FIGURE 21–1. Shapes of bacteria. A. Staphylococcus aureus, cocci arranged in clusters; scanning electron micrograph (SEM). B. Group B streptococci, cocci arranged in chains; SEM. C. Bacillus species, straight rods; Gram stain. D. Spirochete, phase contrast, SEM. E. Vibrio, curved rods, SEM. (Reproduced with permission from Willey JM: Prescott, Harley, & Klein’s Microbiology, 7th edition. McGraw-Hill, 2008.)
Bacteria exhibit sphere, rod, and spiral shapes
Whatever the overall shape of the cell, the 1μm size could not accommodate eukaryotic mitochondria, nucleus, Golgi apparatus, lysosomes, and endoplasmic reticulum in a cell that is itself only as large as an average mitochondrion. The solution is in the unique design of the prokaryotic bacterial cell. A generalized bacterial cell is shown in Figure 21–2. The major structures of the cell belong either to the multilayered envelope and its appendages or to the interior core consisting of the nucleoid (or nuclear body) and the cytoplasm. The cytoplasm is analogous to that of eukaryotic cells, but because there is no nucleus it is not clearly separated from the genetic material. The general chemical nature of the bacterial design includes the familiar macromolecules of life (DNA, RNA, protein, carbohydrate, phospholipid) in addition to some macromolecules unique to bacteria such as the peptidoglycan and lipopolysaccharide of bacterial cell walls. The smallness and simplicity of the bacterial design contribute to the ability of the cytosol to grow at least an order of magnitude faster than eukaryotic cells, a significant feature in producing disease.
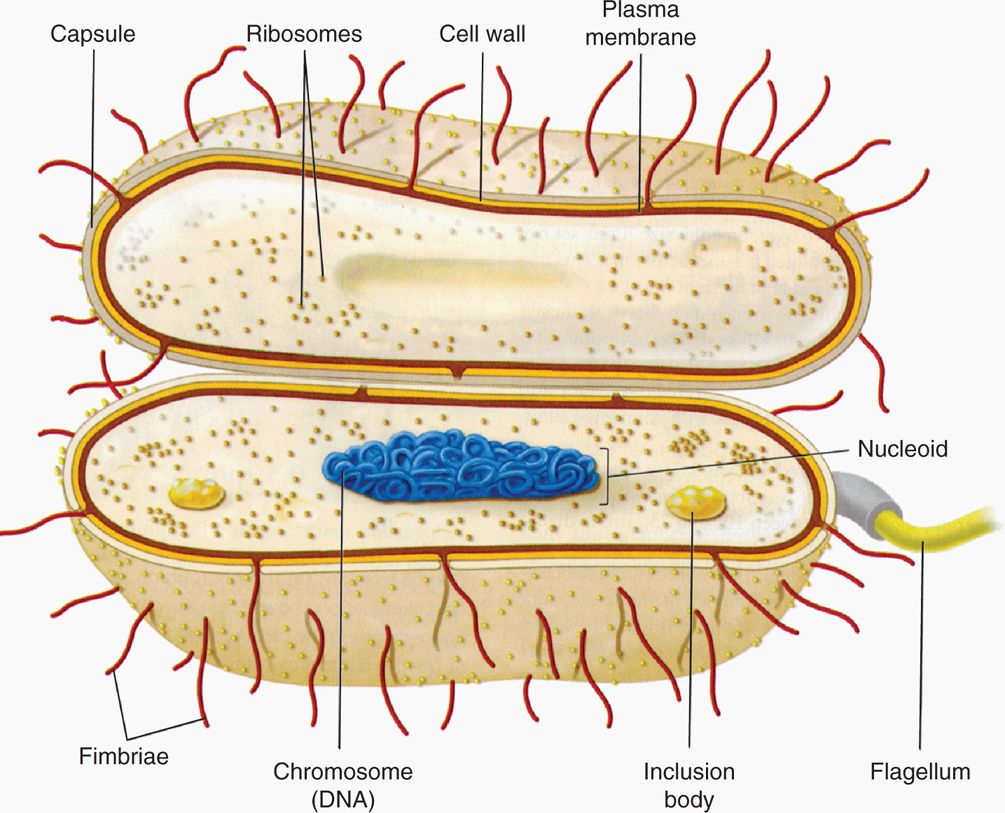
FIGURE 21–2. The prokaryotic bacterial cell. (Reproduced with permission from Willey JM: Prescott, Harley, & Klein’s Microbiology, 7th edition. McGraw-Hill, 2008.)
Prokaryotic design includes envelope, appendages, cytosol, and nucleoid
Chemical nature is similar to eukaryotic cells plus unique components
Design facilitates rapid growth
ENVELOPE AND APPENDAGES
Bacteria have a very plain interior but a complex, even baroque, exterior. This can be readily understood by appreciating that the envelope not only protects the cell against chemical and biologic threats in its environment, but is also responsible for many metabolic processes that are the province of the internal organelles of eukaryotic cells. Structures in the envelope and certain appendages also mediate attachment to human cell surfaces, the first step in disease. Not surprisingly, therefore, more than one-fifth of the specific proteins of well-studied bacteria are located in the envelope. Some of these features are presented in Table 21–1 in relation to the major bacterial cell wall types.
Table 21–1 Components of Bacterial Cells
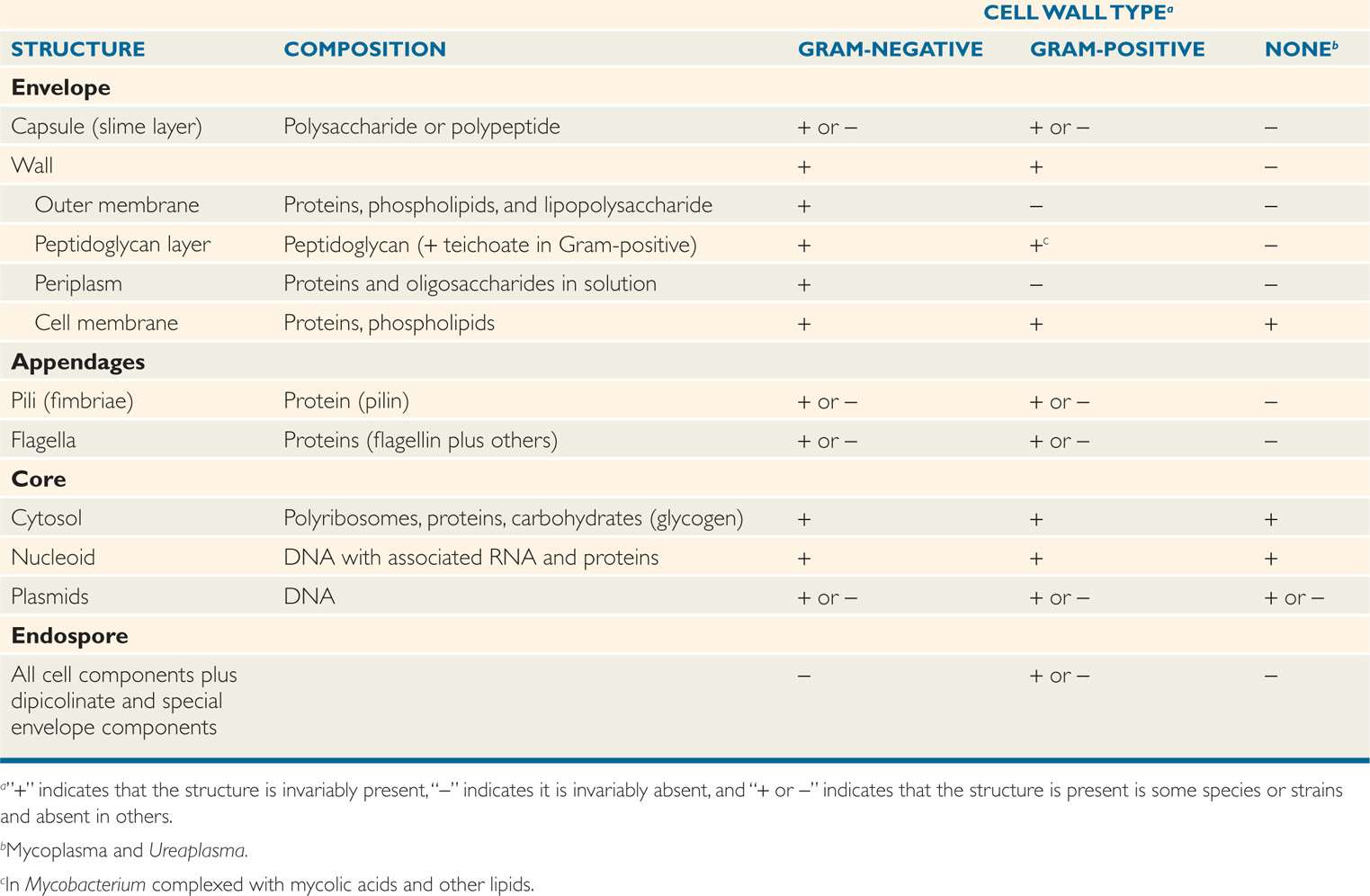
Envelope and appendages carry out multiple functions
 Capsule
Capsule
Many bacterial cells surround themselves with one or other kind of hydrophilic gel. This layer is often thick; commonly it is thicker than the diameter of the cell. Because it is transparent and not readily stained, this layer is usually not appreciated unless made visible by its ability to exclude particulate material, such as India ink or by special capsular stains (Figure 21–3). If the material forms a reasonably discrete layer, it is called a capsule; if it is amorphous in appearance, it is referred to as a slime layer. Almost all bacterial species can synthesize such materials to some degree. Most capsules are polysaccharides made of single or multiple types of sugar residues; a few are simple polypeptides.
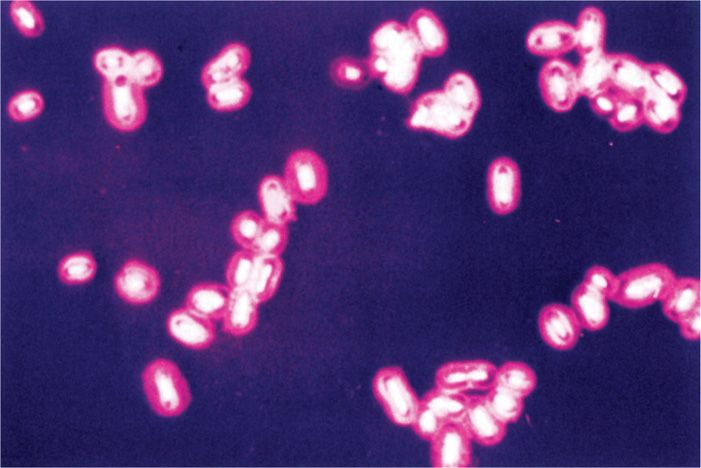
FIGURE 21–3. Bacterial capsule. This capsule surrounding the cells of Klebsiella pneumoniae has been stained red. (Reproduced with permission from Willey JM: Prescott, Harley, & Klein’s Microbiology, 7th edition. McGraw-Hill, 2008.)
Hydrophilic capsules are usually polysaccharides
Capsules provide some general protection for bacteria, but their major function in pathogenic bacteria is protection from the immune system. These features are discussed in Chapter 22. Capsules do not contribute to growth and multiplication and are not essential for cell survival in artificial culture. Capsule synthesis is greatly dependent on growth conditions. For example, the capsule made by the caries-producing Streptococcus mutans consists of a dextran-carbohydrate polymer made in the presence of sucrose.
Capsules protect from immune system
Capsule synthesis depends on growth conditions
 Cell Wall
Cell Wall
Internal to the capsule (if one exists) but still outside the cell proper, a rigid cell wall surrounds all eubacterial cells except wall-less bacteria such as the mycoplasmas and Chlamydia. The structure and function of the bacterial wall is a hallmark of the prokaryotes; nothing like it is found elsewhere. This wall protects the cell from mechanical disruption and from bursting caused by the turgor pressure resulting from the hypertonicity of the cell interior relative to the environment. It also provides a barrier against certain toxic chemical and biologic agents. Its form is responsible for the shape of the cell. Overall, a well-constructed wall protects these minute, fragile cells from chemical and physical assault, while still permitting the rapid exchange of nutrients and metabolic byproducts required for rapid growth.
Unique wall structur prevents osmotic lysis, determines shape
Bacterial evolution has led to two major solutions to cell wall structure. Although the detailed structural basis of the two is now well known, the separation derives from their reaction to a particular staining procedure devised more than a century ago. It is called the Gram stain and is described in Chapter 4. The staining reaction depends on the ability of cells stained with certain dyes to resist extraction of the dye with ethanol-acetone mixtures. The bacteria from which these complexes are readily extracted are called Gram-negative, and those that retain these complexes are termed Gram-positive. Thus, a positive or negative Gram stain response of a cell identifies which of the two types of wall it possesses.
Gram stain distinguishes two major envelope structures
Virtually all bacteria with walls can now be assigned a Gram category even if they cannot be visualized with the stain itself for technical reasons. Examples include the causative agents of tuberculosis and syphilis. Mycobacterium tuberculosis (Gram-positive) has lipids in its cell wall that resist the uptake of most stains. Treponema pallidum (Gram-negative) takes stains poorly but is also too thin to be resolved in the light microscope without special illumination. In these cases, the Gram categorization is based on electron microscopy (Figure 21–4) and chemical analysis of the cell wall.
Poorly staining bacteria still have a Gram category
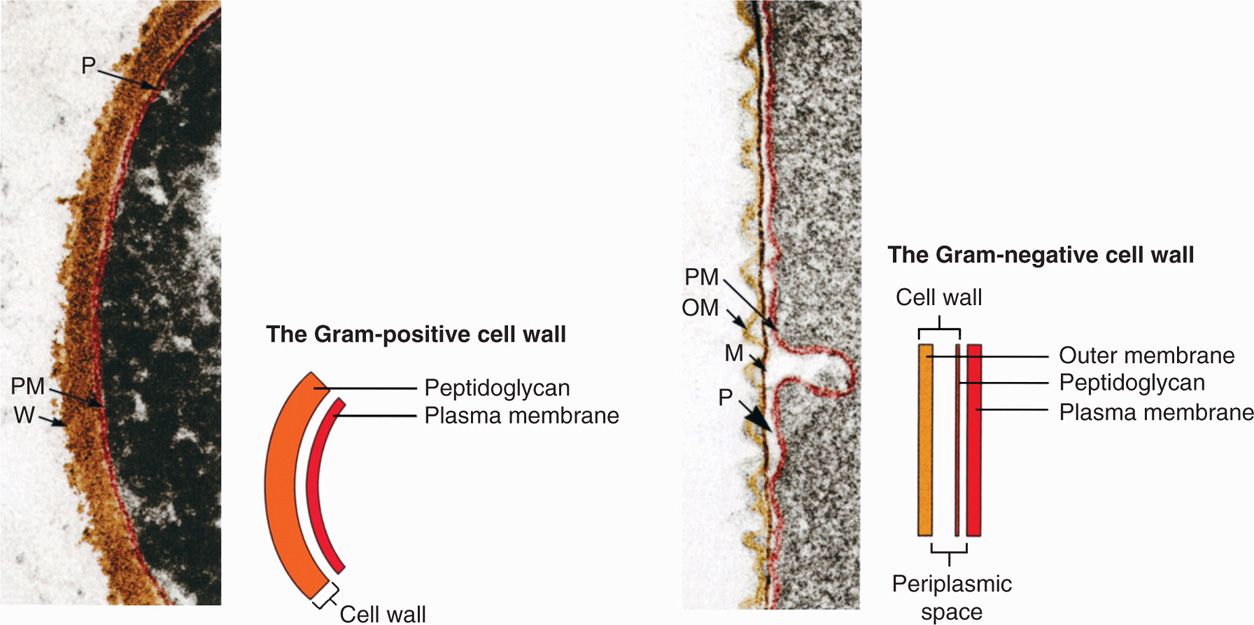
FIGURE 21–4. Gram-positive and Gram-negative cell walls. M, peptidoglycan or murein layer; OM, outer membrane; PM, plasma membrane; P, periplasmic space; W, Gram-positive peptidoglycan wall. (Reproduced with permission from Willey JM: Prescott, Harley, & Klein’s Microbiology, 7th edition. McGraw-Hill, 2008.)
Gram-Positive Cell Wall
The Gram-positive cell wall contains two major components, peptidoglycan and teichoic acids, plus additional carbohydrates and proteins, depending on the species. A generalized scheme illustrating the arrangement of these components is shown in Figure 21–5. The chief component is peptidoglycan, which is found only in prokaryotes. Peptidoglycan consists of a linear glycan chain of two alternating sugars, N-acetylglucosamine (NAG) and N-acetylmuramic acid (NAM) (Figure 21–6). Each muramic acid residue bears a tetra-peptide of alternating L-and D-amino acids. Adjacent glycan chains are cross-linked into sheets by peptide bonds between the third amino acid of one tetrapeptide and the terminal D-alanine of another. The same cross-links between other tetrapeptides connect the sheets to form a three-dimensional, rigid matrix. The cross-links involve perhaps one-third of the tetrapeptides and may be direct or may include a peptide bridge, as, for example, a pentaglycine bridge in Staphylococcus aureus. The cross-linking extends around the cell, producing a scaffold-like giant molecule. Peptidoglycan is much the same in all bacteria, except that there is diversity in the nature and frequency of the cross-linking bridge and in the nature of the amino acids at certain positions of the tetrapeptide.
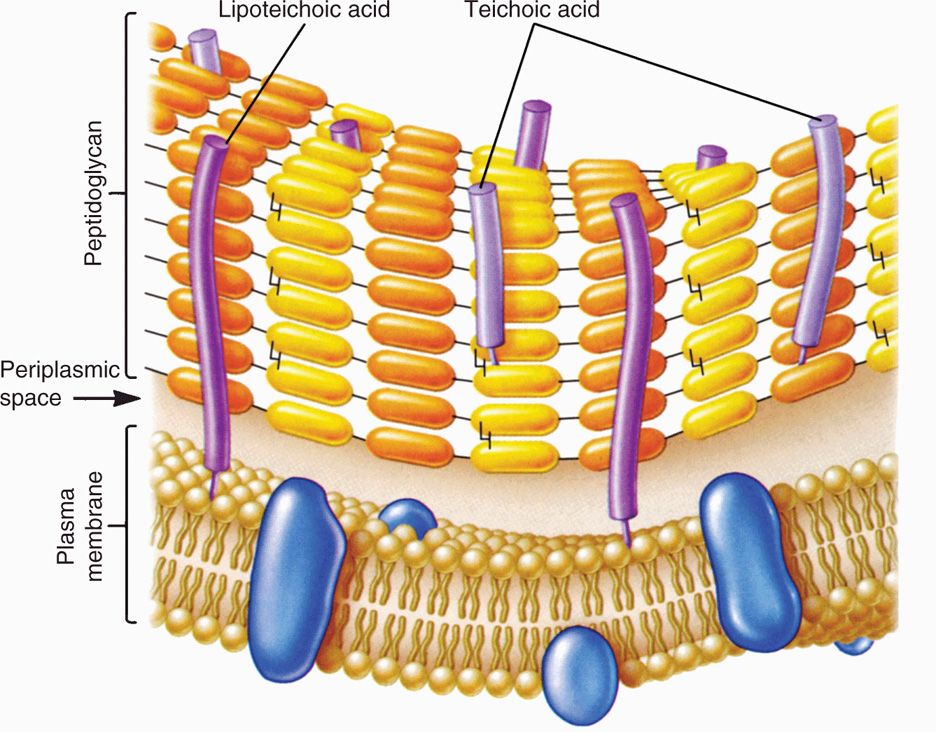
FIGURE 21–5. Gram-positive envelope. (Reproduced with permission from Willey JM: Prescott, Harley, & Klein’s Microbiology, 7th edition. McGraw-Hill, 2008.)
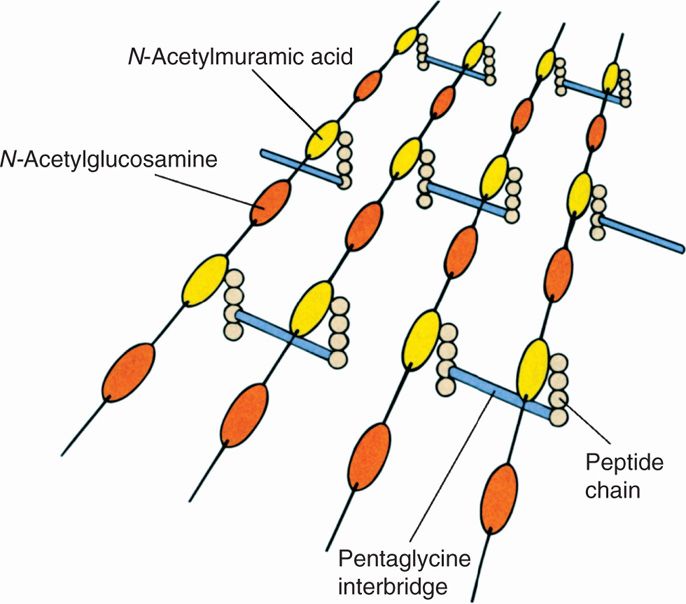
FIGURE 21–6. Peptidoglycan structure. A schematic diagram of one model of peptidoglycan. Shown are the polysaccharide chains, tetrapeptide side chains, and peptide bridges. (Reproduced with permission from Willey JM: Prescott, Harley, & Klein’s Microbiology, 7th edition. McGraw-Hill, 2008.)
Major components of Gram-positive walls are peptidoglycan and teichoic acid
Peptidoglycan comprises glycan chains cross-linked by peptide chains
Scaffold-like sac surrounds cell
The peptidoglycan sac derives its great mechanical strength from the fact that it is a single, covalently bonded structure. Most enzymes found in mammalian hosts and other biologic systems do not degrade peptidoglycan; one important exception is lysozyme, the hydrolase in tears and other secretions, which cleaves the β-1,4 glycosidic bond between muramic acid and glucosamine residues. The role of the peptidoglycan component of the cell wall in conferring osmotic resistance and shape on the cell is easily demonstrated by removing or destroying it. Treatment of a Gram-positive cell with penicillin (which blocks formation of the tetrapeptide cross-links) destroys the peptidoglycan sac, and the wall is lost. Prompt lysis of the cell ensues. If the cell is protected from lysis by suspension in a medium approximately isotonic with the cell interior, such as 20% sucrose, the cell becomes round and forms a sphere called a protoplast.
Components of peptidoglycan provide resistance to most mammalian enzymes
Loss of cell wall leads to lysis or production of protoplasts
A second component of the Gram-positive cell wall is teichoic acid. These compounds are polymers of either glycerol phosphate or ribitol phosphate, with various sugars, amino sugars, and amino acids as substituents. The lengths of the chain and the nature and location of the substituents vary from species to species and sometimes among strains within a species. Up to 50% of the wall may be teichoic acid, some of which is covalently linked to occasional NAM residues of the peptidoglycan. Of the teichoic acids made of polyglycerol phosphate, much is linked not to the wall but to a glycolipid in the underlying cell membrane. This type of teichoic acid is called lipoteichoic acid and appears to play a role in anchoring the wall to the cell membrane and as an epithelial cell adhesin. Besides the major wall components—peptidoglycan and teichoic acids—Gram-positive walls usually have lesser amounts of other molecules characteristic of their species. Some are polysaccharides, such as the group-specific antigens of streptococci; others are proteins, such as the M protein of group A streptococci.
Teichoic and lipoteichoic acids promote adhesion and anchor wall to membrane
Other cell wall components related to species
Gram-Negative Cell Wall
The second kind of cell wall found in bacteria, the Gram-negative cell wall, is depicted in Figure 21–7. Except for the presence of peptidoglycan, there is little chemical resemblance to cell walls of Gram-positive bacteria, and the architecture is fundamentally different. In Gram-negative cells, the amount of peptidoglycan has been greatly reduced, with some of it forming a single-layered sheet around the cell and the rest in a gel-like substance, the periplasmic gel, with little cross-linking. External to this periplasm is an elaborate outer membrane. The proteins in solution in the periplasm consist of enzymes with hydrolytic functions, sometimes antibiotic-inactivating enzymes, and various binding proteins with roles in chemotaxis and in the active transport of solutes into the cell. Oligosaccharides secreted into the periplasm in response to external conditions serve to create an osmotic pressure buffer for the cell.
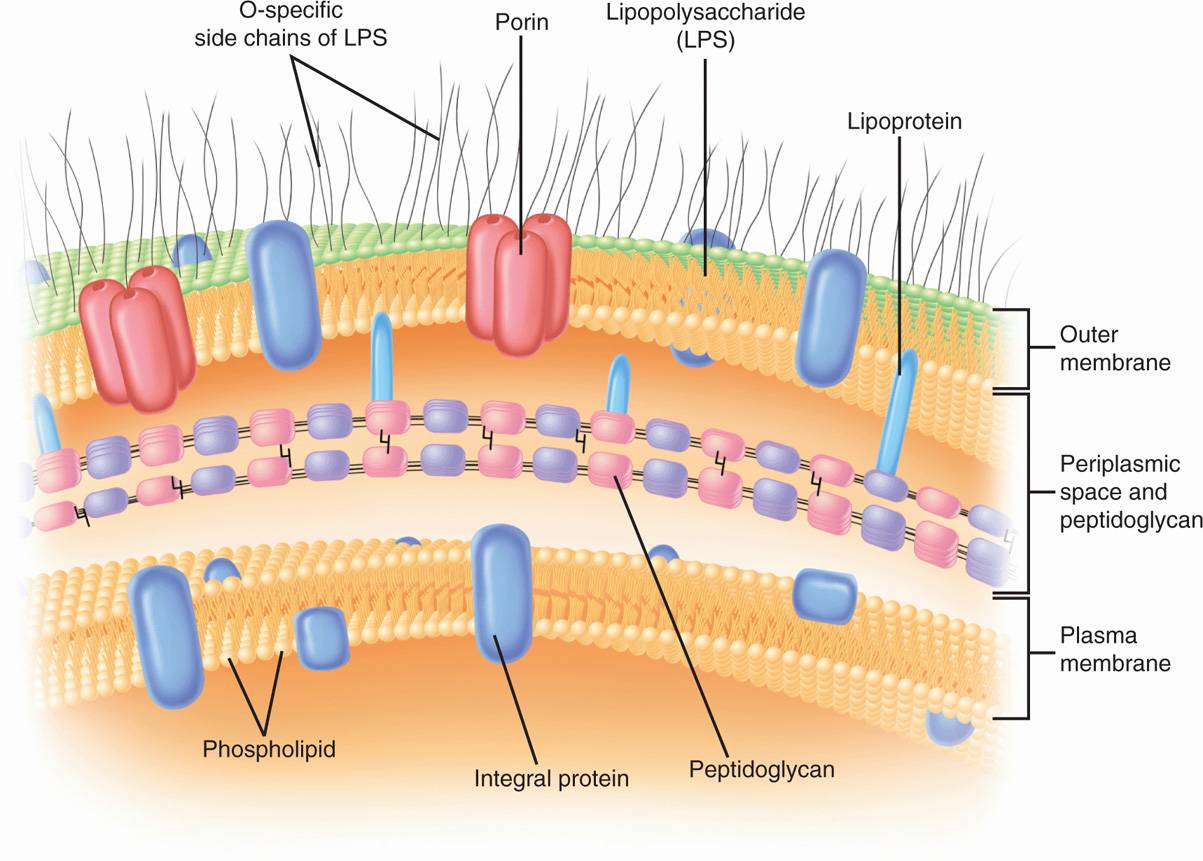
FIGURE 21–7. Gram-negative envelope. (Reproduced with permission from Willey JM: Prescott, Harley, & Klein’s Microbiology, 7th edition. McGraw-Hill, 2008.)
Thin peptidoglycan sac is imbedded in periplasmic gel
Periplasmic proteins have transport, chemotactic, and hydrolytic roles
Gram-negative outer membrane is phospholipoprotein bilayer, of which the outer leaflet is LPS endotoxin
The periplasm is an intermembrane structure, lying between the cell membrane and a special membrane unique to Gram-negative cells, the outer membrane. This has an overall structure similar to most biologic membranes with two opposing phospholipid-protein leaflets. However, in terms of its chemical composition, the outer membrane is unique biologically. Its inner leaflet consists of ordinary phospholipids, but these are replaced in the outer leaflet by a special molecule called lipopolysaccharide (LPS), which is extremely toxic to humans and other animals, and is called an endotoxin. Even in minute amounts, such as the amounts released to circulation during the course of a Gram-negative infection, this substance can produce a fever and shock syndrome called Gram-negative shock, or endotoxic shock.
LPS consists of a toxic lipid A (a phospholipid containing glucosamine rather than glycerol), a core polysaccharide (containing some unusual carbohydrate residues and fairly constant in structure among related species of bacteria), and O antigen polysaccharide side chains (Figure 21–8A and B). The last component constitutes the major surface antigen of Gram-negative cells.
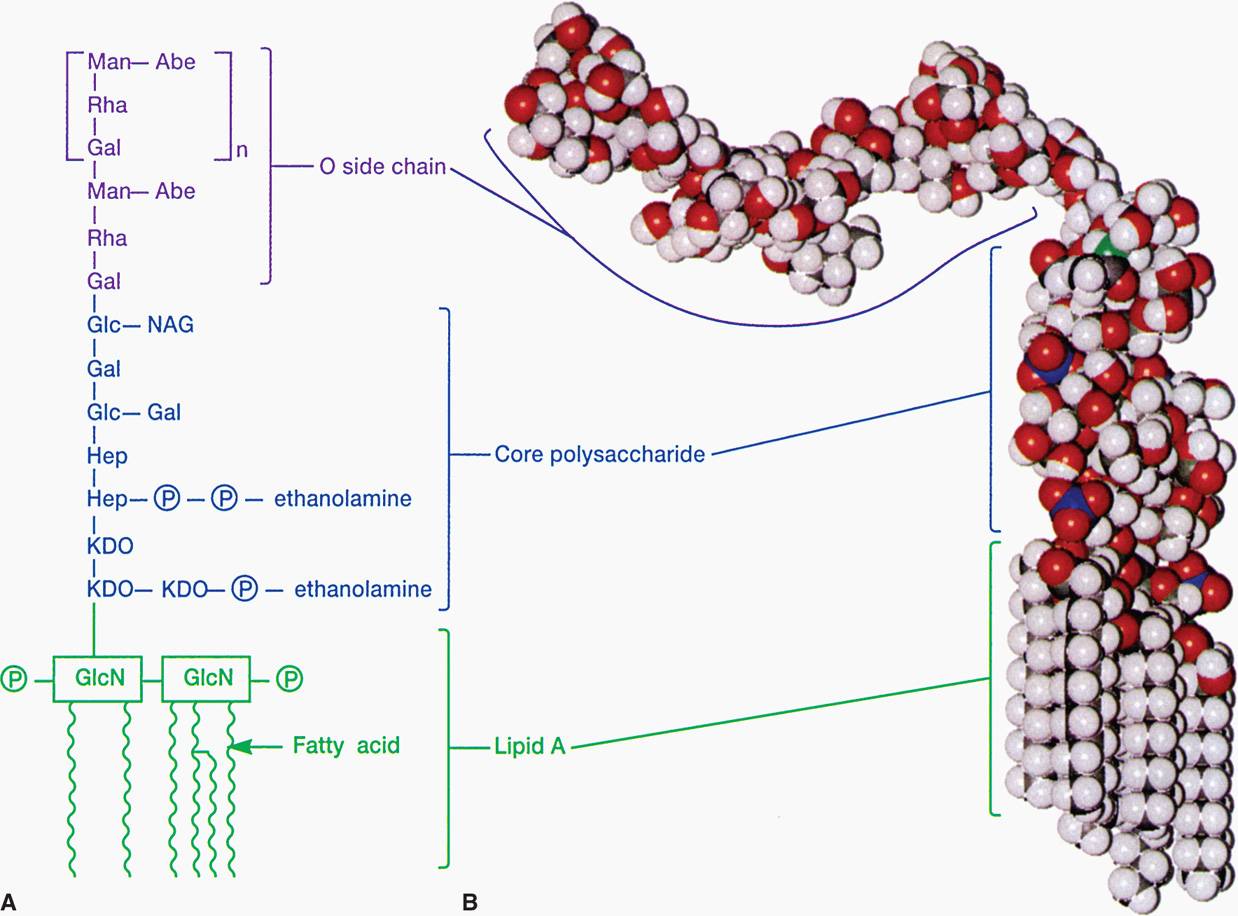
FIGURE 21–8. Lipopolysaccharide structure. A. O side chain—formed by linked sugars. Core polysaccharide—sugars linked to N-acetylglucoseamine (NAG) and keto-deoxycholate (KDO). Lipid A—buried in the outer membrane. B. Molecular model. (Reproduced with permission from Willey JM: Prescott, Harley, & Klein’s Microbiology, 7th edition. McGraw-Hill, 2008.)
Lipid A is the toxic moiety of LPS; polysaccharides are antigenic determinants
The presence of the outer membrane results in the covering of Gram-negative cells by a formidable permeability barrier. For whatever benefit is afforded by possessing a wall with an outer membrane, Gram-negative bacteria must make provision for the entry of nutrients. Special structural proteins, called porins, form pores through the outer membrane that make it possible for hydrophilic solute molecules to diffuse through it and into the periplasm.
Impermeability of outer membrane is overcome by porins
In evolving a cell wall containing an outer membrane, Gram-negative bacteria have succeeded in (1) creating the periplasm, which holds digestive and protective enzymes and proteins important in transport and chemotaxis; (2) presenting an outer surface with strong negative charge, which is important in evading phagocytosis and the action of complement; and (3) providing a permeability barrier against such dangerous molecules as host lysozyme, bile salts, digestive enzymes, and many antibiotics.
Outer membrane has many functions
 Cell Membrane
Cell Membrane
Generally, the cell (plasma) membrane of bacteria (Figure 21–9) is similar to the familiar bileaflet membrane, containing phospholipids and proteins, and which is found throughout the living world. However, there are important differences. The bacterial cell membrane is exceptionally rich in proteins and does not contain sterols (except mycoplasmas). The bacterial chromosome is attached to the cell membrane, which plays a role in the segregation of daughter chromosomes at cell division, analogous to the role of the mitotic apparatus of eukaryotes. The membrane is the site of synthesis of DNA, cell wall polymers, and membrane lipids. It contains the entire electron transport system of the cell (and, hence, is functionally analogous to the mitochondria of eukaryotes). It contains receptor proteins that function in chemotaxis. Similar to the cell membranes of eukaryotes, it is a permeability barrier and contains proteins involved in the selective and active transport of solutes. It is also involved in secretion of the exterior of proteins (exoproteins), including exotoxins and hydrolytic enzymes involved in the pathogenesis of disease. The bacterial cell membrane is therefore the functional equivalent of most of the organelles of the eukaryotic cell and is vital to the growth and maintenance of the cell.
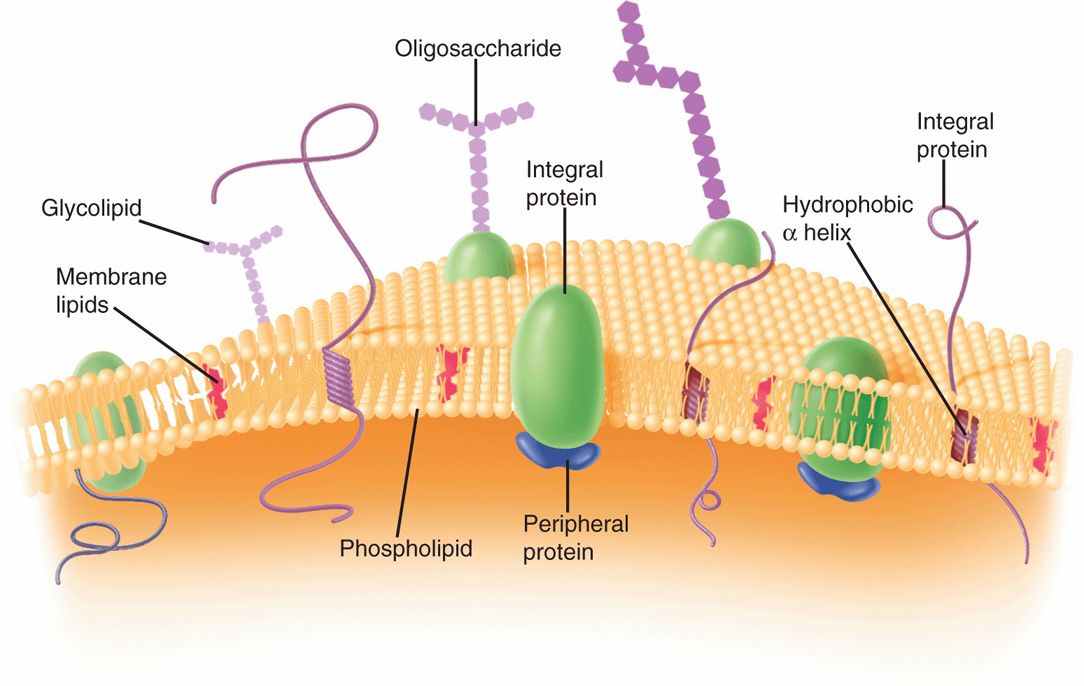
FIGURE 21–9. Bacterial cell membrane. (Reproduced with permission from Willey JM: Prescott, Harley, & Klein’s Microbiology, 7th edition. McGraw-Hill, 2008.)
Phospholipid-protein bilayer lacking sterols
Roles in synthetic, homeostatic, secretory, and electron transport processes
Functional equivalent of many eukaryotic organelles
 Flagella
Flagella
Flagella are molecular organelles of motility found in many species of bacteria, both Gram-positive and Gram-negative. They may be distributed around the cell (an arrangement called peritrichous from the Greek trichos for “hair”), at one pole (polar or monotrichous), or at both ends of the cell (lophotrichous). They are long (up to 20 µim), slender, rigid, and individually helical in shape. Flagella propel the cell by rotating at the point of insertion in the cell envelope. The presence or absence of flagella and their position are important taxonomic characteristics.
Flagella are rotating helical protein structures responsible for locomotion
The flagellar apparatus is complex, but consists entirely of proteins attached to the cell by a basal body consisting of several proteins organized as rings on a central rod. Other structures include a hook that acts as a universal joint and ring-like bushings. All propel the long filament, which consists of polymerized molecules of a single protein species called flagellin. Flagellin varies in amino acid sequence from strain to strain. This makes flagella useful surface antigens for strain differentiation, particularly among the Enterobacteriaceae.
Flagella have bushing rings in cell envelope
Flagellar filament is composed of the protein flagellin
 Pili
Pili
Pili (also called fimbriae) are hair-like projections found on the surface of cells of many Gram-positive and Gram-negative species. They are composed of molecules of a protein called pilin arranged to form a tube with a minute, hollow core. There are two general classes, common pili and sex pili (see Figure 21–33). Up to a thousand common pili cover the surface of the cell (Figure 21–10). They are, in many cases, adhesins, which are responsible for the ability of bacteria to colonize surfaces and cells. These processes are not always passive, since some pili can retract mediating movement across cell surfaces. Some pili are specialized for adherence to certain cell types such as enterocytes or uroepithelial cells. The same cell may have common and specialized pili. The sex pilus is involved in the exchange of genetic material between some Gram-negative bacteria.
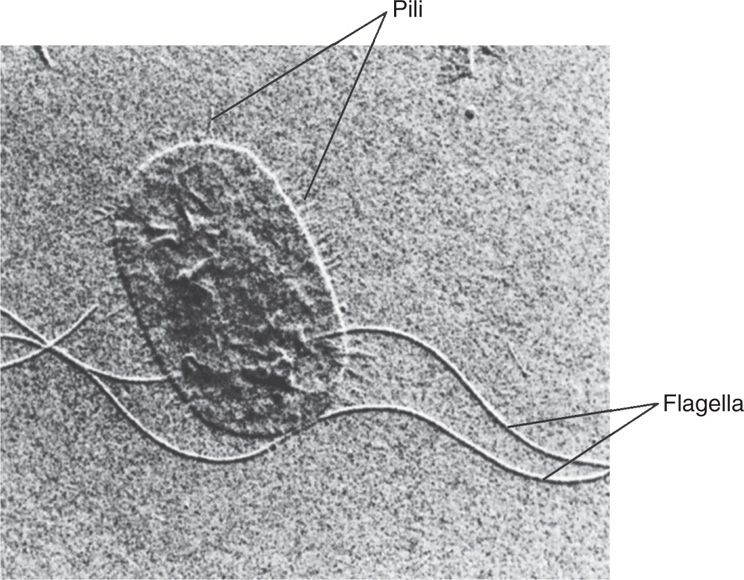
FIGURE 21–10. Flagella and pili. The long flagella and numerous shorter pili are evident in this electron micrograph of Proteus mirabilis. (Reproduced with permission from Willey JM: Prescott, Harley, & Klein’s Microbiology, 7th edition. McGraw-Hill, 2008.)
Pili are tubular hair-like projections
Pili have adherence roles and can retract
Specialized pili mediate selective attachment or genetic transfer
CORE
In contrast to the structural richness of the layers and appendages of the cell envelope, the interior appears relatively simple in transmission electron micrographs of thin sections of bacteria. There are two clearly visible regions, one granular (the cytoplasm) and one fibrous (the nucleoid). In addition, many bacteria possess plasmids that are usually circular, double-stranded DNA bodies in the cytoplasm that are separate from the larger nucleoid; plasmids are too small to be visible in thin sections of bacteria.
 Cytoplasm
Cytoplasm
The dense cytoplasm (cytosol) is bounded by the cell membrane. It appears granular because it is densely packed with ribosomes, which are much more abundant than in the cytoplasm of eukaryotic cells. This is a reflection of the higher growth rate of bacteria. Each ribosome is a ribonucleoprotein particle consisting of three species of rRNA (5 S, 16 S, and 23 S) and over 50 proteins. The overall subunit structure (one 50 S plus one 30 S particle) of the 70 S bacterial ribosome resembles that of eukaryotic ribosomes, but is smaller and differs sufficiently in function that a very large number of antimicrobial agents have the prokaryotic ribosome as their target. The number of ribosomes varies directly with the growth rate of the cell. Except for the functions associated with the cell membrane, all of the metabolic reactions of the cell take place in the cytoplasm.
Cytoplasm contains 70 S ribosomes
Number varies with growth rate
The bacterial cytoplasm has a cytoskeleton which localizes proteins, participates in cell division, and along with the cell wall peptidoglycan, gives shape to the cell. The bacterial cytoskeleton elements are chemical and structural homologs of the microfilaments, microtubules, and intermediate filaments of eukaryotic cells. In the bacterial cell the microfilaments are made from actin and the microtubules from tubulin. Multiple counterparts of intermediate filaments are formed from a mixture of proteins, some of which are unique to bacteria. Modification of the cytoskeleton is a major mechanism of bacterial virulence.
Actin, tubulin, intermediate filaments form cytoskeleton
 Nucleoid
Nucleoid
The nucleoid is a region of the cytoplasm which contains the genome and a collection of related proteins. The bacterial genome resides on a single chromosome and for bacterial pathogens ranges between 600 and 6000 genes encoded in one large, circular molecule of double-stranded DNA. This molecule is more than 1 mm long, and it therefore exceeds the length of the cell by about 1000 times. Tight packing displaces ribosomes and other cytosol components, creating regions that contain a chromosome, coated usually by polyamines and some specialized DNA-binding proteins. The double-helical DNA chain is twisted into supercoils and attached to the cell membrane and/or some central structure at a large number of points. This creates folds of DNA, each of which is independently coiled into a tight bundle. Each nuclear body corresponds to a DNA molecule. The number of nuclear bodies varies as a function of growth rate; resting cells have only one, and rapidly growing cells may have as many as four. Some bacteria have a linear chromosome, and others may have more than one chromosome.
Circular chromosome of supercoiled double-stranded DNA
Attached to cell membrane and central structures
The absence of a nuclear membrane confers on the prokaryotic cell a great advantage for rapid growth in changing environments. Ribosomes can be translating mRNA molecules even as the latter are being made; no transport of mRNA from where it is made to where it functions is needed.
 Plasmids
Plasmids
Many bacteria contain small, usually circular, covalently closed, double-stranded DNA molecules separate from the chromosome. Individual species have regulatory systems controlling plasmids, and more than one type or multiple copies (>40) of a single plasmid may be present in the same cell. Plasmids typically contain up to 30 genes and replicate independent of the chromosome. They are unlikely to contain genes essential for survival of the cell but may have specialized genes such as those mediating virulence or resistance to antimicrobial agents. In fact, many attributes of virulence including, production of pili and exotoxins, and the complex apparatus for injection secretion systems may be determined by plasmid genes.
Plasmids are small, circular, double-stranded DNA molecules
Virulence and resistance genes are present
SPORES
Endospores are small, dehydrated, metabolically quiescent forms that are produced by some bacteria in response to nutrient limitation or a related sign that tough times are coming. Very few species produce spores (the term is loosely used as equivalent to endospores), but they are particularly prevalent in the environment. Some spore-forming bacteria are of great importance in medicine, causing such diseases as anthrax, gas gangrene, tetanus, and botulism. All spore formers are Gram-positive rods. The bacterial endospore is not a reproductive structure. One cell forms one spore under adverse conditions (the process is called sporulation). The spore may persist for a long time (centuries) and then, on appropriate stimulation, give rise to a single bacterial cell (germination). Spores, therefore, are survival rather than reproductive devices.
Endospores are hardy, quiescent forms of some Gram-positives
Spore-forming allows survival under adverse conditions
Spores of some species can withstand extremes of pH and temperature, including boiling water, for surprising periods of time. The thermal resistance is brought about by the low water content and the presence of a large amount of a substance found only in spores, calcium dipicolinate. Resistance to chemicals and, to some extent, radiation is aided by extremely tough, special coats surrounding the spore. These include a spore membrane (equivalent to the former cell membrane); a thick cortex composed of a special form of peptidoglycan; a coat consisting of a cysteine-rich, keratin-like, insoluble structural protein; and, finally, an external lipoprotein and carbohydrate layer called an exosporium.
Resistance of spore is due to dehydrated state, calcium dipicolinate, and specialized coats
Sporulation is under active investigation. The molecular process by which a cell produces a highly differentiated product that is incapable of immediate growth but is able to sustain growth after prolonged periods of nongrowth under extreme conditions of heat, desiccation, and starvation is of great interest. In general, the process involves the initial walling off of a nucleoid and its surrounding cytosol by invagination of the cell membrane, with later additions of special spore layers (Figure 21–11). Germination begins with activation by heat, acid, and reducing conditions. Initiation of germination eventually leads to the outgrowth of a new vegetative cell of the same genotype as the cell that produced the spore.
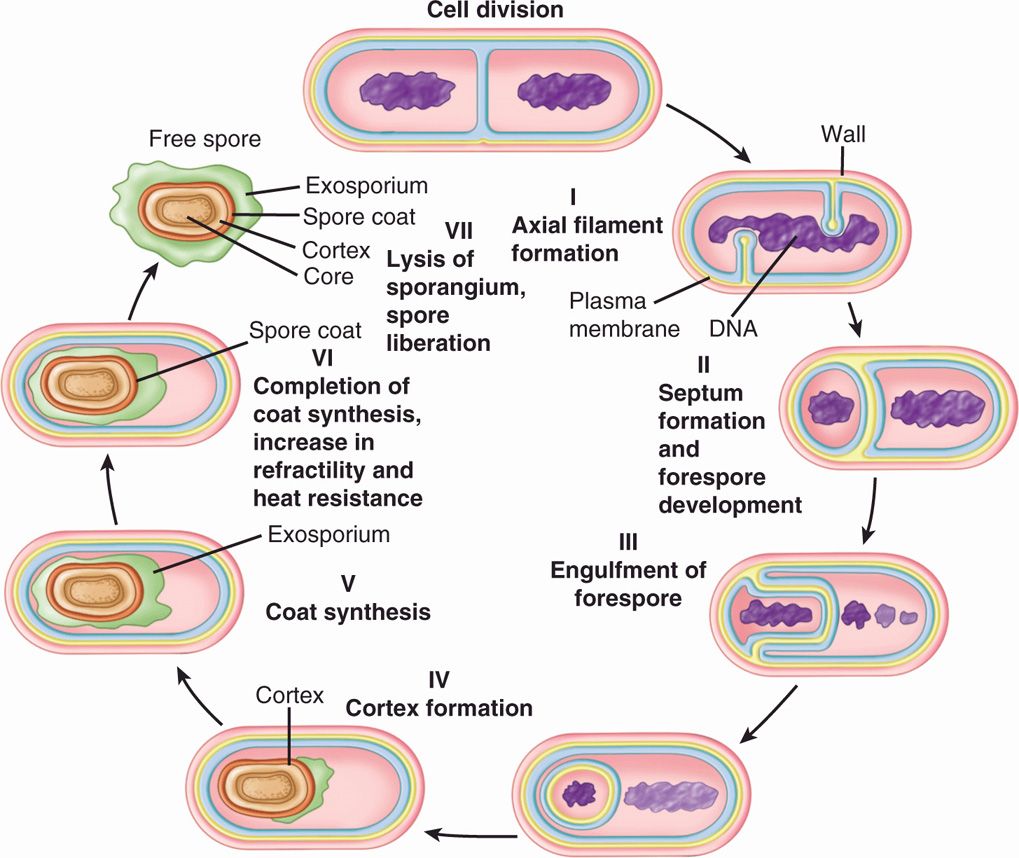
FIGURE 21–11. Stages of bacterial spore formation. (Reproduced with permission from Willey JM: Prescott, Harley, & Klein’s Microbiology, 7th edition. McGraw-Hill, 2008.)
Germination reproduces a cell identical to that which was sporulated
BACTERIAL GROWTH AND METABOLISM
Growth of bacteria is accomplished by an orderly progress of metabolic processes followed by cell division by binary fission. This requires metabolism, which produces cell material from the nutrient substances in the environment; regulation, which coordinates the progress of the hundreds of independent biochemical processes in an orderly way; and, finally, cell division, which produces two independent living units from one.
Growth requires metabolism, regulation, and division by binary fission
BACTERIAL METABOLISM
Many of the principles of metabolism are universal. This section focuses on the unique aspects of bacterial metabolism that are important in medicine. The need to compare bacterial and mammalian pathways is muted by the fact that much of what we understand about human metabolism is derived from work with Escherichia coli.
The broad differences between bacteria and human eukaryotic cells can be summarized as follows:
Speed. Bacteria metabolize at a rate 10 to 100 times faster.
Versatility. Bacteria use more varied compounds as energy sources and are much more diverse in their nutritional requirements.
Simplicity. The prokaryotic body plan makes it possible for bacteria to synthesize macromolecules in a streamlined way.
Uniqueness. Some biosynthetic processes, such as those producing peptidoglycan, lipopolysaccharide, and toxins, are unique to bacteria.
Bacterial metabolism is highly complex. The bacterial cell synthesizes itself and generates energy by as many as 2000 chemical reactions. These reactions can be classified according to their function in the metabolic processes of fueling, biosynthesis, polymerization, and assembly.
 Fueling Reactions
Fueling Reactions
Fueling reactions provide the cell with energy and with the 12 precursor metabolites used in biosynthetic reactions (Figure 21–12). The first step is the capture of nutrients from the environment. Other than water, oxygen, and carbon dioxide, almost no important nutrients enter the cell by simple diffusion, because the cell membrane is too effective a barrier. Some transport occurs by facilitated diffusion in which a protein carrier in the cell membrane, specific for a given compound, participates in the shuttling of molecules of that substance from one side of the membrane to the other (Figure 21–13A and B). Because no energy is involved, this process can work only with, never against, a concentration gradient of the given solute.
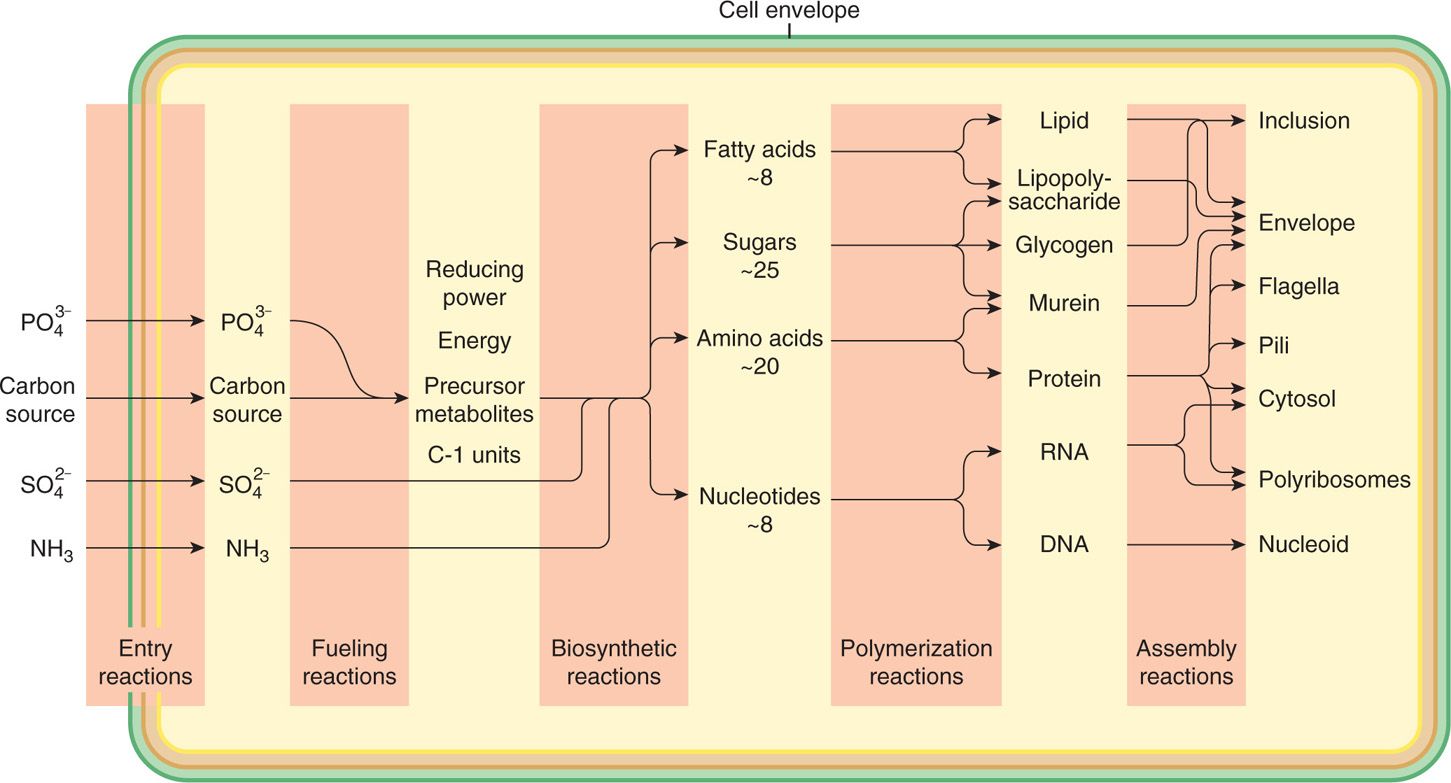
FIGURE 21–12. Bacterial metabolism. General pattern of metabolism leading to the synthesis of a bacterial cell from glucose
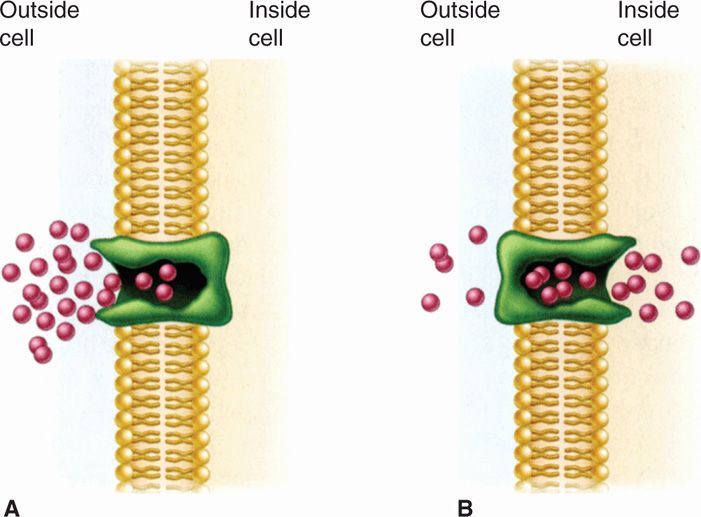
FIGURE 21–13. Facilitated diffusion. A. The membrane carrier can change conformation after binding an external molecule and subsequently release the molecule to the cell interior. B. It then returns to the outward oriented position and is ready to bind another solute molecule. Because there is no energy input, molecules continue to enter only as long as their concentration is greater on the outside. (Reproduced with permission from Willey JM: Prescott, Harley, & Klein’s Microbiology, 7th edition. McGraw-Hill, 2008.)
Substrates enter despite permeability barriers
Facilitated diffusion involves shuttling by carrier protein
Active transport mechanisms involve specific protein molecules as carriers of particular solutes, but the process is energy linked and can therefore establish a concentration gradient. That is, active transport can pump “uphill.” Bacteria have multiple systems of active transport, some of which involve ATP-dependent binding proteins (Figure 21–14) and others that require proton pumps driven by electron transport within the energized cell membrane. Another mechanism called group translocation involves the chemical conversion of the solute into another molecule as it is transported.
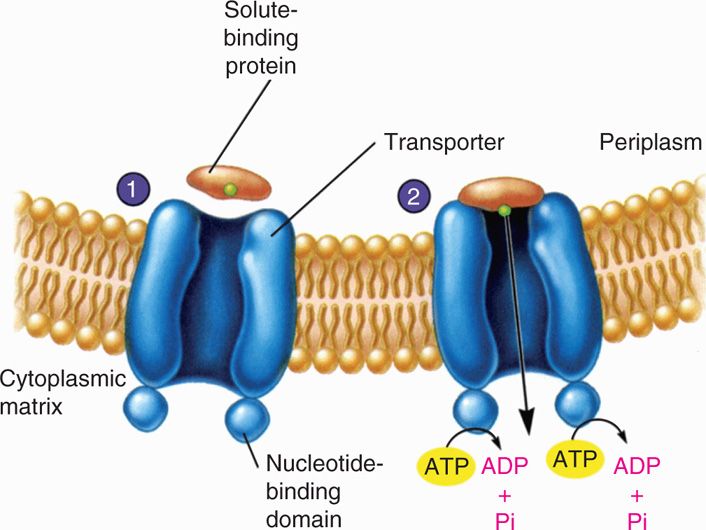
FIGURE 21–14. Active transport. 1. The solute-binding protein binds the substrate to be transported and approaches the transporter complex. 2. The solute binding which is moved across the membrane with the aid of ATP hydrolysis (Reproduced with permission from Willey JM: Prescott, Harley, & Klein’s Microbiology, 7th edition. McGraw-Hill, 2008.)
Active transport involves binding proteins and ATP or proton gradient energy
The transport of iron and other metal ions needed in small amounts for growth is special, and of particular importance in virulence. There is little free Fe3+ in human blood or other body fluids, because it is sequestered by iron-binding proteins (eg, transferrin in blood and lactoferrin in secretions). Bacteria must have iron to grow, and their colonization of the human host requires capture of iron. Bacteria secrete siderophores (iron-specific chelators) to trap Fe3+; the iron-containing chelator is then transported into the bacterium by specific active transport.
Bacterial siderophores chelate iron and are actively transported into cell
Once inside the cell, sugar molecules or other sources of carbon and energy are metabolized by the Embden-Meyerhof glycolytic pathway, the pentose phosphate pathway, and the Krebs cycle to yield the carbon compounds needed for biosynthesis. Some bacteria have central fueling pathways (eg, the Entner-Doudoroff pathway) other than those familiar in mammalian metabolism.
Central fueling pathways produce biosynthetic precursors
Working in concert, the central fueling pathways produce the 12 precursor metabolites. Connections to fermentation and respiration pathways allow the reoxidation of reduced coenzyme nicotinamide adenine dinucleotide (NADH) to NAD+ and the generation of ATP. Bacteria make ATP by substrate phosphorylation in fermentation or by a combination of substrate phosphorylation and oxidative phosphorylation in respiration. (Photosynthetic bacteria are not important in medicine.)
Fermentation and respiration pathways each regenerate ATP and NAD+
Fermentation involves direct transfer of proton and electron to organic receptor acceptor
Fermentation is the transfer of electrons and protons via NAD+ directly to an organic acceptor. Pyruvate occupies a pivotal role in fermentation (Figure 21–15). Fermentation is an inefficient way to generate ATP, and consequently huge amounts of sugar must be fermented to satisfy the growth requirements of bacteria anaerobically. Large amounts of organic acids and alcohols are produced in fermentation. Which compounds are produced depends on the particular pathway of fermentation used by a given species, and therefore the profile of fermentation products is a diagnostic aid in the clinical laboratory.
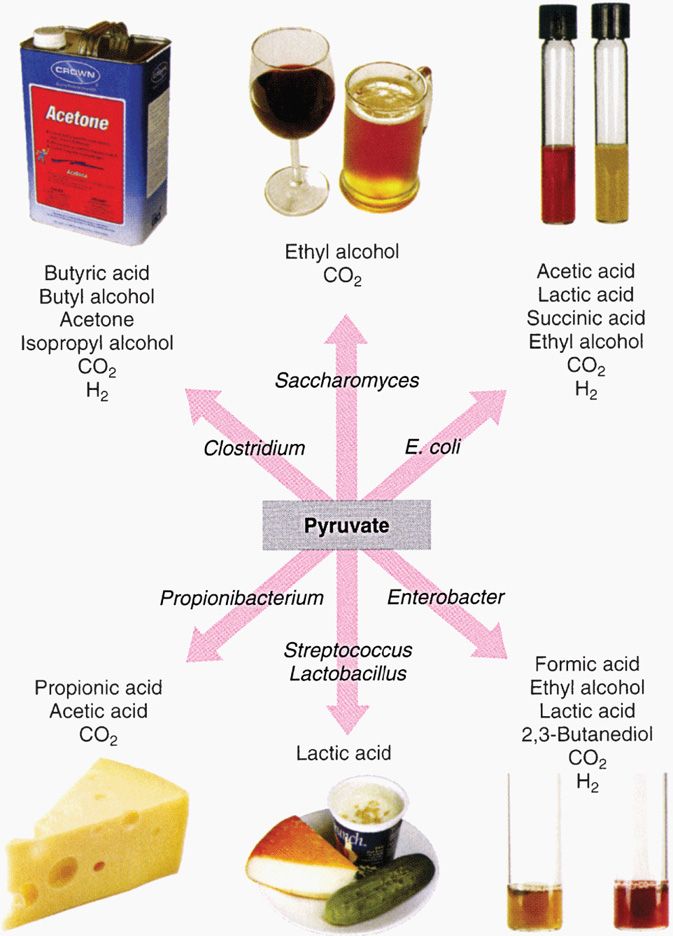
FIGURE 21–15. End products of fermentation pathways. Because a given type of organism uses a characteristic fermentation pathway, the end products can also be used as an identifying marker. (Reproduced with permission from Nester EW: Microbiology: A Human Perspective, 6th edition. 2009.)
Stay updated, free articles. Join our Telegram channel

Full access? Get Clinical Tree


