor can adopt multiple conformations that may alternately help to form the substrate-binding pocket or project outward from the protease fold; it is not yet clear whether these structural rearrangements are biologically significant.
As for other members of the Flaviviridae, the C-terminal region of NS3 encodes a supergroup 2 RNA helicase–nucleoside triphosphatase (NTPase).280 NS3 demonstrates RNA-stimulated NTPase and RNA unwinding activities890,899 and mutagenesis of the active site residues confirmed that these activities are essential for viral replication.549 This region of NS3 also exhibits RNA triphosphatase (RTPase) activity, proposed to dephosphorylate the 5′ end of the genome before cap addition.900 Recent
studies show that RTPase is dependent on the Walker B motif in the helicase–NTPase catalytic core for phosphodiester bond hydrolysis.41,58 Thus, all three nucleic acid–modifying activities of NS3 rely on a common active center. In addition to its roles in RNA replication, the helicase domain of NS3 has been implicated in virus assembly,434,655 a role that is separable from the known enzymatic activities and can function in trans.655
studies show that RTPase is dependent on the Walker B motif in the helicase–NTPase catalytic core for phosphodiester bond hydrolysis.41,58 Thus, all three nucleic acid–modifying activities of NS3 rely on a common active center. In addition to its roles in RNA replication, the helicase domain of NS3 has been implicated in virus assembly,434,655 a role that is separable from the known enzymatic activities and can function in trans.655
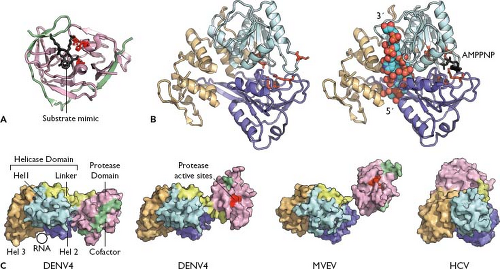 Figure 25.7. Viral NS3 proteins. A: The structure of the West Nile virus (WNV) NS2B-3 serine protease domain, with NS2B co-factor peptide (green), NS3 protease domain (pink), and a substrate-based inhibitor (black). Active-site residues are shown in red. Rendered from PDB number 2FP7.221 B: Structure of the dengue fever 4 (DENV-4) NS3 helicase domain with (left) or without (right) bound RNA substrate (colored spheres) and adenosine triphosphate (ATP) analog (black). Note the adenosine triphosphatase (ATPase) active site (red) becomes structured at the interface of domain I (cyan) and domain II (purple) upon RNA binding. The RNA is bound within a cleft formed by the first two domains and domain III (gold). Rendered from PDB numbers 2JLQ and 2JLV.523 C: Structural interdomain flexibility within full-length NS3. Shown are two conformations of full-length DENV-4 NS3 (PDBs 2VBC522 and 2WHX,521 respectively), Murray Valley encephalitis virus (MVEV) (PDB 2WV931), and hepatitis C virus (HCV) (PDB 1CU1930). Shown is a structural alignment of the helicases, with molecular surfaces colored as in A and B and the protease-helicase linker in yellow. |
Crystal structures of isolated DENV and YFV NS3 helicase domains show three subdomains, two structurally conserved RecA-like domains that are involved in NTP hydrolysis, and a unique C-terminal domain that may be involved in virus-specific RNA and protein recognition.523,916,921 Co-crystallization of the DENV helicase with or without RNA substrates and nucleoside analogs revealed the structural basis for RNA-stimulated adenosine triphosphatase (ATPase) activity and substrate unwinding523 (Fig. 25.7B). The structures of full-length DENV and MVEV NS3 have recently been solved in complex with their corresponding NS2B co-factors.31,521,522 In these structures, the serine protease and helicase regions largely retain their domain folds, forming an elongated binary complex. However, the relative orientation of the domains differ, and greatly differ from the orientation of the related domains in HCV NS3, implying that the flexible linker region may play an important role in coordinating enzyme activities (Fig. 25.7C). The DENV structure also revealed that the serine protease domain can contribute to RNA helicase activity.522
Truncated forms of NS3, which result from alternative serine protease cleavage events in the helicase domain, have been observed in vitro and in vivo. The role of these cleavages is unclear, although it is possible that the products could have a distinct function. In this regard, replication defects caused by deletions in the KUNV helicase domain can be complemented in trans, while deletions in the serine protease domain cannot be complemented.374,396,499
Finally, the NS3 proteins of Langat, DENV-2, and WNV have been shown to induce apoptosis, in some cases through activation of caspase-8.691,704,776 Flaviviruses are often cytopathic in mammalian cells, although whether this is the normal pathway for cell killing requires further study. The DENV NS2B-3 serine protease can also down-regulate the activation of type I IFN in human dendritic cells, although the relevant protease substrate(s) have not been identified.736
The NS4A and NS4B Proteins
NS4A and NS4B are small (16 kD and 27 kD, respectively) hydrophobic proteins. NS4A has been implicated in RNA replication through a genetic interaction with NS1490 and co-localization with replication complexes.535 Similar to the
coordinated processing of C protein, signal peptidase cleavage at the 2K/NS4B junction requires prior cleavage by the NS2B-3 serine protease at a site just upstream of the 2K internal signal peptide.481,690 Overexpression studies showed that NS4A can induce membrane rearrangements and/or formation of autophagosomes, and that regulated NS4A/2K/4B cleavage is necessary for this activity.560,586,738 Mutations in NS4A and 2K have been found to confer resistance to a potent inhibitor of flavivirus RNA replication and to overcome superinfection exclusion, further implicating this region in RNA replication.966,967
coordinated processing of C protein, signal peptidase cleavage at the 2K/NS4B junction requires prior cleavage by the NS2B-3 serine protease at a site just upstream of the 2K internal signal peptide.481,690 Overexpression studies showed that NS4A can induce membrane rearrangements and/or formation of autophagosomes, and that regulated NS4A/2K/4B cleavage is necessary for this activity.560,586,738 Mutations in NS4A and 2K have been found to confer resistance to a potent inhibitor of flavivirus RNA replication and to overcome superinfection exclusion, further implicating this region in RNA replication.966,967
NS4B is a polytopic membrane protein that co-localizes with NS3 at the presumed sites of RNA replication.481,587 NS4B is posttranslationally modified to a form that migrates faster on sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE),123,690 although the identity and function of this modification remain to be determined. Similar to NS2A, DENV NS4A and NS4B can block type I IFN signaling.613 NS4B has the strongest antagonistic effect, which requires either proper processing of the NS4A–NS4B polyprotein or expression of NS4B with an N-terminal signal peptide.612 Similarly, WNV NS4A and NS4B block IFN signaling by inducing an unfolded protein response in the ER, which can down-regulate Jak-STAT signaling.20
NS5 Protein
NS5 is a large (103 kD), highly conserved, multifunctional phosphoprotein with RNA capping and RdRP activities encoded within its N- and C-terminal regions, respectively.181 Formation of a type 1 RNA cap involves multiple steps, including (a) removal of one phosphate from a 5′ triphosphorylated RNA substrate by an RTPase, (b) addition of a 5′-5′ guanosine cap (from guanosine triphosphate [GTP]) by a guanylyltransferase, (c) N7-methylation of the guanylyl cap by a methyltransferase (MTase), and (d) 2′-O methylation of the second residue by the same or another MTase. As mentioned earlier, the NS3 helicase–NTPase exhibits RTPase activity, which suggests that NS3 and NS5 function together during RNA capping.41,900 The flavivirus guanylyltransferase proved to be elusive for many years, although recent evidence indicates that the N-terminal domain of NS5 is capable of performing this reaction.361 Finally, the N-terminal region of NS5 encodes conserved MTase motifs and is capable of performing both N7 and 2′-O methylation in a coordinated fashion.211,417,713 The structure of the NS5 capping domain has been solved by x-ray crystallography under a variety of conditions, revealing high-resolution structures of these reaction pathways181,211,266,931 (Fig. 25.8A). Mutagenesis of the NS5 capping domain showed that the methylation events are separable and that N7 methylation is required for viral translation and replication, while 2′-O methylation allows the virus to avoid innate antiviral defenses.178,195,427,964 Interestingly, cellular casein kinase 1 can phosphorylate YFV NS5 in vitro at a serine residue near the methyltransferase active site, which inhibits 2′-O methylation.67,68 While inhibitors of this kinase affect YFV replication, it is not known if this site is phosphorylated in vivo.
The C-terminal domain of NS5 contains conserved RdRP motifs417,726 and structurally resembles other RNA polymerases, forming a “right hand” structure with palm, fingers, and thumb subdomains181,543,932 (Fig. 25.8B). NS5 RNA polymerase activity has been confirmed with purified, recombinant protein1,181,305,826; mutagenesis of the polymerase active site; and supplying the activity in trans from a KUNV replicon. The major product of in vitro RdRP reactions is often a self-primed copy-back RNA. However, NS5 is capable of initiating RNA synthesis de novo,1,632,948 which likely reflects the authentic mechanism in infected cells. NS5 forms a complex with NS3371,389 and stimulates NS3 NTPase and RTPase activities.175,939 Cross-linking studies have shown that both proteins bind to the 3′ SL of the viral genome.129 Along with genome circularization, this may serve to initiate minus-strand synthesis by 5′ SL–bound NS5.196,237,354,940
WNV and DENV-2 NS5 have been shown to localize at sites of viral RNA synthesis,534,898 although this has been
difficult to show for other viruses. Biochemical studies indicate that only a small fraction of NS5 co-fractionates with replicase activity,298,868 and the protein is frequently localized to the nucleus of flavivirus-infected cells.181,389 These results suggest that NS5 may play additional roles, other than in RNA replication, in the virus life cycle. In this regard, DENV-2 NS5 induces interleukin-8 (IL-8) transcription and secretion, which may enhance viral spread or disease by recruiting inflammatory cells to the site of infection.562 In addition, NS5 has been shown to block the Jak-STAT pathway of IFN signaling.66,181
difficult to show for other viruses. Biochemical studies indicate that only a small fraction of NS5 co-fractionates with replicase activity,298,868 and the protein is frequently localized to the nucleus of flavivirus-infected cells.181,389 These results suggest that NS5 may play additional roles, other than in RNA replication, in the virus life cycle. In this regard, DENV-2 NS5 induces interleukin-8 (IL-8) transcription and secretion, which may enhance viral spread or disease by recruiting inflammatory cells to the site of infection.562 In addition, NS5 has been shown to block the Jak-STAT pathway of IFN signaling.66,181
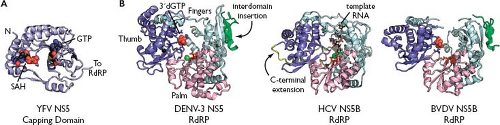 Figure 25.8. Viral NS5 and NS5B proteins. A: The yellow fever virus (YFV) capping domain is shown, with bound methyl donor S-adenosyl-L-homocysteine (SAH) and guanosine triphosphate (GTP). Rendered from PDB number 3EVC.266 B: A structural comparison of RNA-dependent RNA polymerases (RdRP) domains across the Flaviviridae. The dengue virus 3 (DENV-3) RdRP domain is shown, with the canonical finger (cyan), palm (pink), and thumb (purple) domains indicated. A flavivirus-specific interdomain insertion is shown in green. This structure shows the binding site of a GTP analog, which is important for activating de novo RNA synthesis. GTP-binding residues are blue, RdRP active-site residues are red, and a catalytic Mg2+ ion is shown as a green sphere. Structural Zn2+ ions are shown in black. This model is a composite structure, rendered from PDB numbers 2J7U and 2J7W.932 The hepatitis C virus (HCV) NS5B RdRP is shown with bound template RNA; the HCV-specific C-terminal extension is shown in yellow. Rendered from PDB number 1NB7.637 The bovine viral diarrhea virus 1 (BVDV-1) NS5B RdRP is shown modeled in the GTP-bound state by rendering PDB number 2JCQ and the GTP analog from 2J7W. The pestivirus-specific N-terminal extension is shown in green. |
RNA Replication
The flavivirus NS proteins presumably recruit the viral genome out of translation and into a replication complex. Replication begins with the synthesis of a genome-length minus-strand RNA, which then serves as a template for new plus-strand genomes. Minus-strand RNA has been detected as early as 3 hours after infection.492 Viral RNA synthesis is asymmetric, with approximately 10-fold more positive strands accumulating compared to minus strands.152,620 Flavivirus replication can be followed by metabolic labeling of virus-specific RNA in the presence of actinomycin D, an inhibitor of DNA-dependent RNA polymerases. Three major species of labeled flavivirus RNA have been described, including the plus-strand genome, a double-stranded replicative form (RF), and a heterogeneous population of replicative intermediates (RIs) that most likely represent duplex regions and recently synthesized RNAs displaced by nascent strands undergoing elongation.146,152 Pulse-chase analyses indicate that RF and RI are precursors to genome RNA,146,152 indicating semiconservative and asymmetric replication.146
In addition to genome-length plus- and minus-strand products of RNA replication, 0.2- to 0.6-kb subgenomic flavivirus RNAs (sfRNAs) are produced in infected cells.485,762,869 sfRNAs are co-linear with the 3′ end of the genome and are produced through incomplete degradation of the genome by the cellular 5′-3′ exoribonuclease Xrn1.678,785 The Xrn1 resistance of this region is due to conserved secondary structures and pseudoknots located within the 3′ NCR.252,785 While controversy exists over whether sfRNAs are needed for efficient RNA replication, a WNV mutant that does not produce sfRNAs is less cytopathic in cell culture and less pathogenic in mice.252,678,785 Interestingly, supplying sfRNA in trans restored WNV cytopathic effect in cell culture, suggesting that these small RNAs may have a specific but as yet unknown target.678
Membrane Reorganization and the Compartmentalization of Flavivirus Replication
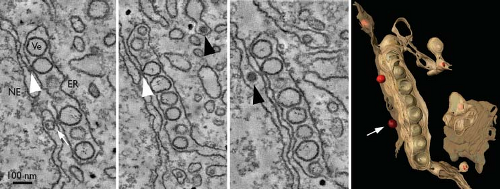
Biochemical studies of flavivirus-infected cells show that replicase activity is concentrated in dense membrane fractions that are enriched for most viral NS proteins. Treatment with nonionic detergents increases sensitivity to nucleases and proteases, indicating that the active replicase resides within a membrane-bound compartment. Consistent with this, ultrastructural changes in perinuclear membranes can be detected in flavivirus-infected cells.898 In general, the earliest event is the proliferation of ER membranes, followed by the appearance of smooth vesicular structures around the time of early logarithmic virus production. These structures, sometimes referred to as double-membrane vesicles or vesicle packets, are small clusters of ∼90-nm vesicles within the lumen of the ER.898,904 They are frequently adjacent to ER-derived convoluted membranes, which can appear as randomly folded membranes or highly ordered “paracrystalline arrays”.454,616,898 Electron tomography revealed that these vesicles are invaginations of the ER and retain connectivity to the cytosol through a neck-like pore, sometimes apposed to sites of virus assembly898 (Fig. 25.9).
Perinuclear vesicle packets have been confirmed as the sites of viral RNA synthesis by metabolic labeling of nascent RNA, in situ hybridization, and immunodetection of double-stranded RNA (dsRNA) (presumably RF and RI).
Perinuclear vesicle packets have been confirmed as the sites of viral RNA synthesis by metabolic labeling of nascent RNA, in situ hybridization, and immunodetection of double-stranded RNA (dsRNA) (presumably RF and RI).
The pathways of flavivirus-induced membrane reorganization are currently being worked out. One important clue comes from the ability of DENV-2 NS3 to recruit the cellular fatty acid synthase to sites of replication, at least by late times of infection.319 In addition, DENV induces autophagosome formation in infected cells,393,463,560,651 leading to turnover of triglycerides into free fatty acids, β-oxidation of lipids, and increased adenosine triphosphate (ATP) levels that promote virus replication.320
Assembly and Release of Particles from Flavivirus-Infected Cells
Similar to replication, early ultrastructural studies indicated that flavivirus morphogenesis occurs in close association with intracellular membranes.615 The assembly process is thought to commence by association of C protein dimers with genomic RNA, followed by budding into ER membranes containing the E–prM glycoprotein complex. Recent electron tomography studies of DENV-infected cells have shown that replicase-containing vesicle packets and ER-associated sites of virus budding are part of one continuous network. Pores within the replication vesicles appear to release newly synthesized viral RNA directly adjacent to budding DENV particles.898 These findings suggest a potential mechanism for coupling of flavivirus replication and assembly.397 Interdependence of these processes is a common theme among RNA viruses and might serve to reduce the propagation of defective genomes. Coupling flavivirus replication and assembly may also explain the roles of NS2A and NS3 proteins in infectious virus assembly.434,465,497,655
Following assembly, nascent virions are transported through the secretory pathway and released at the cell surface.536 It is during secretion that immature particles undergo acid-induced E–prM rearrangement and prM cleavage (Fig. 25.10). Additional virion maturation steps occur during egress, including glycan modification of prM and E (for some viruses) by trimming and terminal addition.123,169,309,547 This implies that virions move through an exocytosis pathway similar to that used for host cell surface glycoproteins. In addition to the secretory pathway machinery, other host factors have been implicated in flavivirus morphogenesis. Inhibitor and RNA interference studies implicate the Src family kinase c-Yes in WNV egress from the ER.333 A serine/threonine protein phosphatase inhibitor, I(2)(PP2A), has also been found to bind the WNV C protein, although its importance for infectivity was minimal.350
Hepatitis C Viruses
Background and Classification
HCV was identified in 1989 through expression cloning of immunoreactive cDNA from the plasma of a chimpanzee infected with an etiologic agent known to cause non-A non-B viral hepatitis.143 The virus currently infects more than 130 million people worldwide and remains a significant public health concern. HCV typically causes chronic hepatotropic infections, which can progress over decades to fibrosis, cirrhosis, and liver cancer. Based on difficulties in detecting HCV in vivo, the virus was originally thought to replicate poorly. Yet mathematical models of HCV dynamics indicate that a chronically infected patient produces approximately 1012 virions per day, with a virion half-life of only a few hours.666
Based on its evolutionary history, HCV is classified into seven genotypes, which differ from each other by more than 30% at the nucleotide level.614,788 Each genotype is further divided into numerous subtypes. HCV genotypes show differences in worldwide distribution, disease progression, and susceptibility to
treatment. In addition, intergenotypic and intersubtype recombinant HCV genomes have been described.788
treatment. In addition, intergenotypic and intersubtype recombinant HCV genomes have been described.788
While no HCV vaccine exists, advances in treatment have been made. Pegylated IFN-α and ribavirin—the standard of care for over a decade—have burdensome side effects and do not cure many of the patients treated. The most common HCV genotype, genotype 1, is also the most difficult to treat with this therapy.334 Recently, however, HCV-specific antiviral drugs—such as protease inhibitors—have begun to reach the clinic. While the new drugs must still be used in combination with IFN and ribavirin, first-generation protease inhibitors have almost doubled the chances of treatment success.
In addition to HCV, the genus Hepacivirus may contain a few related viruses that are awaiting taxonomic classification. An HCV-like virus was recently identified in domestic dogs with respiratory illness.388 This virus appears to be closely related to HCV and has been tentatively designated canine hepacivirus (CHV), although it has not been formally classified within this genus. In addition, GB virus B (see later) appears to be closely related to the hepaciviruses.
Experimental Systems
HCV research was limited for years by the lack of convenient laboratory culture systems and small animal models. However, many of these technical hurdles have been overcome in the past decade.
Shortly after the sequence of the HCV genome was fully elucidated, the first consensus cDNA clones were constructed. RNA transcripts from these cDNAs were shown to be infectious by direct intrahepatic inoculation into chimpanzees.412,925 Chimpanzee infectious clones have been used to show that all viral enzymes, the p7 gene, and 3′ NCR are essential for HCV replication in vivo.414,750,926 Furthermore, the ability to use genetically defined inocula to initiate clonal infections provided a useful tool to study virus evolution and immune responses.106 Despite their demonstrated utility in vivo, however, these infectious transcripts failed to replicate after transfection into cultured cells.
Subgenomic HCV replicons were developed in 1999 and proved to be the first broadly useful system for studying HCV RNA replication in cell culture.509 Subgenomic replicons are bicistronic constructs in which the HCV internal ribosome entry site (IRES) drives expression of the neomycin resistance gene and the encephalomyocarditis virus (EMCV) IRES drives expression of the HCV NS proteins. Following RNA transfection into a human hepatoma line, Huh-7, the original genotype 1b (Con1 strain) replicon replicated to low levels.509 Cell culture–adaptive mutations were later found to increase RNA replication efficiency up to 10,000 times.77 These mutations, many of which cluster in NS5A, are thought to increase RNA accumulation at the expense of infectious virus production, although their mechanisms of action are not well understood (reviewed in43). In addition to Con1, subgenomic replicons have now been constructed for additional genotype 1b strains, as well as genotype 1a and 2a isolates.
Full-length genotype 1a and 1b genomes bearing adaptive mutations were shown to replicate in cell culture but did not produce infectious virus particles.78,355,676 The HCV pseudoparticle (HCVpp) system was therefore developed to examine the role of the viral glycoproteins in entry. HCVpp are defective retrovirus particles expressing a reporter gene and displaying the HCV envelope proteins on their surface.46,201,343 Pseudoparticles undergo low-pH–mediated entry by using the known HCV entry factors and continue to be a valuable system to study entry in isolation from RNA replication. HCVpp differ from authentic particles, however, in their acid susceptibility343 and their assembly in a post-Golgi compartment rather than in the ER.751
In 2005, production of authentic HCV in cell culture (HCVcc) was achieved for the first time. An HCV isolate from an unusual case of acute fulminant hepatitis in Japan (JFH-1) was found to replicate in culture without the need for adaptive mutations.392 Remarkably, full-length JFH-1 also produced low levels of infectious virus.883,959 Infectious titers could be increased with passage or by engineering chimeric genomes based on a related genotype 2a strain, J6.487,675 A cell culture–adapted genotype 1a genome that produces infectious HCVcc has also been developed, albeit with lower infectivity.937
The chimpanzee remains the only animal model that reproduces clinical aspects of HCV infection, including a high rate of viral persistence and the development of HCV-specific innate and adaptive immune responses. Advances have been made, however, in generating small animal models for HCV infection. Immunodeficient mouse strains with liver injury can be repopulated with human hepatocytes and infected with HCV. Two commonly used liver injury models are the toxic urokinase plasminogen activator transgene (Alb-uPA mice)568,570 and a conditionally lethal fumaryl acetoacytate hydrolase deficiency (FAH−/− mice).73 Recently, immunocompetent mice that express human HCV entry factors have been developed as a model for the early steps of infection.198
Structure and Physical Properties of the Virion
HCV particles are enveloped and contain the viral proteins core (C), E1, and E2, as well as the genomic RNA. Virions are between 30 and 80 nm in diameter.88,317,949 For a virus of this size, HCV exhibits an unusually low and heterogeneous buoyant density. In highly infectious acute-phase chimpanzee serum, HCV-specific RNA is detected in fractions with densities ranging from 1.03 to 1.10 g/mL.87,331 The particle density is inversely correlated with infectivity, with low-density virions being more infectious than high-density particles. Similar to serum-derived virus, the peak of HCVcc RNA has a buoyant density of approximately 1.15 g/mL,116,487,883,937,959 whereas the peak infectivity is near 1.10 g/mL.116,487,959 Passage of cell culture–grown virus in animals488 or primary hepatocyte cultures682 increases the proportion of low-density, high-infectivity virus, highlighting the impact of host cell environment on virion composition.
The high density observed for noninfectious particles may come from immunoglobulin binding to the virus and/or represent nonenveloped nucleocapsids.23 The low buoyant density of infectious HCV appears to reflect the ability of the virus to interact with serum lipoproteins.22,631,692,855 The lipidome of HCV particles is very similar to low-density lipoproteins (LDLs) and very-low-density lipoproteins (VLDLs), including an enrichment of cholesteryl esters.569 Immunoprecipitation and immunogold-electron microscopy (EM) studies have detected apolipoprotein (Apo) AI,416 ApoB,416,631,855 ApoC1,572 and ApoE631 in association with serum-derived HCV. Apolipoprotein E and ApoC1 are also associated with HCVcc,126,569,572 but ApoB is generally lacking, leading to controversy about
its functional importance (reviewed in44). The propensity of HCV to interact closely with lipids has led to the hypothesis that at least a portion of circulating virus is internalized in host lipoproteins to form a “lipoviral” particle,22 which may provide camouflage for the virus to escape neutralizing antibodies.
its functional importance (reviewed in44). The propensity of HCV to interact closely with lipids has led to the hypothesis that at least a portion of circulating virus is internalized in host lipoproteins to form a “lipoviral” particle,22 which may provide camouflage for the virus to escape neutralizing antibodies.
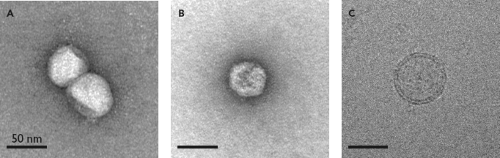
Although unambiguous visualization of HCV from patient samples remains a challenge, negative stain and cryoelectron microscopy studies have started to reveal structural features of HCV grown in cell culture264,569,883 (Fig. 25.11). One recent study detected two predominant classes of HCV particles, each making up approximately half of the preparation. The first were spherical, pleomorphic particles, ∼55 nm in diameter, containing a lipid bilayer and an electron-dense core (Fig. 25.11C). These particles, which are presumably infectious HCV, were enriched in low-density fractions and had a relatively smooth, featureless surface that reacted with E2- and ApoE-specific antibodies. The second class of particles, electron-dense structures ∼45 nm in diameter, is presumed to be naked nucleocapsids. This class was enriched in high-density, low-infectivity fractions and lacked a membrane or visible surface projections. Similar particles were seen after detergent treatment of infectious virus, which is expected to remove the viral envelope.264 Unfortunately, the reconstruction of high-resolution models of the virus particle is currently limited by particle heterogeneity, relatively low yields, and lack of symmetry in both classes of HCV particles.
Binding and Entry
HCV infects target hepatocytes through a multistep process that involves several cell surface molecules. Virus particles initially attach to glycosaminoglycans45,268 and possibly the LDL receptor (LDL-R),4,598 which may interact with virion-associated ApoB and ApoE. Following attachment, virus particles employ four specific entry factors to mediate uptake: scavenger receptor class B type I (SR-BI, aka SCARBI),758 CD81,679 claudin 1 (CLDN1),224 and occludin (OCLN).681
SR-BI, which is highly expressed on hepatocytes, plays important physiologic roles in the selective uptake of cholesterol and cholesteryl esters from lipoproteins. SR-BI can interact with the E2 hypervariable region 1 (HVR1),758 and its overexpression enhances virion binding to CHO cells,224 suggesting a role for SR-BI in primary attachment. Blocking antibody studies have both supported a role for SR-BI in binding,121 as well as shown the importance of SR-B1 at a postbinding step that is closely linked to CD81.386,952 Antibodies directed at E2 HVR1 similarly block a postattachment step of HCV entry.878 The physiologic SR-BI binding partner, PDZK1, is also important for entry.225 Perhaps because the lipoviral particle shares commonalities with the natural ligands of SR-BI, various lipoproteins modulate HCV infectivity. Specifically, high-density lipoproteins (HDLs) can enhance HCV fusion in a manner dependent on SR-BI lipid transfer, ApoCI, and the E2 HVR1.47,199,571,880 In contrast, oxidized LDL,882 VLDL,542 and serum amyloid A114,453 inhibit HCV entry.
The tetraspanin CD81 binds tightly to the glycoprotein E2 via residues in the CD81 large extracellular loop.328,670,679 Antibodies targeting CD81 do not affect HCVpp binding to hepatocytes166 but inhibit HCV infection after virus attachment.421 Because HVR1 masks the CD81 binding site, a conformational change may be required to prime E2 for binding to this co-receptor.37 CD81 interacts with another HCV entry factor, CLDN1,313,929 forming a complex that is essential for HCV infection.312 Receptor tyrosine kinases, such as epidermal growth factor receptor (EGFR), appear to regulate CD81–CLDN1 interactions and to promote HCV membrane fusion.527 Interaction of CD81 with another host factor, EWI-2wint, inhibits virus uptake.734 In addition to its role in entry, interaction of HCV with lymphocyte-expressed CD81 may contribute to the immune dysregulation seen in some patients with chronic infection.173,739,865
Two tight junction proteins, CLDN1224 and OCLN,59,496,681 are essential for postbinding steps of HCV uptake. In addition, CLDN family members-6 and −9 can also mediate entry.563 The involvement of tight junction proteins suggests that HCV could traffic across the surface of the hepatocyte in order to engage all of its entry factors. Indeed, binding to CD81 activates a Rho guanosine triphosphatase (GTPase), which may mediate cytoskeleton rearrangements. Furthermore, recombinant E2 promotes relocalization of CD81 to tight junctions.94 In conflict with this hypothesis, however, is the finding that CLDN1
can form complexes with CD81 on the basolateral surface of hepatocytes.312 In addition, single particle tracking has visualized HCV internalization outside tight junctions—although this may represent the nonpolarized nature of the cultured hepatoma cells.159
can form complexes with CD81 on the basolateral surface of hepatocytes.312 In addition, single particle tracking has visualized HCV internalization outside tight junctions—although this may represent the nonpolarized nature of the cultured hepatoma cells.159
Following attachment, HCV is taken up via clathrin-mediated endocytosis.76,564 Internalized virions co-localize with clathrin light chain and the E3 ubiquitin ligase c-Cbl prior to uncoating.159 Intracellular HCV is delivered to Rab5a-positive early endosomes,159 which likely provide the acidic environment necessary to induce rearrangement of the HCV glycoproteins into their fusogenic form.343,864 Surprisingly, extracellular HCVcc particles are resistant to inactivation by low pH, suggesting that interactions at or near the cell surface prime the virus for fusion.864 This is functionally similar to pestiviruses425 but in contrast to flavivirus particles, which are primed during maturation,946 and to HCVpp,343,644 which have structural and compositional differences from authentic HCV particles.
Genome Structure
The HCV genome is an uncapped, 9.6-kb RNA containing highly structured 5′ and 3′ ends (Fig. 25.12A). The 5′ NCR is a well-conserved, 341-nt sequence element that folds into a complex structure consisting of four major domains and a pseudoknot.324 The first 120 nt serves as a minimal replication element, although compelling genetic evidence shows that the entire 5′ NCR (or its complement in the negative strand) plays an important role in efficient positive-strand RNA synthesis.70,247,282,290,404,525,721 The 5′ NCR also contains an IRES, which directs the cap-independent translation of the single large ORF.
Interestingly, the liver-specific micro-RNA (miR)-122 binds to two sites within the HCV 5′ NCR, and these interactions are required for efficient viral replication.377,469 While miR-122 can stimulate IRES-mediated translation in an Argonaute 2-dependent manner,325,366,732,907 elegant genetic experiments revealed that a major function of miR-122 is to sequester the 5′ end of the genome, perhaps protecting it from RNA degradation or from activation of innate antiviral responses.532 Nuclease-resistant oligonucleotides that antagonize miR-122 function in vivo profoundly decrease HCV replication in experimentally infected chimpanzees and show promise for future therapeutics,447 although viral escape can occur in cell culture.474
A number of functionally important RNA structures have been identified within the HCV ORF.867 Genetic analysis indicated that an RNA stem-loop within the C gene is important for viral fitness.561,876 An RNA structure in NS5B, termed 5BSL3.2, forms a long-distance base-pairing interaction with upstream RNA sequences with the 3′ NCR.246 This “kissing” interaction is essential for replication, although its function remains to be determined. In addition, the HCV ORF contains fewer UA and UU dinucleotides than expected
by chance.308 This is significant because these dinucleotides are the preferred cleavage sites of the IFN-stimulated RNase L. Based on the large-scale computerized folding of numerous positive-strand RNA viral genomes, the HCV ORF is also predicted to contain an unusually high rate of internal base pairing.789 Intriguingly, this feature is common among viruses that cause persistent infections. Thus, the observed bias in nucleotide composition and propensity for internal base pairing within the HCV ORF may reflect adaptation to evolutionary pressure exerted by innate antiviral pathways.
by chance.308 This is significant because these dinucleotides are the preferred cleavage sites of the IFN-stimulated RNase L. Based on the large-scale computerized folding of numerous positive-strand RNA viral genomes, the HCV ORF is also predicted to contain an unusually high rate of internal base pairing.789 Intriguingly, this feature is common among viruses that cause persistent infections. Thus, the observed bias in nucleotide composition and propensity for internal base pairing within the HCV ORF may reflect adaptation to evolutionary pressure exerted by innate antiviral pathways.
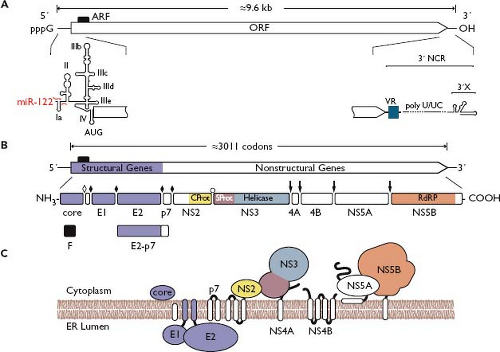 Figure 25.12. Hepacivirus genome structure and protein expression. A: Hepatitis C virus (HCV) genome structure and RNA elements. Important RNA elements are indicated as in Figure 25.4. The binding sites of micro-RNA (miR)-122 are indicated in red; the alternative reading frame (ARF) is indicated by a black box. B: Polyprotein processing and cleavage products. Cleavage sites are indicated as in Figure 25.4 except the NS2/3 cleavage, which is mediated by the NS2 cysteine autoprotease (open bullet). C: Polyprotein membrane topology. See the legend of Figure 25.4 for symbol definitions and the text for further details. |
The 3′ NCR was originally thought to terminate in polyadenosine or polyuridine. Improved methods for producing 3′ terminal cDNAs later revealed, however, that the HCV 3′ NCR actually consists of a short (∼40 nt) variable domain and a polyuridine/polypyrimidine (polyU/UC) tract, followed by a highly conserved 98-nt 3′ X domain.413,827 Mutagenesis studies revealed that the 3′ X domain is essential for RNA replication and that the polyU/UC tract must be at least 26 nt in length.245,926,935,941 A number of cellular factors have been found to bind to the HCV 3′ NCR, including RNA-binding proteins polypyrimidine-tract binding (PTB) protein,278,363,866 heterogeneous ribonuclear protein C,150,277 glyceraldehyde-3-phosphate dehydrogenase,671 HuR,801 and La autoantigen.799
Translation and Proteolytic Processing
In addition to its roles in replication, the HCV 5′ NCR acts as an IRES to direct cap-independent translation of the viral genome. The IRES is encoded by 5′ NCR domains II through IV (Fig. 25.12A), although flanking sequences can influence activity. In the current model of HCV IRES function, free 40S ribosomal subunits directly bind to the 5′ NCR domains IIId through f and adopt an mRNA-bound conformation.645,669,798 The IRES–40S complex then binds eIF3 and the ternary complex eIF2–GTP–Met-tRNAi to form a 48S intermediate complex, in which the initiating AUG codon at nt 342 is positioned within the ribosomal P site.367,645 The HCV IRES directly interacts with eIF3 via determinants in domain IIIb164 and thereby functionally mimics the 5′ cap-binding complex eIF4F.793 Following GTP hydrolysis and release of eIF2, the 60S ribosomal subunit is recruited to form a translationally active 80S complex.367,645,669 Under some conditions the HCV IRES does not require eIF3 or ternary complex and is also capable of directing translation from non-AUG initiation sites.444,846 This mechanism, which utilizes eIF5B to recruit Met-tRNAi, may allow the virus to initiate translation despite cellular eIF2 phosphorylation by the viral RNA sensor protein kinase R (PKR).
A number of cellular factors have been reported to contribute to HCV IRES activity. The human La protein stimulates translation by binding near the initiator AUG and recruiting the 40S ribosomal subunit168,693; inhibiting La strongly decreases HCV gene expression and replication.168,194 Other host factors that stimulate HCV IRES activity include proteosome α-subunit PSMA7428 and nucleolin.365 In addition, polycytidine binding proteins 1 and 2 and PTB were shown to bind the 5′ NCR, although their functions in the virus life cycle remain to be defined.43 IRES-mediated translation can be inhibited by long-distance base pairing of 5′ NCR nt 24 to 38 to a stem-loop region (nt 428 to 442) within the C gene region.405 Interestingly, this part of the 5′ NCR overlaps with one of the miR-122 binding sites.377
Translation of the HCV genome produces a large polyprotein (∼3,000 amino acids) that is proteolytically cleaved to produce 10 viral proteins (Fig. 25.12B): the virion structural components core (C) and glycoproteins E1 and E2, and the presumed NS proteins p7, NS2, NS3, NS4A, NS4B, NS5A, and NS5B. Mature forms of the HCV proteins arise via co- and posttranslational cleavage mediated by cellular and host-encoded proteases. During structural region processing, cleavages between C/E1, E1/E2, and E2/p7 are mediated by signal peptidase; mature C protein is released from the E1 signal peptide via signal peptide peptidase (SPP). Within the NS region, signal peptidase mediates cleavage of p7/NS2. The remainder of the NS region is processed by two virus-encoded proteases: the NS2 cysteine autoprotease mediates cleavage at the NS2/3 junction, whereas the NS3-4A serine protease cleaves at all downstream junctions (Fig. 25.12C).
In addition to proteins encoded by the large ORF, it appears that small protein products can be produced from the +1 reading frame of the C gene. At least three different forms of alternative reading frame protein (ARFP) have been described: ARFP/F (frameshift), ARFP/DF (double frameshift), or ARFP/S (short form). Although it is thought that these products are produced via ribosomal frameshifting, alternate translational initiation sites may also be involved.89 It is equally unclear what role, if any, ARFP expression plays in the virus life cycle. Mutant genomes with a disrupted +1 reading frame are highly attenuated in vivo,561,876 although this may be due to disruption of conserved RNA secondary structures within the C gene.
Features of the Structural Proteins
C Protein
Core (C) is the first protein encoded by the HCV genome. It is among the most conserved of the viral proteins and is thought to multimerize and bind RNA to form the viral nucleocapsid. Proteolytic processing of the C protein occurs in two steps. First, the C/E1 junction is cleaved by host signal peptidase to yield a 191–amino acid, membrane-anchored form of C and the N-terminus of E1.329 Subsequent cleavage within the C-terminal membrane anchor by SPP liberates the mature form of C559,640,752,933 and is essential for infectivity.832 The authentic SPP cleavage site has not yet been determined but likely occurs somewhere between residues 173 and 182351; recent genetic evidence suggests that the C-terminus is at position 177.419 In addition, N-terminally truncated forms of C have been described, but their mechanism of production is unknown.218
Following removal of the membrane anchor, the mature C protein consists of two domains.86 The N-terminal domain I (DI, ∼120 amino acids) is hydrophilic and can bind RNA nonspecifically,752 associate with the 5′ NCR,783,828 and mediate dimerization of the RNA 3′X region.171 In vitro, interaction of DI with structured RNA can induce the assembly of nucleocapsid-like particles.431 C protein likely dimerizes through homotypic binding elements in domains I and II,86,406,518,548 as well as a potential disulfide bond at Cys 128.432 C protein has also been shown to associate with E1 and E2.49,502,623 Domain I may also be the target of phosphorylation by protein kinases A and C.517
Domain II (∼50 amino acids) contains primarily hydrophobic residues. In vitro, this domain is important for folding of truncated recombinant protein (residues 1 to 169) into an
α-helix–rich dimer.86 DII is also important for localizing C to ER-associated lipid droplets,38,85,604 which play an important but unclear role in virus particle assembly.588,777 Lipid droplet association requires prior cleavage with SPP559 and is enhanced by the actions of diacylglycerol acyltransferase-1 (DGAT-1), an ER-resident enzyme that catalyzes a final step in triglyceride synthesis.326 It has been proposed that trafficking of C to lipid droplets increases the risk of liver steatosis in patients with chronic HCV, as well as promotes the development of steatosis in certain transgenic mice that overexpress the protein. Indeed, C binding to lipid droplets displaces adipocyte differentiation–related protein (ADRP) and leads to redistribution, clustering, and increased synthesis of these organelles in cell culture.84,190
α-helix–rich dimer.86 DII is also important for localizing C to ER-associated lipid droplets,38,85,604 which play an important but unclear role in virus particle assembly.588,777 Lipid droplet association requires prior cleavage with SPP559 and is enhanced by the actions of diacylglycerol acyltransferase-1 (DGAT-1), an ER-resident enzyme that catalyzes a final step in triglyceride synthesis.326 It has been proposed that trafficking of C to lipid droplets increases the risk of liver steatosis in patients with chronic HCV, as well as promotes the development of steatosis in certain transgenic mice that overexpress the protein. Indeed, C binding to lipid droplets displaces adipocyte differentiation–related protein (ADRP) and leads to redistribution, clustering, and increased synthesis of these organelles in cell culture.84,190
Envelope Glycoproteins
The HCV glycoproteins E1 (∼30 kD) and E2 (∼70 kD) form a noncovalent heterodimer that mediates viral attachment and membrane fusion. Both E1 and E2 are type I TM glycoproteins that contain a large extracellular domain and a single C-terminal TM domain. To generate this topology, the E1 and E2 TM segments, which each contain two short (<20 amino acids) hydrophobic stretches separated by charged residues, adopt an intramembrane hairpin structure during translocation. Following signal peptidase cleavage, the luminal C-termini reorient to face the cytoplasm, yielding tandem proteins each with a single membrane anchor.156 Following biogenesis, the glycoproteins are retained in the ER.203,207,740
Stay updated, free articles. Join our Telegram channel

Full access? Get Clinical Tree