35
DNA Organization, Replication, & Repair
OBJECTIVES
After studying this chapter, you should be able to:
![]() Appreciate that roughly 3 × 109 base pairs of DNA that compose the haploid genome of humans are divided uniquely between 23 linear DNA units, the chromosomes. Humans, being diploid, have 23 pairs of chromosomes: 22 autosomes and 2 sex chromosomes.
Appreciate that roughly 3 × 109 base pairs of DNA that compose the haploid genome of humans are divided uniquely between 23 linear DNA units, the chromosomes. Humans, being diploid, have 23 pairs of chromosomes: 22 autosomes and 2 sex chromosomes.
![]() Understand that human genomic DNA, if extended end-to-end, would be meters in length, yet still fits within the nucleus of the cell, an organelle that is only microns (μ; 10–6 meters) in diameter. Such condensation in DNA length is induced following its association with the highly positively charged histone proteins resulting in the formation of a unique DNA-histone complex termed the nucleosome. Nucleosomes have DNA wrapped around the surface of an octamer of histones.
Understand that human genomic DNA, if extended end-to-end, would be meters in length, yet still fits within the nucleus of the cell, an organelle that is only microns (μ; 10–6 meters) in diameter. Such condensation in DNA length is induced following its association with the highly positively charged histone proteins resulting in the formation of a unique DNA-histone complex termed the nucleosome. Nucleosomes have DNA wrapped around the surface of an octamer of histones.
![]() Explain that strings of nucleosomes form along the linear sequence of genomic DNA to form chromatin, which itself can be more tightly packaged and condensed, which ultimately leads to the formation of the chromosomes.
Explain that strings of nucleosomes form along the linear sequence of genomic DNA to form chromatin, which itself can be more tightly packaged and condensed, which ultimately leads to the formation of the chromosomes.
![]() Appreciate that while the chromosomes are the macroscopic functional units for DNA recombination, gene assortment, and cellular division, it is DNA function at the level of the individual nucleotides that composes regulatory sequences linked to specific genes that are essential for life.
Appreciate that while the chromosomes are the macroscopic functional units for DNA recombination, gene assortment, and cellular division, it is DNA function at the level of the individual nucleotides that composes regulatory sequences linked to specific genes that are essential for life.
![]() Explain the steps, phase of the cell cycle, and the molecules responsible for the replication, repair, and recombination of DNA, and understand the negative effects of errors in any of these processes upon cellular and organismal integrity and health.
Explain the steps, phase of the cell cycle, and the molecules responsible for the replication, repair, and recombination of DNA, and understand the negative effects of errors in any of these processes upon cellular and organismal integrity and health.
BIOMEDICAL IMPORTANCE*
The genetic information in the DNA of a chromosome can be transmitted by exact replication or it can be exchanged by a number of processes, including crossing over, recombination, transposition, and conversion. These provide a means of ensuring adaptability and diversity for the organism but, when these processes go awry, can also result in disease. A number of enzyme systems are involved in DNA replication, alteration, and repair. Mutations are due to a change in the base sequence of DNA and may result from the faulty replication, movement, or repair of DNA and occur with a frequency of about one in every 106 cell divisions. Abnormalities in gene products (either in RNA, protein function, or amount) can be the result of mutations that occur in coding or regulatory-region DNA. A mutation in a germ cell is transmitted to offspring (so-called vertical transmission of hereditary disease). A number of factors, including viruses, chemicals, ultraviolet light, and ionizing radiation, increase the rate of mutation. Mutations often affect somatic cells and so are passed on to successive generations of cells, but only within an organism (ie, horizontally). It is becoming apparent that a number of diseases—and perhaps most cancers—are due to the combined effects of vertical transmission of mutations as well as horizontal transmission of induced mutations.
CHROMATIN IS THE CHROMOSOMAL MATERIAL IN THE NUCLEI OF CELLS OF EUKARYOTIC ORGANISMS
Chromatin consists of very long double-stranded DNA (dsDNA) molecules and a nearly equal mass of rather small basic proteins termed histones as well as a smaller amount of nonhistone proteins (most of which are acidic and larger than histones) and a small quantity of RNA. The nonhistone proteins include enzymes involved in DNA replication and repair, and the proteins involved in RNA synthesis, processing, and transport to the cytoplasm. The dsDNA helix in each chromosome has a length that is thousands of times the diameter of the cell nucleus. One purpose of the molecules that comprise chromatin, particularly the histones, is to condense the DNA; however, it is important to note that the histones also integrally participate in gene regulation (Chapters 36, 38, and 42); indeed histones contribute importantly to all DNA-directed molecular transactions. Electron microscopic studies of chromatin have demonstrated dense spherical particles called nucleosomes, which are approximately 10 nm in diameter and connected by DNA filaments (Figure 35–1). Nucleosomes are composed of DNA wound around a collection of histone molecules.
FIGURE 35–1 Electron micrograph of nucleosomes (white, ball-shaped) attached to strands of DNA (thin, gray line); see also Figure 35–2. (Reproduced, with permission, from Shao Z: Probing nanometer structures with atomic force microscopy. News Physiol Sci, 1999;14:142–149. Courtesy of Professor Zhifeng Shao, University of Virginia.)
Histones Are the Most Abundant Chromatin Proteins
Histones are a small family of closely related basic proteins. H1 histones are the ones least tightly bound to chromatin (Figures 35–1, 35–2, and 35–3) and are, therefore, easily removed with a salt solution, after which chromatin becomes more soluble. The organizational unit of this soluble chromatin is the nucleosome. Nucleosomes contain four major types of histones: H2A, H2B, H3, and H4. The structures of all four histones—H2A, H2B, H3, and H4, the so-called core histones that form the nucleosome—have been highly conserved between species, although variants of the histones exist and are used for specialized purposes. This extreme conservation implies that the function of histones is identical in all eukaryotes and that the entire molecule is involved quite specifically in carrying out this function. The carboxyl terminal two-thirds of the histone molecules are hydrophobic, while their amino terminal thirds are particularly rich in basic amino acids. These four core histones are subject to at least six types of covalent modification or posttranslational modifications (PTMs): acetylation, methylation, phosphorylation, ADP-ribosylation, monoubiquitylation, and sumoylation. These histone modifications play an important role in chromatin structure and function, as illustrated in Table 35–1.
TABLE 35–1 Possible Roles of Modified Histones
The histones interact with each other in very specific ways. H3 and H4 form a tetramer containing two molecules of each (H3–H4)2, while H2A and H2B form dimers (H2A–H2B). Under physiologic conditions, these histone oligomers associate to form the histone octamer of the composition (H3–H4)2–(H2A–H2B)2.
The Nucleosome Contains Histone & DNA
When the histone octamer is mixed with purified dsDNA under appropriate ionic conditions, the same X-ray diffraction pattern is formed as that observed in freshly isolated chromatin. Electron microscopic studies confirm the existence of reconstituted nucleosomes. Furthermore, the reconstitution of nucleosomes from DNA and histones H2A, H2B, H3, and H4 is independent of the organismal or cellular origin of the various components. Neither the histone H1 nor the nonhistone proteins are necessary for the reconstitution of the nucleosome core.
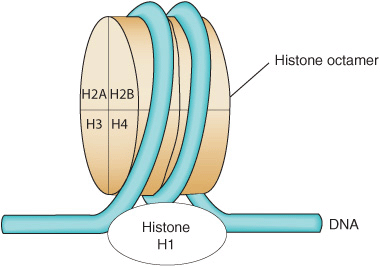
In the nucleosome, the DNA is supercoiled in a left-handed helix over the surface of the disk-shaped histone octamer (Figure 35–2). The majority of core histone proteins interact with the DNA on the inside of the supercoil without protruding, although the amino terminal tails of all the histones are thought to extend outside of this structure and are available for regulatory PTMs (see Table 35–1).
FIGURE 35–2 Model for the structure of the nucleosome, in which DNA is wrapped around the surface of a flat protein cylinder consisting of two each of histones H2A, H2B, H3, and H4 that form the histone octamer. The ~145 bp of DNA, consisting of 1.75 superhelical turns, are in contact with the histone octamer. The position of histone H1, when it is present, is indicated by the dashed outline at the bottom of the figure. Histone H1 interacts with DNA as it enters and exits the nucleosome.
The (H3–H4)2 tetramer itself can confer nucleosome-like properties on DNA and thus has a central role in the formation of the nucleosome. The addition of two H2A–H2B dimers stabilizes the primary particle and firmly binds two additional half-turns of DNA previously bound only loosely to the (H3–H4)2. Thus, 1.75 superhelical turns of DNA are wrapped around the surface of the histone octamer, protecting 145–150 bp of DNA and forming the nucleosome core particle (Figure 35–2). In chromatin, core particles are separated by an about 30-bp region of DNA termed “linker.” Most of the DNA is in a repeating series of these structures, giving the so-called beads-on-a-string appearance when examined by electron microscopy (see Figure 35–1).
The assembly of nucleosomes is mediated by one of several nuclear chromatin assembly factors facilitated by histone chaperones, a group of proteins that exhibit high-affinity histone binding. As the nucleosome is assembled, histones are released from the histone chaperones. Nucleosomes appear to exhibit preference for certain regions on specific DNA molecules, but the basis for this nonrandom distribution, termed phasing, is not yet completely understood. Phasing is likely related both to the relative physical flexibility of particular nucleotide sequences to accommodate the regions of kinking within the supercoil, as well as the presence of other DNA-bound factors that limit the sites of nucleosome deposition.
HIGHER ORDER STRUCTURES PROVIDE FOR THE COMPACTION OF CHROMATIN
Electron microscopy of chromatin reveals two higher orders of structure—the 10-nm fibril and the 30-nm chromatin fiber—beyond that of the nucleosome itself. The disk-like nucleosome structure has a 10-nm diameter and a height of 5 nm. The 10-nm fibril consists of nucleosomes arranged with their edges separated by a small distance (30 bp of DNA) with their flat faces parallel to the fibril axis (Figure 35–3). The 10-nm fibril is probably further supercoiled with six or seven nucleosomes per turn to form the 30-nm chromatin fiber (Figure 35–3). Each turn of the supercoil is relatively flat, and the faces of the nucleosomes of successive turns would be nearly parallel to each other. H1 histones appear to stabilize the 30-nm fiber, but their position and that of the variable length spacer DNA are not clear. It is probable that nucleosomes can form a variety of packed structures. In order to form a mitotic chromosome, the 30-nm fiber must be compacted in length another 100-fold (see below).
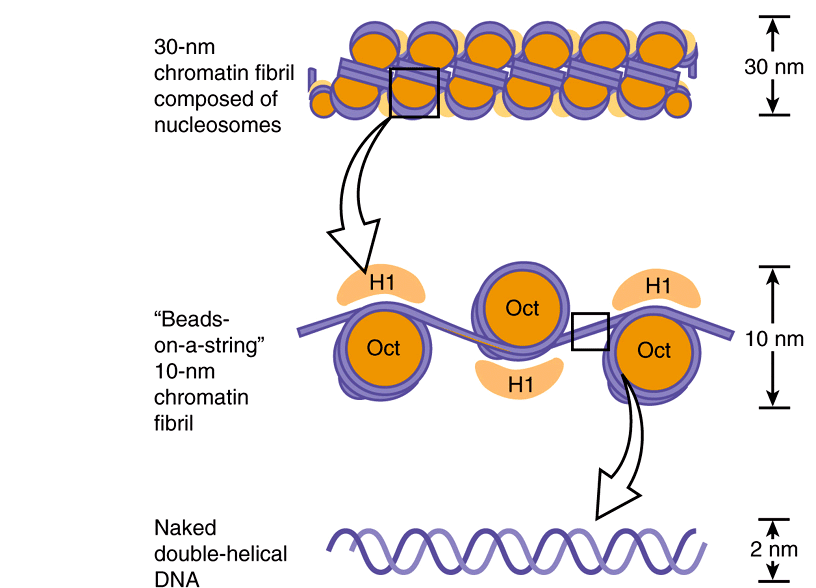
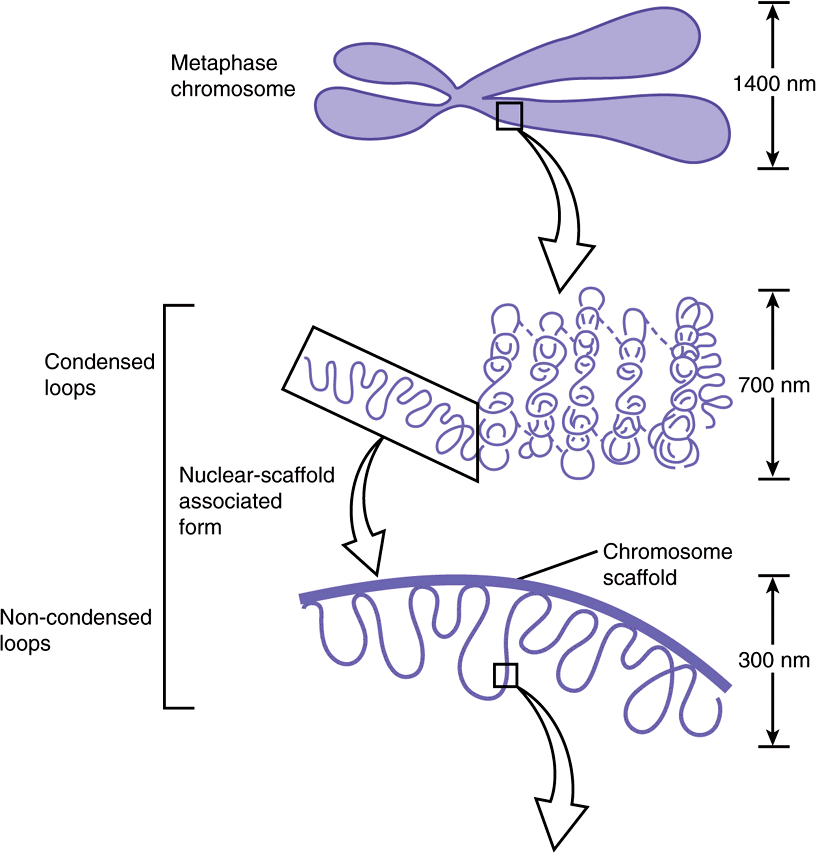
FIGURE 35–3 Shown is the extent of DNA packaging in metaphase chromosomes (top) to noted duplex DNA (bottom). Chromosomal DNA is packaged and organized at several levels as shown (see Table 35–2). Each phase of condensation or compaction and organization (bottom to top) decreases overall DNA accessibility to an extent that the DNA sequences in metaphase chromosomes are almost totally transcriptionally inert. In toto, these five levels of DNA compaction result in nearly a 104-fold linear decrease in end-to-end DNA length. Complete condensation and decondensation of the linear DNA in chromosomes occur in the space of hours during the normal replicative cell cycle (see Figure 35–20).
TABLE 35–2 The Packing or Compaction Ratios of Each of the Orders of DNA Structure
In interphase chromosomes, chromatin fibers appear to be organized into 30,000–100,000 bp loops or domains anchored in a scaffolding (or supporting matrix) within the nucleus, the so-called nuclear matrix. Within these domains, some DNA sequences may be located nonrandomly. It has been suggested that each looped domain of chromatin corresponds to one or more separate genetic functions, containing both coding and noncoding regions of the cognate gene or genes. This nuclear architecture is likely dynamic, having important regulatory effects upon gene regulation. Recent data suggest that certain genes or gene regions are mobile within the nucleus, moving obligatorily to discrete loci within the nucleus upon activation. Further work will determine both if this is a general phenomenon, and what molecular mechanisms are responsible.
SOME REGIONS OF CHROMATIN ARE “ACTIVE” & OTHERS ARE “INACTIVE”
Generally, every cell of an individual metazoan organism contains the same genetic information. Thus, the differences between cell types within an organism must be explained by differential expression of the common genetic information. Chromatin containing active genes (ie, transcriptionally or potentially transcriptionally active chromatin) has been shown to differ in several ways from that of inactive regions. The nucleosome structure of active chromatin appears to be altered, sometimes quite extensively, in highly active regions. DNA in active chromatin contains large regions (about 100,000 bases long) that are relatively more sensitive to digestion by a nuclease such as DNase I. DNase I makes single-strand cuts in nearly any segment of DNA (ie, low-sequence specificity). It will digest DNA that is not protected, or bound by protein, into its component deoxynucleotides. The sensitivity to DNase I of active chromatin regions reflects only a potential for transcription rather than transcription itself and in several systems can be correlated with a relative lack of 5-methyldeoxycytidine (meC) in the DNA and particular histone variants and/or PTMs (phosphorylation, acetylation, etc; see Table 35–1).
Within the large regions of active chromatin there exist shorter stretches of 100–300 nucleotides that exhibit an even greater (another 10-fold) sensitivity to DNase I. These hypersensitive sites probably result from a structural conformation that favors access of the nuclease to the DNA. These regions are often located immediately upstream from the active gene and are the location of interrupted nucleosomal structure caused by the binding of nonhistone regulatory transcription factor proteins (see Chapters 36 and 38). In many cases, it seems that if a gene is capable of being transcribed, it very often has a DNase-hypersensitive site(s) in the chromatin immediately upstream. As noted above, nonhistone regulatory proteins involved in transcription control and those involved in maintaining access to the template strand lead to the formation of hypersensitive sites. Such sites often provide the first clue about the presence and location of a transcription control element.
By contrast, transcriptionally inactive chromatin is densely packed during interphase as observed by electron microscopic studies and is referred to as heterochromatin; transcriptionally active chromatin stains less densely and is referred to as euchromatin. Generally, euchromatin is replicated earlier than heterochromatin in the mammalian cell cycle (see below). The chromatin in these regions of inactivity is often high in meC content, and histones therein contain relatively lower levels of covalent modifications.
There are two types of heterochromatin: constitutive and facultative. Constitutive heterochromatin is always condensed and thus essentially inactive. It is found in the regions near the chromosomal centromere and at chromosomal ends (telomeres). Facultative heterochromatin is at times condensed, but at other times it is actively transcribed and, thus, uncondensed and appears as euchromatin. Of the two members of the X chromosome pair in mammalian females, one X chromosome is almost completely inactive transcriptionally and is heterochromatic. However, the heterochromatic X chromosome decondenses during gametogenesis and becomes transcriptionally active during early embryogenesis—thus, it is facultative heterochromatin.
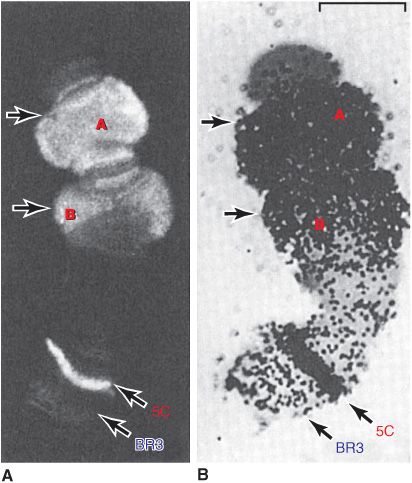
Certain cells of insects, for example, Chironomus and Drosophila, contain giant chromosomes that have been replicated for multiple cycles without separation of daughter chromatids. These copies of DNA line up side by side in precise register and produce a banded chromosome containing regions of condensed chromatin and lighter bands of more extended chromatin. Transcriptionally active regions of these polytene chromosomes are especially decondensed into “puffs” that can be shown to contain the enzymes responsible for transcription and to be the sites of RNA synthesis (Figure 35–4). Using highly sensitive fluorescently labeled hybridization probes, specific gene sequences can be mapped, or “painted,” within the nuclei of human cells, even without polytene chromosome formation, using FISH (fluorescence in situ hybridization; Chapter 39) techniques.
FIGURE 35–4 Illustration of the tight correlation between the presence of RNA polymerase II (Table 36–2) and messenger RNA synthesis. A number of genes, labeled A, B (top), and 5C, but not genes at locus (band) BR3 (5C, BR3, bottom) are activated when Chironomus tentans larvae are subjected to heat shock (39°C for 30 min). (A) Distribution of RNA polymerase II in isolated chromosome IV from the salivary gland (at arrows). The enzyme was detected by immunofluorescence using an antibody directed against the polymerase. The 5C and BR3 are specific bands of chromosome IV, and the arrows indicate puffs. (B) Autoradiogram of a chromosome IV that was incubated in 3H-uridine to label the RNA. Note the correspondence of the immunofluorescence and presence of the radioactive RNA (black dots). ![]() . (Reproduced, with permission, from Sass H: RNA polymerase B in polytene chromosomes. Cell 1982;28:274. Copyright © 1982. Reprinted with permission from Elsevier.)
. (Reproduced, with permission, from Sass H: RNA polymerase B in polytene chromosomes. Cell 1982;28:274. Copyright © 1982. Reprinted with permission from Elsevier.)
DNA IS ORGANIZED INTO CHROMOSOMES
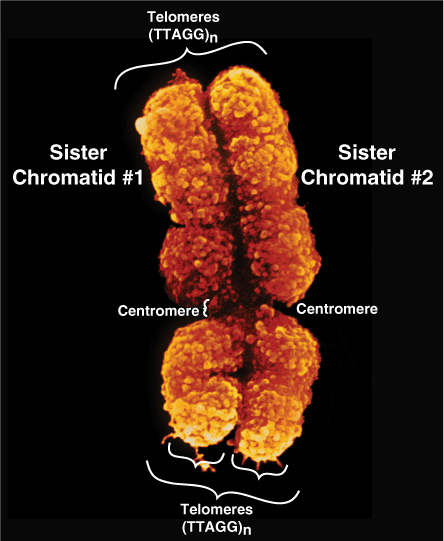
At metaphase, mammalian chromosomes possess a twofold symmetry, with the identical duplicated sister chromatids connected at a centromere, the relative position of which is characteristic for a given chromosome (Figure 35–5). The centromere is an adenine-thymine (A–T)-rich region containing repeated DNA sequences that range in size from 102 (brewers’ yeast) to 106 (mammals) base pairs (bp). Metazoan centromeres are bound by nucelosomes containing the histone H3 variant protein CENP-A and other specific centromere-binding proteins. This complex, called the kine-tochore, provides the anchor for the mitotic spindle. It thus is an essential structure for chromosomal segregation during mitosis.
FIGURE 35–5 The two sister chromatids of mitotic human chromosome 12. The location of the A+T-rich centromeric region connecting sister chromatids is indicated, as are two of the four telomeres residing at the very ends of the chromatids that are attached one to the other at the centromere. (Courtesy of Biophoto Associates/Photo Researchers, Inc.)
The ends of each chromosome contain structures called telomeres. Telomeres consist of short TG-rich repeats. Human telomeres have a variable number of repeats of the sequence 5′-TTAGGG-3’, which can extend for several kilobases. Telomerase, a multisubunit RNA template-containing complex related to viral RNA-dependent DNA polymerases (reverse transcriptases), is the enzyme responsible for telomere synthesis and thus for maintaining the length of the telomere. Since telomere shortening has been associated with both malignant transformation and aging, this enzyme has become an attractive target for cancer chemotherapy and drug development. Each sister chromatid contains one dsDNA molecule. During interphase, the packing of the DNA molecule is less dense than it is in the condensed chromosome during metaphase. Metaphase chromosomes are nearly completely transcriptionally inactive.
The human haploid genome consists of about 3 × 109 bp and about ![]() nucleosomes. Thus, each of the 23 chromatids in the human haploid genome would contain on the average
nucleosomes. Thus, each of the 23 chromatids in the human haploid genome would contain on the average ![]() nucleotides in one dsDNA molecule. Therefore, the length of each DNA molecule must be compressed about 8000-fold to generate the structure of a condensed metaphase chromosome. In metaphase chromosomes, the 30-nm chromatin fibers are also folded into a series of looped domains, the proximal portions of which are anchored to a nonhistone proteinaceous nuclear matrix scaffolding within the nucleus (Figure 35–3). The packing ratios of each of the orders of DNA structure are summarized in Table 35–2. The packaging of nucleoproteins within chromatids is not random, as evidenced by the characteristic patterns observed when chromosomes are stained with specific dyes such as quinacrine or Giemsa stain (Figure 35–6).
nucleotides in one dsDNA molecule. Therefore, the length of each DNA molecule must be compressed about 8000-fold to generate the structure of a condensed metaphase chromosome. In metaphase chromosomes, the 30-nm chromatin fibers are also folded into a series of looped domains, the proximal portions of which are anchored to a nonhistone proteinaceous nuclear matrix scaffolding within the nucleus (Figure 35–3). The packing ratios of each of the orders of DNA structure are summarized in Table 35–2. The packaging of nucleoproteins within chromatids is not random, as evidenced by the characteristic patterns observed when chromosomes are stained with specific dyes such as quinacrine or Giemsa stain (Figure 35–6).
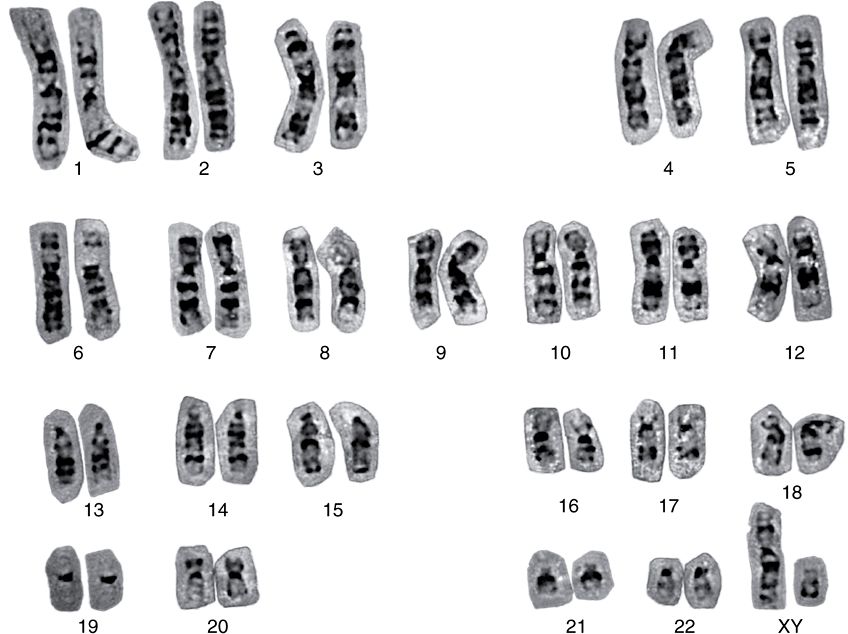
FIGURE 35–6 A human karyotype (of a man with a normal 46,XY constitution), in which the metaphase chromosomes have been stained by the Giemsa method and aligned according to the Paris Convention. (Courtesy of H Lawce and F Conte.)
From individual to individual within a single species, the pattern of staining (banding) of the entire chromosome complement is highly reproducible; nonetheless, it differs significantly between species, even those closely related. Thus, the packaging of the nucleoproteins in chromosomes of higher eukaryotes must in some way be dependent upon species-specific characteristics of the DNA molecules.
A combination of specialized staining techniques and high-resolution microscopy has allowed cytogeneticists to quite precisely map many genes to specific regions of mouse and human chromosomes. With the recent elucidation of the human and mouse genome sequences (among others), it has become clear that many of these visual mapping methods were remarkably accurate.
Coding Regions Are Often Interrupted by Intervening Sequences
The protein coding regions of DNA, the transcripts of which ultimately appear in the cytoplasm as single mRNA molecules, are usually interrupted in the eukaryotic genome by large intervening sequences of nonprotein-coding DNA. Accordingly, the primary transcripts of DNA, mRNA precursors, (originally termed hnRNA because this species of RNA was quite heterogeneous in size [length] and mostly restricted to the nucleus), contain noncoding intervening sequences of RNA that must be removed in a process which also joins together the appropriate coding segments to form the mature mRNA. Most coding sequences for a single mRNA are interrupted in the genome (and thus in the primary transcript) by at least one—and in some cases as many as 50—noncoding intervening sequences (introns). In most cases, the introns are much longer than the coding regions (exons). The processing of the primary transcript, which involves precise removal of introns and splicing of adjacent exons, is described in Chapter 36.
The function of the intervening sequences, or introns, is not totally clear. Introns may serve to separate functional domains (exons) of coding information in a form that permits genetic rearrangement by recombination to occur more rapidly than if all coding regions for a given genetic function were contiguous. Such an enhanced rate of genetic rearrangement of functional domains might allow more rapid evolution of biologic function. In some instances other protein or noncoding RNAs are localized within the intronic DNA of certain genes (Chapter 34). The relationships among chromosomal DNA, gene clusters on the chromosome, the exon–intron structure of genes, and the final mRNA product are illustrated in Figure 35–7.
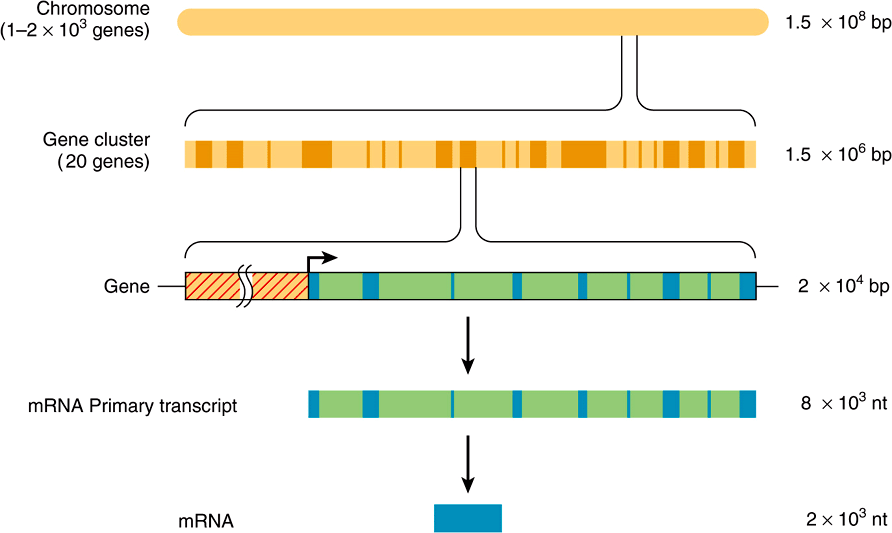
FIGURE 35–7 The relationship between chromosomal DNA and mRNA. The human haploid DNA complement of 3 × 109 bp is distributed between 23 chromosomes. Genes are often clustered on these chromosomes. An average gene is ![]() bp in length, including the regulatory region (red-hatched area), which is usually located at the 5′ end of the gene. The regulatory region is shown here as being adjacent to the transcription initiation site (arrow). Most eukaryotic genes have alternating exons and introns. In this example, there are nine exons (blue colored areas) and eight introns (green colored areas). The introns are removed from the primary transcript by the processing reactions, and the exons are ligated together in sequence to form the mature mRNA. (nt, nucleotides.)
bp in length, including the regulatory region (red-hatched area), which is usually located at the 5′ end of the gene. The regulatory region is shown here as being adjacent to the transcription initiation site (arrow). Most eukaryotic genes have alternating exons and introns. In this example, there are nine exons (blue colored areas) and eight introns (green colored areas). The introns are removed from the primary transcript by the processing reactions, and the exons are ligated together in sequence to form the mature mRNA. (nt, nucleotides.)
MUCH OF THE MAMMALIAN GENOME APPEARS REDUNDANT & MUCH IS NOT HIGHLY TRANSCRIBED
The haploid genome of each human cell consists of 3 × 109 bp of DNA subdivided into 23 chromosomes. The entire haploid genome contains sufficient DNA to code for nearly 1.5 million average-sized genes. However, studies of mutation rates and of the complexities of the genomes of higher organisms strongly suggest that humans have significantly fewer than 100,000 proteins encoded by the ~1% of the human genome that is composed of exonic DNA. Indeed current estimates suggest there are 25,000 or less protein-coding genes in humans. This implies that most of the DNA is nonprotein-coding—that is, its information is never translated into an amino acid sequence of a protein molecule. Certainly, some of the excess DNA sequences serve to regulate the expression of genes during development, differentiation, and adaptation to the environment, either by serving as binding sites for regulatory proteins or by encoding regulatory RNAs (ie, miRNAs and ncRNAs). Some excess clearly makes up the intervening sequences or introns (24% of the total human genome) that split the coding regions of genes, and another portion of the excess appears to be composed of many families of repeated sequences for which clear functions have not yet been defined, though some small RNAs transcribed from these repeats can modulate transcription, either directly by interacting with the transcription machinery or indirectly by affecting the activity of the chromatin template. A summary of the salient features of the human genome is presented in Chapter 39. Interestingly, the ENCODE Project Consortium (Chapter 39) has shown that for the 1% of the genome studied most of the genomic sequence was indeed transcribed at a low rate. Further research will elucidate the role(s) played by such transcripts.
The DNA in a eukaryotic genome can be divided into different “sequence classes.” These are unique-sequence DNA, or nonrepetitive DNA and repetitive-sequence DNA. In the haploid genome, unique-sequence DNA generally includes the single copy genes that code for proteins. The repetitive DNA in the haploid genome includes sequences that vary in copy number from 2 to as many as 107 copies per cell.
More Than Half the DNA in Eukaryotic Organisms Is in Unique or Nonrepetitive Sequences
This estimation (and the distribution of repetitive-sequence DNA) is based on a variety of DNA–RNA hybridization techniques and, more recently, on direct DNA sequencing. Similar techniques are used to estimate the number of active genes in a population of unique-sequence DNA. In brewers’ yeast (Saccharomyces cerevisiae,
Stay updated, free articles. Join our Telegram channel

Full access? Get Clinical Tree