7.1 OVERVIEW OF VIRUS GENOME REPLICATION
In this chapter we consider the fifth step of our generalized replication cycle: genome replication. The genome of the infecting virus is replicated so that viral transcription can be amplified and to provide copies of the genome for progeny virions.
Generally, DNA viruses copy their genomes directly to DNA, and RNA viruses copy their genomes directly to RNA. There are, however, some DNA viruses that replicate their genomes via an RNA intermediate and some RNA viruses that replicate their genomes via a DNA intermediate. The various replication modes of virus genomes are summarized in Figure 7.1.
Figure 7.1 Replication of virus genomes in the seven Baltimore Classes. (+) RNA and (+) DNA have the same sequence as the mRNA (except that in DNA thymine replaces uracil). (–) RNA and (–) DNA have the sequence complementary to the mRNA (except that in DNA thymine replaces uracil). (+) and (–) strands are not indicated for the dsDNA of the class I viruses because the genomes of most of these viruses have ORFs in both directions. (+) and (–) strands are indicated for the ssDNA of the class II viruses. Most of these viruses have either a (+) or a (–) strand genome. Some ssDNA viruses and some ssRNA viruses have ambisense genomes.
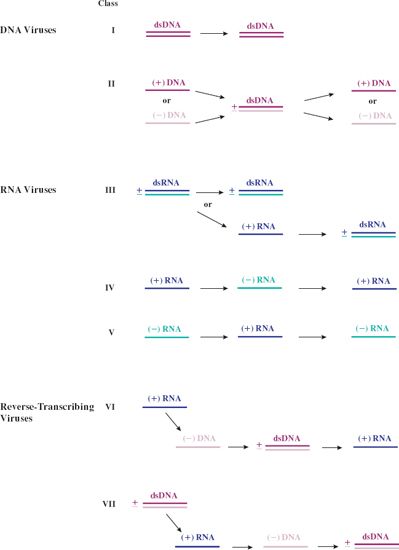
Single-stranded genomes are designated as plus or minus depending on their relationship to the virus mRNA. Plus-strand genomes have the same sequence as the mRNA (except that in DNA thymine replaces uracil), while minus-strand genomes have the sequence complementary to the mRNA. Single-stranded DNA is converted to dsDNA prior to copying.
There are two classes of viruses with (+) RNA genomes (Figure 7.1). Class IV viruses copy their (+) RNA genomes via a (–) RNA intermediate, while class VI viruses replicate via a DNA intermediate. The synthesis of DNA from an RNA template (reverse transcription) is also a characteristic of class VII viruses.
In this chapter we shall look at some general aspects of virus genome replication, and then we shall give individual attention to the DNA viruses, the RNA viruses and the reverse-transcribing viruses.
7.2 LOCATIONS OF VIRUS GENOME REPLICATION IN EUKARYOTIC CELLS
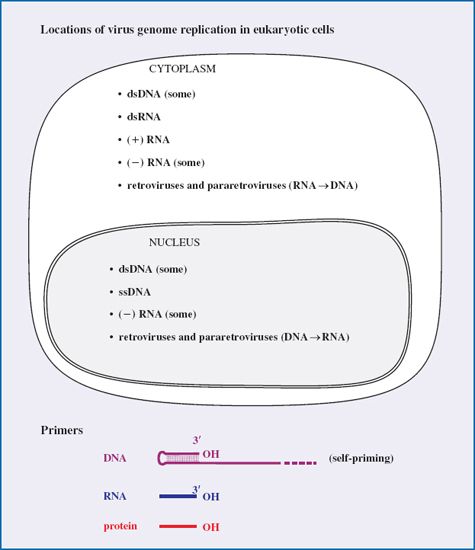
As we saw in Chapter 5, when viruses infect eukaryotic cells the genomes of some are delivered to the cytoplasm and some are conveyed to the nucleus. The destination of a virus genome, and hence the location in which it is replicated, varies with the type of genome (Table 7.1).
Table 7.1 Locations of virus genome replication in eukaryotic cells
| Virus genome | Cytoplasm | Nucleus |
| dsDNA (except pararetroviruses) | Some | Some |
| ssDNA | All | |
| dsRNA | All | |
| (+) RNA (except retroviruses) | All | |
| (−) RNA | Some | Some |
| (+) RNA (retroviruses)} | ||
| dsDNA (pararetroviruses) } | ssRNA → dsDNA | dsDNA → ssRNA |
The genomes of most DNA viruses are replicated in the nucleus, but those of some dsDNA viruses are replicated in the cytoplasm. The genomes of most RNA viruses are replicated in the cytoplasm, but there are exceptions amongst the minus-strand RNA viruses (Table 7.1). The retroviruses and pararetroviruses are special cases: each replicates RNA to DNA in the cytoplasm and DNA to RNA in the nucleus.
For those viral genomes that are replicated in the cytoplasm, nucleic acid synthesis usually takes place inside structures within the cytoplasm. For the dsRNA viruses, the retroviruses and the pararetroviruses these structures are composed of viral proteins; for the remaining plus-strand RNA viruses they are composed of cell-derived membranes. These viral and cell-derived structures are described in later chapters. One probable reason why viruses avoid replicating their nucleic acids free in the cytoplasm is to protect them from defense mechanisms of the host cells. Obviously there must be an adequate supply of nucleotides into these structures in order for nucleic acid synthesis to proceed.
7.3 INITIATION OF GENOME REPLICATION
The replication of a virus genome is initiated at a specific nucleotide sequence that is recognized by the proteins that initiate replication. For most linear genomes this sequence is at one of the ends of the nucleic acid.
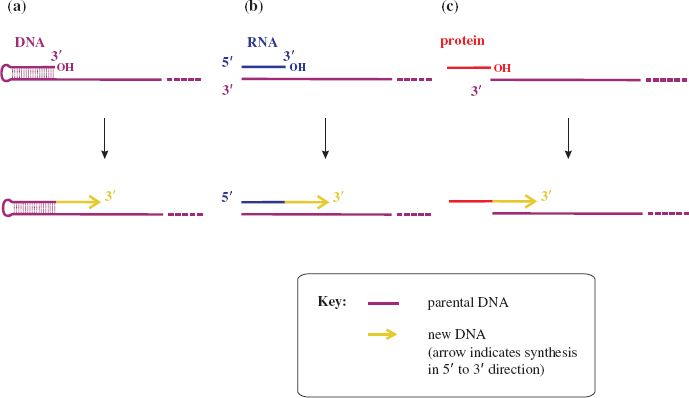
Replication of DNA, and of some viral RNAs, requires priming, which is the reaction of a nucleotide with an —OH group on a molecule at the initiation site. Initiation of genome replication for many RNA viruses, including rotaviruses and rhabdoviruses, does not require a primer; replication initiates when the first ribonucleotide of the new strand base pairs with a ribonucleotide in the template RNA. This first ribonucleotide effectively acts as a primer when its 3′ —OH group becomes linked to the second ribonucleotide.
Some ssDNA viruses, such as parvoviruses, use self-priming. At the 3′ end of the DNA there are regions with complementary sequences that can base pair (Figure 7.2(a)). The —OH group of the nucleotide at the 3′ end forms a linkage with the first nucleotide, then DNA synthesis proceeds to copy the whole genome.
Stay updated, free articles. Join our Telegram channel

Full access? Get Clinical Tree