Parvoviridae
Kenneth I. Berns
Colin R. Parrish
Introduction and History
Parvoviruses are small, nonenveloped viruses with a linear, single-strand DNA (ssDNA) genome of about 5,000 bases. The family Parvoviridae (Table 57.1) contains two subfamilies: Parvovirinae and Densovirinae. The latter infect invertebrates and will not be described in detail in this chapter. (The interested reader is referred to the latest report of the International Committee on Taxonomy of Viruses [234]). The Parvovirinae are divided into five genera: Amdovirus, Bocavirus, Dependovirus, Erythrovirus, and Parvovirus. The adeno-associated viruses (AAVs) among the dependoviruses require co-infection with a helper virus for productive infection, whereas the others are all autonomous, although they require the host cell to go through S phase for viral DNA replication but do not induce cell division.234 The AAV helper virus is most commonly an adenovirus, or less frequently a herpesvirus.
In the first half of the 20th century, feline diseases characterized by enteritis, panleukopenia, and cerebellar ataxia were recognized as being of viral origin. In the late 1940s, a similar disease was noted in mink; in 1952, it was determined that those diseases were caused by similar small, DNA-containing viruses.399,431 Parvoviruses were isolated from rats (Kilham rat virus),233 and the H-1 virus isolated from human tumor tissue infected hamsters.430 Between 1960 and 1962, small particles were observed within Adenovirus preparations by electron microscopy, and the smaller virus (AAV) depended on co-infection of the cells with adenovirus for replication24,196,335 or on herpesviruses.68 The human parvovirus B19 was identified in 1974 during screening of blood, and the name derives from the code for that sample. The B19 virus is clearly associated with transient aplastic crisis (TAC) in patients with sickle cell disease, as well as with the childhood fifth disease and with postinfection arthralgia. In 1978, a new virus of dogs (canine parvovirus [CPV]) caused myocarditis of neonatal puppies and enteritis in older dogs, and CPV proved to be a variant of the long-known feline panleukopenia virus (FPV).340 Over the years, many different parvoviruses have been isolated from animals and cell cultures, and by discovery of viral DNA using direct cloning or sequencing approaches.223,224,395 Many of the latter viruses grow poorly or not at all in cultured cells, or are not clearly associated with disease in animals.
Biology of the Parvoviruses
The parvoviruses have small (∼25 nm diameter) and structurally stable capsids that bind host cell receptors, small ssDNA genomes with few genes that control their interactions with their hosts, and they replicate only in dividing cells or in the presence of a helper virus. Parvoviruses are widespread, and the presence of integrated viral DNA in the genomes of many different vertebrates and invertebrates indicates that related viruses have likely been infecting animals for millions of years.41,222,226,267 The capsids transmit between hosts by routes that include fecal–oral, urine, and respiratory spread, and they are stable in the environment and are readily transmitted by contaminated fomites. The routes of entry into the body likely involve infection of dividing epithelial or lymphoid cells of the upper respiratory tract, oropharynx, or intestine. The viruses are relatively simple and do not induce the host cell to proliferate or manipulate the immune responses directly through viral gene products. Where
the mechanisms have been defined, the host and tissue tropisms are determined by the requirement for dividing cells of the autonomous parvoviruses, by host receptor binding, or by the cell-specific control of viral gene expression.
the mechanisms have been defined, the host and tissue tropisms are determined by the requirement for dividing cells of the autonomous parvoviruses, by host receptor binding, or by the cell-specific control of viral gene expression.
Table 57.1 Classification of Parvoviridae | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
AAVs are cryptic viruses and are not clearly associated with any pathology. Natural hosts likely include most vertebrates, and the viruses are widespread and have been isolated from humans and other primates, horses, birds, cows, and sheep (see Table 57.1). Viral persistence is likely related to the ability of the DNA to integrate and establish a latent infection, and AAV sequences are found in many tissues.163,365 If AAV infects a healthy cell in culture in the absence of a helper virus, the viral genome does not replicate but may establish a persistent infection by chromosomal integration, and stable concatemers of AAV genomes in an extrachromosomal state are frequently found. In cell culture, the integrated AAV genome can be rescued by superinfection of the cells with a helper virus.197 Extrachromosomal or integrated latency are likely functionally equivalent if the infected cell does not divide; however, in dividing cells, the nonreplicating extrachromosomal AAV DNA would likely be diluted out. Stressing host cells allows some limited AAV replication. In cell culture, this can be achieved by exposure of the cell to genotoxic conditions that include ultraviolet irradiation, ionizing radiation, and cycloheximide.386,470
There are complex two-way relationships between AAVs and their helper viruses. AAVs can inhibit the replication of adenovirus in a process that depends on the relative amounts of the two viruses and on the temporal relationship of the co-infection.81,82 Adenovirus inhibition likely results from AAV Rep protein inhibition of heterologous promoters, and that inhibition is also seen in co-infections of AAV and SV40, papillomavirus, and some herpesviruses.192,238,254,473
Taxonomy and Classification
The Family Parvoviridae has unenveloped capsids of about 25 nm diameter with icosahedral symmetry composed of 60 copies of the capsid proteins. The nucleic acid is single-stranded linear DNA between 4 and 6 kb in length. The Parvovirinae infect vertebrates, and the Densovirinae infect insects and other invertebrates. The Parvovirinae are subdivided into five genera: Amdovirus, Bocavirus, Dependovirus, Erythrovirus, and Parvovirus. The Densovirinae are subdivided into four genera: Brevidensovirus, Densovirus, Iteravirus, and Pefudensovirus (see Table 57.1). Genera of the Parvovirinae are defined by their genomic and biological properties, whereas species are defined by their DNA sequence relatedness, biological host ranges, and/or capsid antigenicity.
There is significant variation in the details of the viral genomes and gene expression. Parvoviruses may package mostly negative-strand DNA, or both DNA strands in variable proportions, and the ssDNA genome has different hairpin structures at each end. There are generally two messenger RNA (mRNA) promoters and a single polyadenylation site near the 3′ end. The erythroviruses are related to the B19 human parvovirus, and mature virions contain equivalent proportions of positive and negative sense ssDNA, approximately 5 kb in size. The DNA molecules contain inverted terminal repeats (ITRs), and there is a single transcriptional promoter as well as two polyadenylation signals. Amdoviruses are related to the Aleutian mink disease virus (AMDV), whereas the bocaviruses are related to bovine parvovirus and the canine minute virus and include many viruses of humans and other hosts. Dependovirus virions package equivalent numbers of positive- or negative-strand ssDNA genomes, with ITRs of approximately 145 nucleotides—of which the first nearly 125 nucleotides form a palindromic sequence.
Structure of Capsid and General Properties
The particle has a molecular weight (MW) of 5.5 to 6.2 × 106 daltons. The buoyant density of the intact virion in cesium chloride (CsCl) is 1.39 to 1.42 g/cm3,424 and the sedimentation coefficient of the virion in neutral sucrose gradients is 110 to 122. The virions are resistant to inactivation, being largely stable for 60 minutes between pH 3 and 9 and at 56°C. The viruses can be inactivated by formalin, β-propiolactone, hydroxylamine, and oxidizing agents.
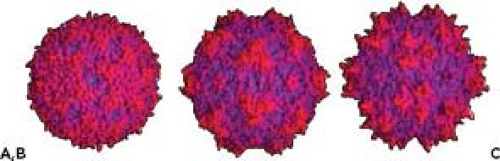
The capsids have T = 1 icosahedral structures assembled from 60 copies of between 2 and 4 forms of a single structural protein (virus protein [VP]: VP1, VP2, and VP3), which encapsidate the linear, ssDNA genome435 (Fig. 57.1). The atomic
structures of the capsids of many parvoviruses and AAVs share a similar structure5,229,402,435,447,469 (see Fig. 57.1). An eight-stranded, antiparallel β-barrel makes up one-third of the sequence of the major structural protein, and large loops connecting the strands of the β-barrel make up the capsid surface and determine host and tissue interactions, bind receptors, and form the epitopes recognized by antibodies. Antiparallel β-ribbons form a cylinder about the icosahedral fivefold axes and make a channel through the capsid in some of the viruses.118,150,229 N-terminal sequences of some VP2 molecules are externalized through that channel, allowing them to be cleaved to VP3, and DNA packaging occurs through that channel, leaving sequences from the 5′ end of the genome outside the capsid.118,123,350 Other features of the capsid may include one large or three smaller spike-like protrusions surrounding the icosahedral threefold axes of symmetry, depressions about the fivefold axes, and dimple-like depressions at the icosahedral twofold axes87,435,469 (Fig. 57.1).
structures of the capsids of many parvoviruses and AAVs share a similar structure5,229,402,435,447,469 (see Fig. 57.1). An eight-stranded, antiparallel β-barrel makes up one-third of the sequence of the major structural protein, and large loops connecting the strands of the β-barrel make up the capsid surface and determine host and tissue interactions, bind receptors, and form the epitopes recognized by antibodies. Antiparallel β-ribbons form a cylinder about the icosahedral fivefold axes and make a channel through the capsid in some of the viruses.118,150,229 N-terminal sequences of some VP2 molecules are externalized through that channel, allowing them to be cleaved to VP3, and DNA packaging occurs through that channel, leaving sequences from the 5′ end of the genome outside the capsid.118,123,350 Other features of the capsid may include one large or three smaller spike-like protrusions surrounding the icosahedral threefold axes of symmetry, depressions about the fivefold axes, and dimple-like depressions at the icosahedral twofold axes87,435,469 (Fig. 57.1).
Conformation-dependent neutralizing epitopes are important targets of neutralizing antibodies on the exposed surface of CPV, minute virus of mice (MVM), AAV, and B19 capsids, and the raised regions around the threefold axes are favored binding sites.272,381,415,460,477 Linear epitopes include the exposed N-terminus of VP2 of parvovirus full capsids and the N-terminus of VP1 for the B19 parvovirus.113,141,257,307,381
Functions of the Capsid Proteins
Most autonomous parvovirus and AAV capsids assemble from combinations of three overlapping proteins—MWs of 80,000 to 86,000 (VP1), 64,000 to 75,000 (VP2), and 60,000 to 62,000 (VP3)—although AMDV, B19, and simian parvovirus assemble from only VP1 and VP2.424 For the autonomous viruses, VP1 and VP2 are derived from alternatively spliced viral messages, and VP3 is generated in DNA-containing capsids by proteolytic cleavage of VP2. AAVs have three coat proteins: VP1 (∼87,000), VP2 (∼73,000), and VP3 (∼62,000). The capsid proteins are not glycosylated but are variably phosphorylated.
The unique regions of VP1 of most or all parvoviruses contain a calcium-dependent phospholipase A2 (PLA2) enzymatic activity that is buried within the capsid but becomes exposed during cell entry.151,418,480 The VP1 unique sequence is exposed on the surface of B19 capsids, which may reflect a difference in the biology of that virus.142 The PLA2 activities of the different viruses differ significantly, with the highest activity seen for porcine parvovirus (PPV).71 The N-terminus of VP1 contains basic amino acid motifs that likely function as nuclear localization motifs during cell entry and capsid assembly.199,270,444,459
Genome Structures and Organizations
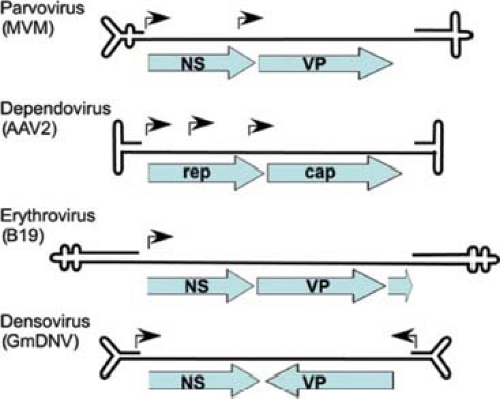
All parvoviruses of vertebrates have similar genome structures, with terminal repeats required for DNA replication, nonstructural protein genes on the left half of the genome (the 3′ end of the –ve strand DNA), and capsid protein genes on the right half of the genome (Fig. 57.2). Smaller proteins are produced by alternative splicing of some viruses and include the NS2 protein in the viruses similar to MVM and the NP1 protein in the bocaviruses (see later discussion). The functions of the noncoding sequences between the capsid protein and the 5′ terminal hairpin of the autonomous parvoviruses are less well understood; however, in the B19 viruses, there is a short open reading frame (ORF) encoding a protein (p11), and that region may also be involved in regulating the packaging of the
viral DNA.118,119 The smaller ORFs include the NS2 protein in several viruses, the 11KDa protein in the B19 human parvovirus, and a small ORF (SAT) that overlaps the VP1 and VP2 N-terminal sequence in an alternative reading frame.481 Various alternative splice products have been identified for many viruses, although the predicted proteins produced have generally not been characterized.
viral DNA.118,119 The smaller ORFs include the NS2 protein in several viruses, the 11KDa protein in the B19 human parvovirus, and a small ORF (SAT) that overlaps the VP1 and VP2 N-terminal sequence in an alternative reading frame.481 Various alternative splice products have been identified for many viruses, although the predicted proteins produced have generally not been characterized.
Epidemiology and Antiviral Immunity
Many autonomous parvoviruses, including CPV, FPV, porcine parvovirus (PPV), and B19 human parvovirus, cause acute infections of their hosts that last for fewer than 10 days. The virus is cleared by the host immunity, and infectious virus is not subsequently shed.336 However, some parvoviruses show prolonged replication and persistence: Rodent parvoviruses may persist in the kidneys and be shed in the urine, whereas AMDV persists in mink and continues to replicate for the life of the animal.10,52,354 For B19 in humans, rare chronic persistent infections occur in individuals who are immunosuppressed, or who do not develop effective immunity.63,158,478
The mechanisms of transmission among the parvoviruses vary and in many cases are not well defined. Carnivore parvoviruses such as CPV and FPV are spread by fecal–oral transmission.283,284,336 Some rodent parvoviruses also replicate in the intestine but may also be transmitted through urine after replication in the kidney.28,214 The human B19 virus replicates primarily in the bone marrow and is thought to be transmitted by respiratory routes.61
Cell-mediated immunity likely assists in recovery from infection; however, humoral immunity alone can protect animals against infection and is also important for recovery from infection, as immune immunoglobulin G (IgG) can arrest CPV replication in dogs and can terminate chronic human infections by the B19 parvovirus.61,63 Antibodies produced in older mink reduce titers of AMDV but do not eliminate the virus, thus a persistent infection continues.9,12,52
Any role for antigenic variation in the epidemiology of the parvoviruses is not well understood. Viruses of different genera or species are distinguished by polyclonal sera; however, little antigenic variation is present within virus species, and any epidemiologic significance is not known, and the same changes often alter host range or other properties of the virus.85,287,310,333,338 For MVM, antigenic variants were readily selected with neutralizing monoclonal antibodies in tissue culture or in persistent infections of SCID mice,272 although it is not known whether similar variation occurs in nature.
Genetic Variation and Evolution
These DNA viruses are replicated by host cell DNA polymerases; however, the single-stranded nature of the genome may make it susceptible to modification, and the mode of DNA replication may lead to significant sequence variation. The temporal rate of sequence variation is high (>4 × 10−4 variations per site per year) when measured over defined periods or during the growth of CPV through serial passage in tissue culture.25,276,393,394 MVM mutant strains grown persistently in mice or under monoclonal antibody selection showed variation at several sites owing to host or immune selection.272,275
All parvoviruses are related through a distant common ancestor, and the sequences can be subdivided into several clades that show some correlations with the hosts of origin,280 although there are many variants. Parvovirus-related sequences are integrated into the genomes of several animals.41,222,226,267 Most were only partial or degenerate sequences, which would not generate infectious virus or recombine with infectious virus sequences. The finding of some syntenic sequences in related mammals shows that the integration occurred prior to the host evolutionary divergence, thus at least some of the integrations occurred millions of years ago.
Various different viruses have been seen to infect single host species. For example, within cows, three parvovirus sequences have been discovered by searching for nonhost DNAs.14 The several erythroviruses from primates (B19 and related human and simian parvoviruses) are most closely related to each other and to the chipmunk parvovirus; however, those are distinct from other human parvoviruses.109,198,391,475 Most viruses from rats, mice, and hamsters were found to be within the same clade along with CPV and the related viruses of carnivores and PPV,27,448 whereas AMDV appears to infect mink and grey foxes and is only distantly related to other parvoviruses of vertebrates. Recombination is quite common among the parvoviruses.280,392
The human B19-like parvoviruses include three distinct clades that differ by 5% to 20% in sequence; within each clade, the viruses differ by less than 1% to 4%.72,162,198,391 Viruses from patients with persistent infections may show more variation than viruses from patients with acute infections. Many parvovirus strains have worldwide distributions, although some geographic clustering occurs.
The emergence of CPV illustrates specific features of the parvoviral evolution. The most recent common ancestor to all CPVs likely arose during the early 1970s.394,434 The CPV lineage included two major variants, and related viruses recovered from raccoons and other hosts suggested extensive exchange among the related hosts.16 Virus strains differed in various combinations of the two neutralizing epitopes, which were in many cases associated with capsid changes that influence receptor binding and host range.16,69,338,339
Significant variation (of up to 16%) was seen in the genomes of AMDV isolates, most likely owing to the co-circulation and co-infection of animals with variant virus strains. Because that virus can form persistent infections with continuously circulating virus, mixed infections likely occur.322 A hypervariable sequence within the capsid protein gene alters residues on the surface of the VP2 protein structure.51,291,321
Infection Pathways
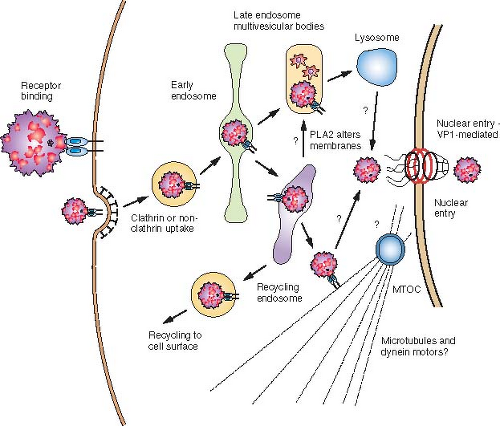
Parvoviruses bind to one or more cell surface receptors followed by endosomal trafficking within the cell within vesicles, release into the cytoplasm, and active or passive nuclear entry. There appear to be no substantial differences between the infections by the autonomous viruses and AAV (Fig. 57.3).
Capsid Structures and Cell Infection
The parvovirus capsid is a stable sphere of about 26 nm in diameter (see Fig. 57.1). Flexibility is required during infection to allow genome release and to expose protein structures from within the capsid. The VP1 unique region of 40 to 230 residues is not required for capsid formation or DNA packaging,439,444 and that sequence includes a basic amino acid motif, which likely acts as a nuclear localization sequence, and a PLA2, which is active when released from the capsid.71,270,418,444,480 The
unique region of PPV showed approximately 100-fold higher PLA2 activity than those of AAV2 or the human B19 parvovirus.71 The PLA2 modification of the lipid components of the endosome may allow more efficient viral release.151
unique region of PPV showed approximately 100-fold higher PLA2 activity than those of AAV2 or the human B19 parvovirus.71 The PLA2 modification of the lipid components of the endosome may allow more efficient viral release.151
The viruses of vertebrates show various surface features, including raised regions around the threefold axis of symmetry, a depressed region spanning the twofold axis, and in many cases a pore at the fivefold axis of symmetry that runs through the capsid6,402,403,435 (see Fig. 57.1). The insect parvoviruses show a relatively smooth surface.228,402 The host ranges and tissue tropisms of CPV and FPV, and of MVM variants, are controlled by a small number of residues on the capsid surface.4,6,29,85,436,469
Variability and flexibility in the capsid structure are important for viral infectivity and capsid functions—in particular, the externalization of VP1 and VP2 N-terminal sequences. Flexibility of surface loops was seen in CPV, FPV, and MVM capsids at different pH, and removal of Ca++ or other cations from capsids altered their structures.114,401 For many parvoviruses and AAVs, the VP1 unique region can be exposed to the exterior after certain mild treatments, and that exposure was increased by changes of residues surrounding the fivefold axis of icosahedral symmetry.113,150,247,350,444
Cell Receptor Binding
Various molecules mediate parvovirus cell binding and infection. A cellular receptor for parvovirus B19 is globoside or erythrocyte P antigen (glycosphingolipid tetrahexoseceramide), and people who genetically lack P antigen (p phenotype) are not susceptible to parvovirus B19 infection.57,59 The role of globoside binding is not well understood, and in some studies the capsids did not bind the globoside in vitro.227 Globoside is expressed on mature erythrocytes and erythroid precursors; in tissue culture, excess soluble globoside or anti–P monoclonal antibody can protect erythroid progenitors from infection. However, on other cells, it is uncertain whether the globoside is displayed on the cell surface or functions as a B19 receptor. The α5β1 integrin is a co-receptor for infection of erythroid progenitor cells.454,455
Transferrin receptor type 1 (TfR) is used by CPV and FPV for cell infection, and receptor binding controls differences in the host ranges of those viruses.202,332 Capsids bind through contacts between the TfR apical domain and the capsid threefold spike.201,331 CPV and FPV also bind to sialic acids on some erythrocytes and host cells, although that binding does not mediate infection.34,37 MVM capsids bind sialic acids and
hemagglutinate mouse erythrocytes, and cell infection can be blocked by neuraminidase treatment of the cells prior to virus binding.116 Differences in affinity of binding result in differences in plaque size and pathogenicity in mice.6,273
hemagglutinate mouse erythrocytes, and cell infection can be blocked by neuraminidase treatment of the cells prior to virus binding.116 Differences in affinity of binding result in differences in plaque size and pathogenicity in mice.6,273
The AAVs can have broad cellular tropisms and infect various hosts. Infection may involve participation of more than one cell surface molecule. Receptors or co-receptors for AAV2 include heparan sulfate proteoglycan,231,421 human fibroblast growth factor receptor 1,359 and αVβ5 integrin.420 The differences in tropisms of AAV serotypes allow matching receptors on desired target cells with specific forms of transducing virions. Retargeting AAV capsids to alternative receptors using a variety of approaches has allowed transduction of cells that are normally not susceptible.189,308,344,353,371,467
Endocytosis and Endosomal Release
Parvoviruses require receptor-mediated endocytosis for cell infection. Capsids of CPV and MVM are taken up rapidly into cells by clathrin-mediated endocytosis, and the capsids become associated with clathrin-coated pits and vesicles during uptake.266,334 Infection is inhibited by treatment of cells with lysosomotropic agents, including ammonium chloride (NH4Cl), chloroquine, or bafilomycin A1, indicating that low endosomal pH is required for infection.36,334,373,418 After uptake, capsids are detected in endosomes by antibody staining for several hours187,334,419,453; by in situ hybridization, viral DNA was localized with the capsids in perinuclear compartments for at least several hours.419 The infectious process appears to be slow, and CPV infection could be blocked by anticapsid antibodies injected into the cytoplasm of cells 4 or more hours after virus uptake.444,445
Uptake and endocytosis of the AAV capsids follow similar pathways, and infection may require signaling after receptor binding.137,384 Infection can be affected by the activity of the proteosome system, as protease inhibitors can enhance the infection of AAV2 and AAV5 vectors.373,472 The capsids may be retained for long periods in the endosomal system before escaping into the cytoplasm and being transported to the nucleus.35,138,186
The mechanisms of escape from endocytic vesicles into the cytosol are not fully understood, although wholesale lysis of the endosomal vesicles does not occur.334,418 The PLA2 activity of the VP1 unique region is essential for infection, and that enhances the release of the capsids into the cytoplasm.71,142,151,171,418,480
Transport Within the Cytoplasm
Capsids are likely released into the cytoplasm from a vesicle in a perinuclear location, and further processing and trafficking events in the cytoplasm likely occur before transport to the nucleus. Infection of cells by MVM is affected by the activity of the proteasome, because infection can be reduced by some proteosome inhibitors, including those of the chymotrypsin-like activity (N-tosyl-L-phenylalanine chloromethyl ketone and aclarubin) but not by inhibitors of the trypsin-like activity.373 Mechanisms involved in that activity are not clear; however, there is no clear evidence for ubiquitination of the capsid within the cells or direct proteolytic digestion of the capsids.374
Active mechanisms likely transport capsids to the vicinity of the nucleus. Transport to the perinuclear region and nuclear transport are blocked by treatment of cells with nocodazole to depolymerize microtubules and by injection into the cells of an antibody against the intermediate chain of dynein.417,419,445 By electron microscopy, capsids were seen associated with tubulin and dynein structures in vitro, and viral capsids were also precipitated from infected cells along with intermediate chain of dynein.
Nuclear Transport
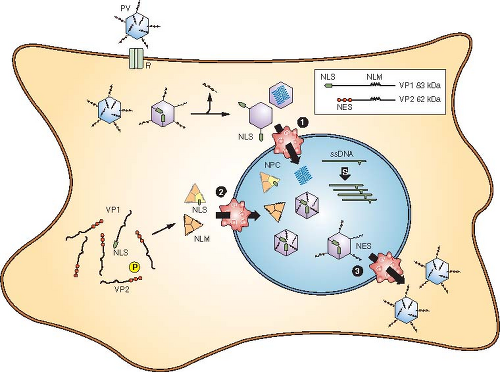
Nuclear transport is thought to involve the nuclear pore complex, although this has not been strictly proven, and injection of MVM capsids into the cytoplasm of Xenopus oocytes was shown to affect the structure of the nuclear envelope.106 However, other studies suggest that parvovirus capsids pass through the nuclear pore intact.417,445 Nuclear entry may require modification of the capsid to expose nuclear localization sequences (NLS) in the VP1 unique region. A VP1 sequence can function for nuclear transport when conjugated to bovine serum albumin (BSA),443 which appears to be exposed on incoming virions, and antibodies to the VP1 unique region blocked infection when injected into cells before virus inoculation.444 MVM capsids have NLS in both VP1 and VP2, with two NLS mapped near the VP1 N-terminus (Fig. 57.4). A VP1 unique region that could mediate nuclear transport is required for efficient cell infection by MVM capsids, and infection was reduced significantly by mutations in the basic sequences in the VP1 N-terminal sequence.270,438 An internal basic sequence acts in a conformation-dependent manner when present in a trimer of VP2 proteins.269,270,372
Viral DNA Release from the Capsid and Initiation of Replication
It is not clear how the viral DNA genome is released from the capsid for replication. Full capsids of MVM and CPV, and likely other viruses including AAV2, have 20 to 30 nucleotides of the 5′ end of the viral genome exposed on the outside of the capsid, and the NS1 or Rep protein is covalently attached to the 5′ end of that DNA in newly produced capsids.123,449 The 3′ end of the viral DNA becomes exposed outside the capsid without capsid disintegration.113,114,444 In the nucleus, that extra-capsid DNA could be used to initiate DNA replication by the host cell DNA polymerase, where the DNA could be removed without disassembly of the stable capsid.
Autonomous Virus DNA Replication
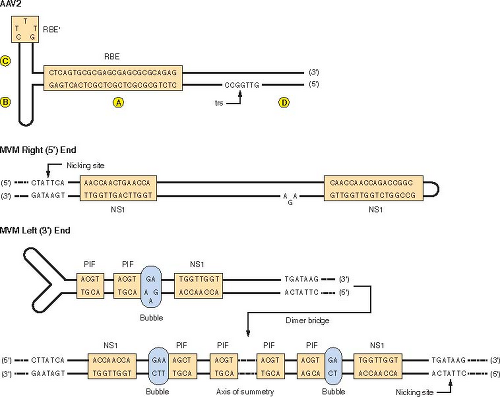
The autonomous parvovirus genome is a linear ssDNA with terminal palindromes (Fig. 57.5), and many viruses have different palindromes at each end22,368 and primarily package the minus DNA strand.424 DNA replication has been most extensively studied for the rodent parvoviruses.23,425 Only two nonstructural (NS) proteins are encoded by most parvoviruses. Replication depends on the DNA replication machinery of the cell and various cellular proteins, and the cell must pass through S phase.369,423,461
The current model for autonomous rodent parvovirus DNA replication is shown in Figures 57.6 and 57.7.119,121 A hairpin formed by the palindrome at the 3′ end of the viral genome serves as a primer for complementary strand synthesis (see step 1, Fig. 57.6). The elongating strand becomes covalently linked to the hairpin form of the 5′ end of the template to form a linear duplex model covalently cross-linked at both ends by DNA hairpins (see step 2, Fig. 57.6). The hairpin formed at the right end is nicked on the newly synthesized strand by NS1
at a site 18 nucleotides downstream from the original 5′ end, and NS1 is covalently linked to the 5′ end formed by the nick (see step 3, Fig. 57.6). The 3′ OH allows repair synthesis to fill the gap at the 3′ end of the newly synthesized strand so that the right end of the intermediate is now a double-stranded form of the original 5′ palindrome with NS1 covalently linked to the 5′ end of the parental strand. The restored 5′ end is 18 nucleotides longer than the 5′ end of virion DNA (step 4, Fig. 57.6). The extended form of the right end palindromic sequence is denatured so that both strands can form hairpins (see step 5, Fig. 57.6), and the 3′ OH at the end of the hairpin on the progeny strand primes DNA synthesis that extends through both strands of the linear duplex monomer so that a linear duplex dimeric structure is formed (see step 7, Fig. 57.6). The left end palindromic sequence is now in a linear double-stranded form at the center of the dimer intermediate (the dimer bridge), with the GAA sequence from the bubble present in the left end palindrome hairpin on the left side of the dimer bridge and the GA sequence to the right (see Figs. 57.5 and 57.7), and those sequences determine the generation of virion DNA strands as outlined later.
at a site 18 nucleotides downstream from the original 5′ end, and NS1 is covalently linked to the 5′ end formed by the nick (see step 3, Fig. 57.6). The 3′ OH allows repair synthesis to fill the gap at the 3′ end of the newly synthesized strand so that the right end of the intermediate is now a double-stranded form of the original 5′ palindrome with NS1 covalently linked to the 5′ end of the parental strand. The restored 5′ end is 18 nucleotides longer than the 5′ end of virion DNA (step 4, Fig. 57.6). The extended form of the right end palindromic sequence is denatured so that both strands can form hairpins (see step 5, Fig. 57.6), and the 3′ OH at the end of the hairpin on the progeny strand primes DNA synthesis that extends through both strands of the linear duplex monomer so that a linear duplex dimeric structure is formed (see step 7, Fig. 57.6). The left end palindromic sequence is now in a linear double-stranded form at the center of the dimer intermediate (the dimer bridge), with the GAA sequence from the bubble present in the left end palindrome hairpin on the left side of the dimer bridge and the GA sequence to the right (see Figs. 57.5 and 57.7), and those sequences determine the generation of virion DNA strands as outlined later.
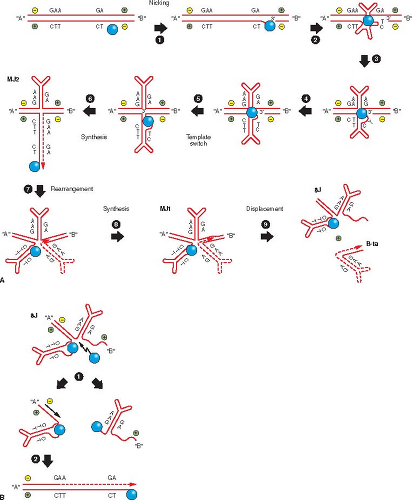
Three sites in the dimeric intermediate are potentially susceptible to single-strand nicking by NS1: two in the covalently closed right end hairpins at each end of the intermediate and a third on the right side of the dimer bridge at GA/TC (nicked at TC; the GAA/TCC sequence on the left side of the dimer bridge is not a site for cleavage)121 (see Figs. 57.6 and 57.7). The resolution process is shown in Figure 57.7.121 Nicking at TC leaves NS1 covalently linked to the 5′ side of the nick and a free 3′ OH that can serve as a primer for DNA synthesis. The helicase activity of the bound NS1 unwinds the helix in the 3′ to 5′ direction, allowing the palindromic sequence of the dimer
bridge to form a cruciform structure that can function as a template for the synthesis. In vitro studies of dimer bridge resolution suggest that when the newly synthesized strand extends beyond the GA bubble sequence, there is a template strand switch—that is, the newly synthesized strand is displaced from the template, forms a hairpin, and copies the strand to the right of the dimer bridge. This creates a new left terminal palindrome in the original flip orientation and displaces a single-stranded monomer equivalent, which is still linked to a duplex, unit length component. This proposed synthesis involves two template strand switches. The original double-stranded replicative form (RF) can be recreated by forming a hairpin of the free right end palindrome to prime synthesis of the complementary strand in the single-stranded region. Alternatively, the right end palindrome to the left of the dimer bridge could be cleaved by NS1 and converted to the extended duplex form, then the strands separated to provide a 3′ OH to prime synthesis, which would displace a single strand containing both the plus and minus strands linked by the extended left end palindrome (Fig. 57.7). The original nick in the dimer bridge leaves NS1 covalently linked at the 5′ end of the template strand for the last reaction and thus provides a stopping point for synthesis. The duplex molecule created is cross-linked at one end by the right palindrome and could go through multiple rounds of displacement synthesis to produce minus strands with the original flip orientation of the left palindrome for packaging into the virion, as seen in MVM.23,117 This model conforms to the observation that NS1 cannot cleave oriL when the left end palindrome is in the hairpin conformation but only when the left palindrome is in the extended duplex form of the dimer
bridge in the RF. However, NS1 can nick oriR when the right end palindrome is hairpinned.
bridge to form a cruciform structure that can function as a template for the synthesis. In vitro studies of dimer bridge resolution suggest that when the newly synthesized strand extends beyond the GA bubble sequence, there is a template strand switch—that is, the newly synthesized strand is displaced from the template, forms a hairpin, and copies the strand to the right of the dimer bridge. This creates a new left terminal palindrome in the original flip orientation and displaces a single-stranded monomer equivalent, which is still linked to a duplex, unit length component. This proposed synthesis involves two template strand switches. The original double-stranded replicative form (RF) can be recreated by forming a hairpin of the free right end palindrome to prime synthesis of the complementary strand in the single-stranded region. Alternatively, the right end palindrome to the left of the dimer bridge could be cleaved by NS1 and converted to the extended duplex form, then the strands separated to provide a 3′ OH to prime synthesis, which would displace a single strand containing both the plus and minus strands linked by the extended left end palindrome (Fig. 57.7). The original nick in the dimer bridge leaves NS1 covalently linked at the 5′ end of the template strand for the last reaction and thus provides a stopping point for synthesis. The duplex molecule created is cross-linked at one end by the right palindrome and could go through multiple rounds of displacement synthesis to produce minus strands with the original flip orientation of the left palindrome for packaging into the virion, as seen in MVM.23,117 This model conforms to the observation that NS1 cannot cleave oriL when the left end palindrome is in the hairpin conformation but only when the left palindrome is in the extended duplex form of the dimer
bridge in the RF. However, NS1 can nick oriR when the right end palindrome is hairpinned.
The genomic ssDNA is generated and packaged when capsids are present. The RF structures giving ssDNA production for encapsidation are undetermined but do not appear to be simple linear duplex monomers; they are likely complex NS1/DNA complexes that are associated with capsid structures.119,121
NS1 is the major parvovirus regulatory protein and is a helicase, adenosine triphosphatase (ATPase), and site-specific nickase that binds to the sequence 5′ACCA3′.101,111,458 When NS1 nicks, it becomes covalently attached to the 5′ side of the nick through residue Y210.26,123,317 This site in the protein is in a motif characteristic of proteins involved in rolling circle replication (RCR motif).206 NS1 cleaves at a site within the extended duplex form of oriL in the dimer bridge and at a site within the hairpinned form of the right end palindrome, but not the hairpinned form of the left terminal palindrome (see Figs. 57.5–57.7). NS1 can presumably cleave the extended duplex form of oriR that would exist in a concatemer bridge with a tail-to-tail junction of monomeric units. Whether the type of asymmetry observed in the cleavage of the oriL bridge would occur has not been reported. The left end palindrome contains a short sequence that is not complementary; when extended in the dimer bridge, there is a 5′GA/TC to the right of the axis of symmetry and a 5′GAA/CTT to the left. Cleavage only occurs at the 5′GA/TC site, owing at least in part to a requirement for a cellular protein (parvovirus initiation factor [PIF]) to be part of the cleavage reaction.97,99 The sequence of the nucleotides in the “bubble” mismatch is not critical; however, the number of nucleotides is, owing to the need for interaction between NS1 and the cellular protein so that adenosine triphosphate (ATP) hydrolysis can occur, allowing the double helix to be unwound and NS1 cleavage of its single-stranded substrate. Both NS1 and PIF can bind to both sides of the dimer bridge, although only the dinucleotide bubble sequence allows formation of the necessary ternary complex (see Fig. 57.5). PIF is a cellular transcriptional factor of the KDWK motif family, which functions as a heterodimer with subunits of 96 and 79 kda.100
Three sequences are required for NS1 cleavage of the covalently closed hairpinned form of oriR found in the dimer RF. An NS1 binding site is near the axis of symmetry in the right end palindrome and another just outboard of the nicking site, and NS1 binding to the two sites leads to looping of the double helix of the hairpin and positions NS1 relative to the nicking site to allow the helicase to expose the site and NS1 to cleave.112,120 A cellular protein, either HMG1 or 2,120 acts in concert with NS1. HMG1/2 preferentially binds to bent DNA, heterocruciforms, and Holliday junctions, and likely assists NS1 in creating the bend in the double helical hairpin to allow oriR cleavage.
Two additional sequences near the right end of the MVM genome affect DNA replication. There is a 65-nucleotide repeat from 4760 to 4850; deletion of one copy of the repeat reduces replication by 100-fold.382 There is also a so-called internal replication sequence from 4489 to 4695.67 Deletion of this sequence reduces the extent of DNA replication by 5-fold. Whether these regions function as internal oris is unknown.
Additional cellular proteins are required for MVM DNA replication, including phosphokinase C (PKC), where PKC eta is required for NS1 to function in RCR,107,316,319 and PKC lambda phosphorylates NS1 at T435 and S473 so that NS1 can function as a helicase.255 G1 phase cell extracts do not support MVM DNA replication in vitro; however, the addition of cyclin A to the G1 phase extract allows DNA replication.
In vivo MVM replication has been found to evoke a DNA damage response (similar to AAV); H2AX, Nbs1, RPA32, Chk2, and p53 are phosphorylated and are recruited to MVM replication centers, where they co-localize with NS1.3,378 Parvovirus DNA replication takes place in discrete subnuclear structures sometimes known as PARs, which contain DNA polymerases α and d.38,128,205
In vivo MVM replication has been found to evoke a DNA damage response (similar to AAV); H2AX, Nbs1, RPA32, Chk2, and p53 are phosphorylated and are recruited to MVM replication centers, where they co-localize with NS1.3,378 Parvovirus DNA replication takes place in discrete subnuclear structures sometimes known as PARs, which contain DNA polymerases α and d.38,128,205
Adeno-Associated Virus DNA Replication
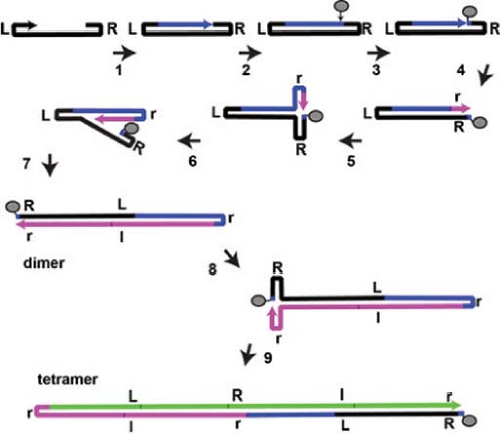
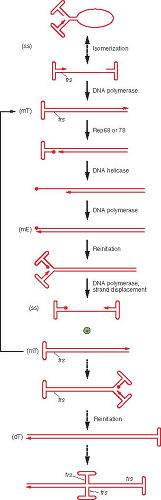
The AAV genome is 4,500 to 5,000 nucleotides long and has palindromic terminal sequences that serve as the primers for DNA replication of AAV (see Fig. 57.5). AAVs have several forms of Rep with multiple functions in DNA replication, but no DNA polymerase or ssDNA binding protein, and additional factors are supplied by either the helper virus and/or the cell. Because of the AAV ITR and the evidence for inversion of the palindromic sequences at both termini, replication can occur by single-strand displacement190,416 (Fig. 57.8). The 3′ terminal repeats fold over to form hairpin structures281 that serve as primers for DNA synthesis416 (see Fig. 57.8). Complementary strand synthesis continues either to the end of the parental template strand, forming a linear duplex molecule cross-linked at one end by the terminal hairpin, or the growing end of the complementary strand may become covalently linked to the hairpinned form of the ITR at the 5′ end of the parental strand template to form a linear duplex cross-linked at both ends. Resolution is achieved by Rep (68 or 78) binding to the hairpin (which has a Rep binding sequence [RBS], as well as a secondary binding site on one of the cross-arms [see Fig. 57.5]) and nicking the original parental strand at the terminal resolution site (TRS), which is opposite the phosphodiester bond between the 3′OH of the primer and the 5′PO4 of the first nucleotide inserted into the progeny strand.54,92,290,380 Rep covalently binds to the 5′PO4 created at the nick,133,207,405 and the 3′OH serves as a primer for “repair” synthesis, filling in the resultant gap at the 5′ end of the parental strand using the displaced hairpin sequence as the template.208 Thus, the original 3′ end of the parental strand is transferred to become the 5′ end of the new strand, and the terminal palindromic sequence is inverted. The 3′ end of the newly synthesized strand may fold over to prime synthesis of another new strand, which can use both complementary strands of the original unit length duplex as a template to produce a linear, duplex dimer-length
molecule.313,416 The structure of the initial primer strand may potentially be either a linear single strand with a hairpinned 3′ end or a single-stranded circle held together by H-bonding between the ITRs (see Fig. 57.8).
molecule.313,416 The structure of the initial primer strand may potentially be either a linear single strand with a hairpinned 3′ end or a single-stranded circle held together by H-bonding between the ITRs (see Fig. 57.8).
The large Rep proteins introduce site-specific nicks at the TRS, and their helicase activities are necessary for the hairpin unwinding during resolution of the hairpinned termini.208,408,462 The endonuclease activity selectively works on the hairpin structure, but poorly, if at all, on the linear duplex form of the same sequence.92,290 The Rep protein binds ATP, and its ATPase activity is required to unwind the double helix, but not for the endonucleolytic activity.208,408,462 The site-specific DNA binding and endonuclease activities have been mapped to the N-terminal domain of the larger Rep 68/78 proteins.406 The RBS is a tandem repeat of four copies of GAGC.92,290,380,407,457 Only two copies of the GAGC sequence are required to bind Rep, and there is apparently wide latitude in the number of copies of the tetranucleotide that are required for Rep to function.93,290,462 Rep 68/78 functions as a multimeric complex that binds to the RBS in the stem of the T-shaped structure of the folded ITR and also to a second site (RBS′, GTTTC) that is at the tip of the cross arm farthest removed from the TRS54,380 (see Fig. 57.5). The TRS consists of a seven base sequence (3′CCGGT/TG5′) that is recognized only on the correct strand.55 The functional complex of Rep is a pentamer.193 Once Rep is bound to the hairpin, Rep unwinds the stem of the ITR in a reaction that requires ATP hydrolysis to generate a single-stranded TRS,55,133,407 then the endonuclease active site carries out a transesterification to link tyrosine 156 of Rep to the 5′ phosphate end of the nick.133,208,405 Rep remains covalently bound until packaging occurs and can be detected on the surface of the viral particle.357
Stay updated, free articles. Join our Telegram channel

Full access? Get Clinical Tree