29
Replication
CHAPTER CONTENTS
INTRODUCTION
The viral replication cycle is described in this chapter in two different ways. The first approach is a growth curve, which shows the amount of virus produced at different times after infection. The second is a stepwise description of the specific events within the cell during virus growth.
VIRAL GROWTH CURVE
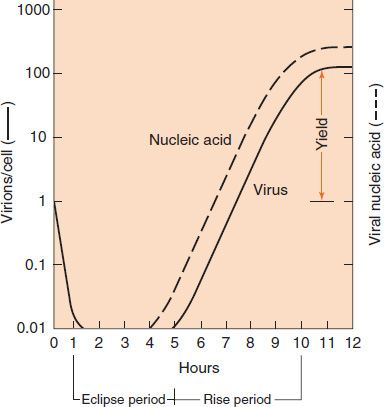
The growth curve depicted in Figure 29–1 shows that when one virion (one virus particle) infects a cell, it can replicate in approximately 10 hours to produce hundreds of virions within that cell. This remarkable amplification explains how viruses spread rapidly from cell to cell. Note that the time required for the growth cycle varies; it is minutes for some bacterial viruses and hours for some human viruses.
FIGURE 29–1 Viral growth curve. The figure shows that one infectious virus particle (virion) entering a cell at the time of infection results in more than 100 infectious virions 10 hours later, a remarkable increase. Note the eclipse period during which no infectious virus is detectable within the infected cells. In this growth curve, the amount of infecting virus is 1 virion/cell (i.e., 1 infectious unit/cell). (Modified and reproduced with permission from Joklik WK et al. Zinsser Microbiology. 20th ed. Originally published by Appleton & Lange. Copyright 1992 by McGraw-Hill.)
The first event shown in Figure 29–1 is quite striking: the virus disappears, as represented by the solid line dropping to the x axis. Although the virus particle, as such, is no longer present, the viral nucleic acid continues to function and begins to accumulate within the cell, as indicated by the dotted line. The time during which no virus is found inside the cell is known as the eclipse period. The eclipse period ends with the appearance of virus (solid line). The latent period, in contrast, is defined as the time from the onset of infection to the appearance of virus extracellularly. Note that infection begins with one virus particle and ends with several hundred virus particles having been produced; this type of reproduction is unique to viruses.
Alterations of cell morphology accompanied by marked derangement of cell function begin toward the end of the latent period. This cytopathic effect (CPE) culminates in the lysis and death of cells. CPE can be seen in the light microscope and, when observed, is an important initial step in the laboratory diagnosis of viral infection. Not all viruses cause CPE; some can replicate while causing little morphologic or functional change in the cell.
SPECIFIC EVENTS DURING THE GROWTH CYCLE
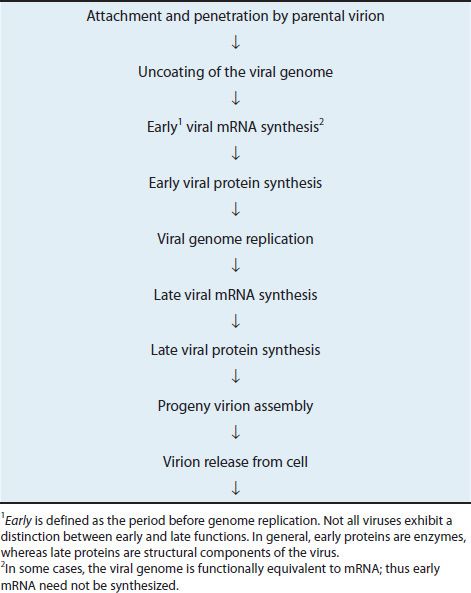
An overview of the events is described in Table 29–1 and presented in diagrammatic fashion in Figure 29–2. The infecting parental virus particle attaches to the cell membrane and then penetrates the host cell. The viral genome is “uncoated” by removing the capsid proteins, and the genome is free to function. Early mRNA and proteins are synthesized; the early proteins are enzymes used to replicate the viral genome. Late mRNA and proteins are then synthesized. These late proteins are the structural, capsid proteins. The progeny virions are assembled from the replicated genetic material, and newly made capsid proteins and are then released from the cell.
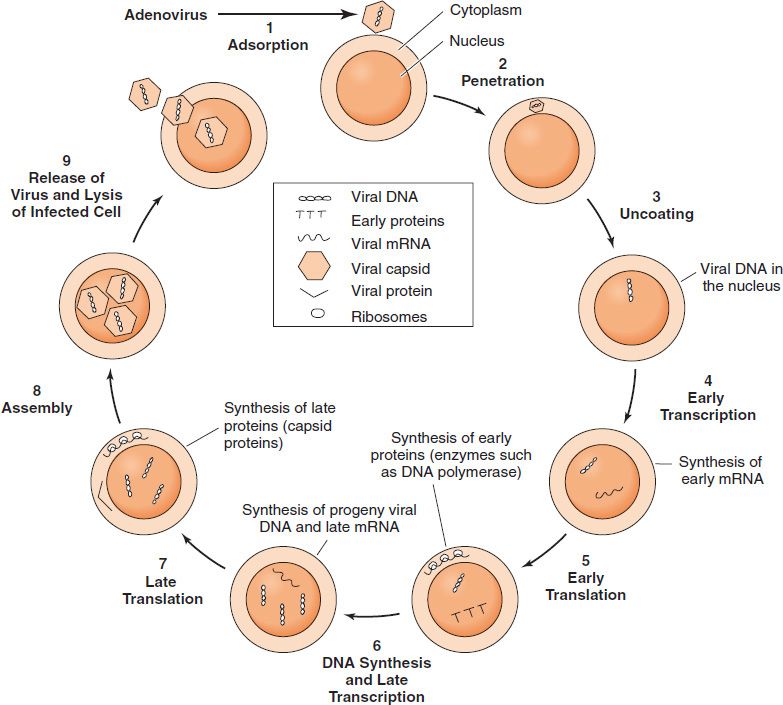
FIGURE 29–2 Viral growth cycle. The growth cycle of adenovirus, a nonenveloped DNA virus, is shown. (Modified and reproduced with permission from Jawetz E, Melnick JL, Adelberg EA. Review of Medical Microbiology. 16th ed. Originally published by Appleton & Lange. Copyright 1984 by McGraw-Hill.)
Another, more general way to describe the growth cycle is as follows: (1) early events (i.e., attachment, penetration, and uncoating); (2) middle events (i.e., gene expression and genome replication); and (3) late events (i.e., assembly and release). With this sequence in mind, each stage will be described in more detail.
Attachment, Penetration, & Uncoating
The proteins on the surface of the virion attach to specific receptor proteins on the cell surface through weak, noncovalent bonding. The specificity of attachment determines the host range of the virus. Some viruses have a narrow range, whereas others have quite a broad range. For example, poliovirus can enter the cells of only humans and other primates, whereas rabies virus can enter all mammalian cells. The organ specificity of viruses is governed by receptor interaction as well. Those cellular receptors that have been identified are surface proteins that serve various other functions. For example, herpes simplex virus type 1 attaches to the fibroblast growth factor receptor, rabies virus to the acetylcholine receptor, and human immunodeficiency virus (HIV) to the CD4 protein on helper T lymphocytes.
The virus particle penetrates by being engulfed in a pinocytotic vesicle, within which the process of uncoating begins. A low pH within the vesicle favors uncoating. Rupture of the vesicle or fusion of the outer layer of virus with the vesicle membrane deposits the inner core of the virus into the cytoplasm.
The receptors for viruses on the cell surface are proteins that have other functions in the life of the cell. Probably the best known is the CD4 protein that serves as one of the receptors for HIV but whose normal function is the binding of class 2 major histocompatibility complex (MHC) proteins involved in the activation of helper T cells. A few other examples will serve to illustrate the point: rabies virus binds to the acetylcholine receptor, Epstein–Barr virus binds to a complement receptor, and vaccinia virus binds to the receptor for epidermal growth factor,
Certain bacterial viruses (bacteriophages) have a special mechanism for entering bacteria that has no counterpart in either human viruses or those of animals or plants. Some of the T group of bacteriophages infect Escherichia coli by attaching several tail fibers to the cell surface and then using lysozyme from the tail to degrade a portion of the cell wall. At this point, the tail sheath contracts, driving the tip of the core through the cell wall. The viral DNA then enters the cell through the tail core, whereas the capsid proteins remain outside.
It is appropriate at this point to describe the phenomenon of infectious nucleic acid, because it provides a transition between the concepts of host specificity described earlier and early genome functioning, which is discussed later. Note that we are discussing whether the purified genome is infectious. All viruses are “infectious” in a person or in cell culture, but not all purified genomes are infectious.
Infectious nucleic acid is purified viral DNA or RNA (without any protein) that can carry out the entire viral growth cycle and result in the production of complete virus particles. This is interesting from three points of view:
(1) The observation that purified nucleic acid is infectious is the definitive proof that nucleic acid, not protein, is the genetic material.
(2) Infectious nucleic acid can bypass the host range specificity provided by the viral protein–cell receptor interaction. For example, although intact poliovirus can grow only in primate cells, purified poliovirus RNA can enter nonprimate cells, go through its usual growth cycle, and produce normal poliovirus. The poliovirus produced in the nonprimate cells can infect only primate cells because it now has its capsid proteins. These observations indicate that the internal functions of the nonprimate cells are capable of supporting viral growth once entry has occurred.
(3) Only certain viruses yield infectious nucleic acid. The reason for this is discussed later. Note that all viruses are infectious, but not all purified viral DNAs or RNAs (genomes) are infectious.
Gene Expression & Genome Replication
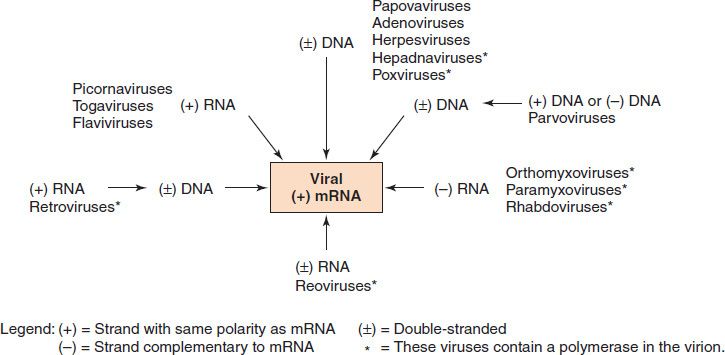
The first step in viral gene expression is mRNA synthesis. It is at this point that viruses follow different pathways depending on the nature of their nucleic acid and the part of the cell in which they replicate (Figure 29–3).
FIGURE 29–3 Synthesis of viral mRNA by medically important viruses. The following information starts at the top of the figure and moves clockwise: Viruses with a double-stranded DNA genome (e.g., papovaviruses such as human papillomavirus) use host cell RNA polymerase to synthesize viral mRNA. Note that hepadnaviruses (e.g., hepatitis B virus) contain a virion DNA polymerase that synthesizes the missing portion of the DNA genome, but the viral mRNA is synthesized by host cell RNA polymerase. Parvoviruses use host cell DNA polymerase to synthesize viral double-stranded DNA and host cell RNA polymerase to synthesize viral mRNA. Viruses with a single-stranded, negative-polarity RNA genome (e.g., orthomyxoviruses such as influenza virus) use a virion RNA polymerase to synthesize viral mRNA. Viruses with a double-stranded RNA genome (e.g., reoviruses) use a virion RNA polymerase to synthesize viral mRNA. Some viruses with a single-stranded, positive-polarity RNA genome (e.g., retroviruses) use a virion DNA polymerase to synthesize a DNA copy of the RNA genome but a host cell RNA polymerase to synthesize the viral mRNA. Some viruses with a single-stranded, positive-polarity RNA genome (e.g., picornaviruses) use the virion genome RNA itself as their mRNA. (Modified and reproduced with permission from Ryan K et al. Sherris Medical Microbiology. 3rd ed. Originally published by Appleton & Lange. Copyright 1994 by McGraw-Hill.)
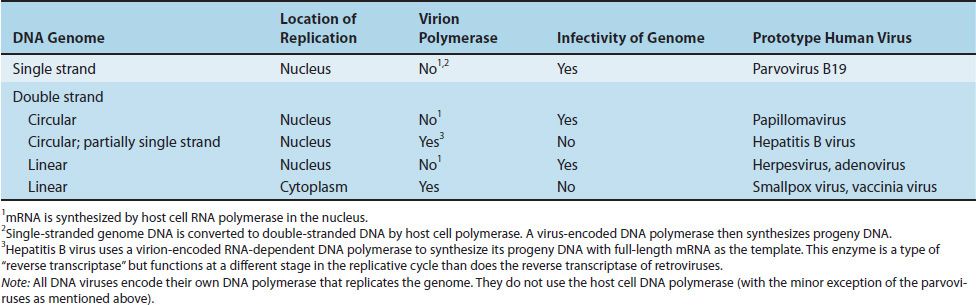
DNA viruses, with one exception, replicate in the nucleus and use the host cell DNA-dependent RNA polymerase to synthesize their mRNA. The poxviruses are the exception because they replicate in the cytoplasm, where they do not have access to the host cell RNA polymerase. They therefore carry their own polymerase within the virus particle. The genome of all DNA viruses consists of double-stranded DNA, except for the parvoviruses, which have a single-stranded DNA genome (Table 29–2).
Most RNA viruses undergo their entire replicative cycle in the cytoplasm. The two principal exceptions are retroviruses and influenza viruses, both of which have an important replicative step in the nucleus. Retroviruses integrate a DNA copy of their genome into the host cell DNA, and influenza viruses synthesize their progeny genomes in the nucleus. In addition, the mRNA of hepatitis delta virus is also synthesized in the nucleus of hepatocytes.
The genome of all RNA viruses consists of single-stranded RNA, except for members of the reovirus family, which have a double-stranded RNA genome. Rotavirus is the important human pathogen in the reovirus family.
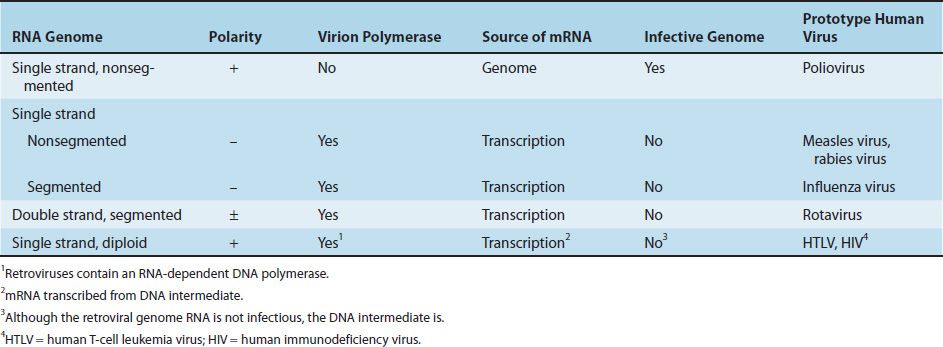
RNA viruses fall into four groups with quite different strategies for synthesizing mRNA (Table 29–3).
(1) The simplest strategy is illustrated by poliovirus, which has single-stranded RNA of positive polarity1 as its genetic material. These viruses use their RNA genome directly as mRNA.
(2) The second group has single-stranded RNA of negative polarity as its genetic material. An mRNA must be transcribed by using the negative strand as a template. Because the cell does not have an RNA polymerase capable of using RNA as a template, the virus carries its own RNA-dependent RNA polymerase. There are two subcategories of negative-polarity RNA viruses: those that have a single piece of RNA (e.g., measles virus [a paramyxovirus] or rabies virus [a rhabdovirus]) and those that have multiple pieces of RNA (e.g., influenza virus [a myxovirus]).
Certain viruses, such as arenaviruses and some bunyaviruses, have a segmented RNA genome, most of which is negative stranded, but there are some positive strand regions as well. RNA segments that contain both positive polarity and negative polarity regions are called “ambisense.”
(3) The third group has double-stranded RNA as its genetic material. Because the cell has no enzyme capable of transcribing this RNA into mRNA, the virus carries its own polymerase. Note that plus strand in double-stranded RNA cannot be used as mRNA because it is hydrogen-bonded to the negative strand. Rotavirus, an important cause of diarrhea in children, has 11 segments of double-stranded RNA.
(4) The fourth group, exemplified by retroviruses, has single-stranded RNA of positive polarity that is transcribed into double-stranded DNA by the RNA-dependent DNA polymerase (reverse transcriptase) carried by the virus. This DNA copy is then transcribed into viral mRNA by the regular host cell RNA polymerase (polymerase II). Retroviruses are the only family of viruses that are diploid (i.e., that have two copies of their genome RNA).
These differences explain why some viruses yield infectious nucleic acid and others do not. Viruses that do not require a polymerase in the virion can produce infectious DNA or RNA. By contrast, viruses such as the poxviruses, the negative-stranded RNA viruses, the double-stranded RNA viruses, and the retroviruses, which require a virion polymerase, cannot yield infectious nucleic acid. Several additional features of viral mRNA are described in the “Viral mRNA” box.
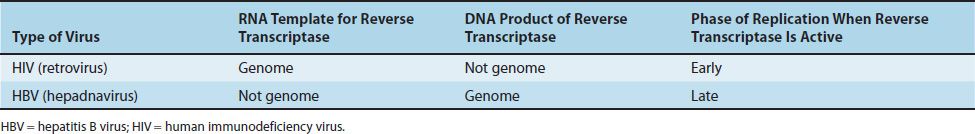
Note that two families of viruses utilize a reverse transcriptase (an RNA-dependent DNA polymerase) during their replicative cycle, but the purpose of the enzyme during the cycle is different. As described in Table 29–4, retroviruses, such as HIV, use their genome RNA as the template to synthesize a DNA intermediate early in the replicative cycle. However, hepadnaviruses, such as hepatitis B virus (HBV), use an RNA intermediate as the template to produce their DNA genome late in the replicative cycle.
TABLE 29–4 Comparison of Reverse Transcriptase Activity of HIV (Retroviruses) and HBV (Hepadnaviruses)
Stay updated, free articles. Join our Telegram channel

Full access? Get Clinical Tree