CHAPTER 8 Membrane Pumps
Membrane Permeability: An Introduction
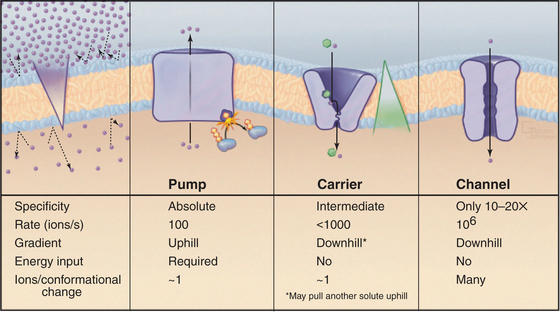
Although lipid bilayers provide a barrier to diffusion of ions and polar molecules larger than about 150 D, protein pores provide selective passages for ions, and other larger molecules across membranes. Integral proteins that control membrane permeability fall into three broad classes—pumps, carriers, and channels—each with distinct properties (Fig. 8-1). These proteins allow cells to control solute traffic across membranes, an essential feature of many physiological processes.
This chapter and Chapters 9, 10, and 11 consider, in turn, the three classes of proteins that control membrane permeability. Pumps are discussed first because they create the solute gradients required for the function of carriers and channels. The concluding chapter in this section, Chapter 11, illustrates how pumps, carriers, and channels work together to perform a remarkable variety of functions. An impor-tant point is that differential expression of a subset of isoforms of these proteins in specific membranes allows differentiated cells to perform a wide range of complex functions.
Membrane Pumps
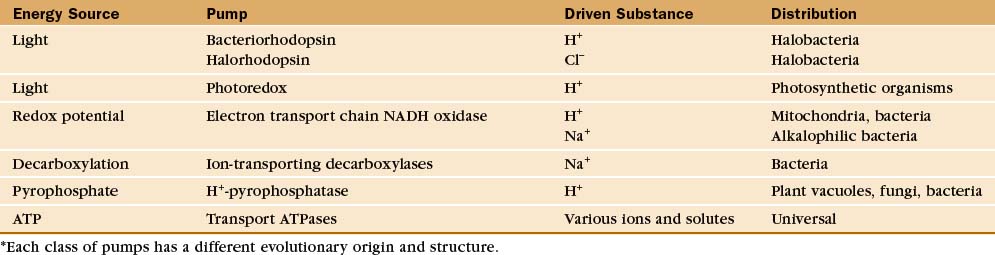
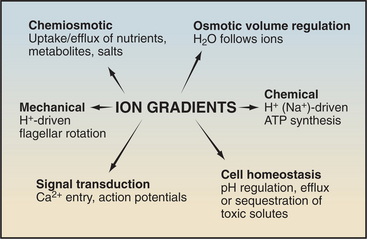
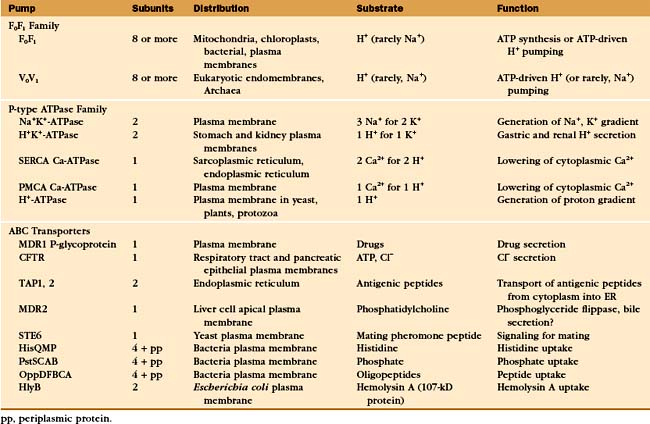
Protein pumps transport ions and other solutes across membranes up concentration gradients as great as 1 million-fold. Energy for this task can come from a variety of sources: light, oxidation-reduction reactions, or, most commonly, hydrolysis of ATP (Table 8-1). Energy is conserved in the form of transmembrane electrical or chemical gradients of the transported ion or solute. The potential energy in these ion gradients drives a variety of energy-requiring processes (Fig. 8-2). Most known biological pumps translocate cations. Although they could just as well move anions, cations were selected during the evolution of early life forms 3 billion years ago.
Pumps are also called primary active transporters because they transduce electromagnetic or chemical energy directly into transmembrane concentration gradients. Some carriers use ion gradients created by pumps to drive the uphill movement of other ions or solutes, so these are called secondary transporters (see Chapter 9). Channels are passive transporters, allowing net diffusion of ions and water only down their concentration gradients (see Chapter 10).
Diversity of Membrane Pumps
A vast array of integral membrane proteins can capture energy from an external source to pump ions and other solutes across biological membranes (Table 8-1). The protein families differ in their energy sources and transported materials. Fortunately, these pumps had a limited number of common ancestors, providing a relatively simple classification and generalizations about their structures and mechanisms. Given the importance of pumps in establishing transmembrane electrochemical gradients, the simplicity of this list is remarkable. Its brevity may be attributable to the fact that a single pump can drive a whole host of secondary reactions mediated by different carriers.
This chapter considers four representative pumps, emphasizing examples in which both high-resolution structures and detailed biochemical analysis of pathways are available. Chapter 19 provides additional details on H+ translocation by redox-driven cytochrome c oxidase and the role of F-type pumps in ATP synthesis by mitochondria and chloroplasts. Microbiology texts provide more information on pumps driven by decarboxylases and pyrophosphatases.
Light-Driven Proton Pumping by Bacteriorhodopsin
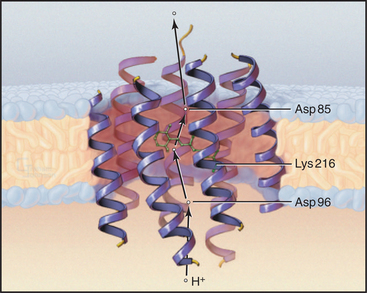
Owing to its simplicity, its small size, and the availability of a high-resolution structure (Fig. 8-3), more is known about light-driven transport of protons by bacteriorhodopsin than about any other pump. This pump allows the halophilic (salt-loving) Archaea Halobacterium halobium to convert light energy into a proton gradient across its plasma membrane. The 26-kD pump packs into two-dimensional crystalline arrays in the plasma membrane. The polypeptide is folded into seven α-helices that cross the lipid bilayer. The light-absorbing chromophore retinal (vitamin A aldehyde) is bound covalently to the side chain of lysine 216 (Lys216) via a Schiff base. This chromophore makes the protein and the membrane purple.
Bacteriorhodopsin absorbs light and uses the energy to pump protons out of the cell. A proton-driven ATP synthase uses this proton gradient to make ATP (Fig. 8-5). The proton pathway includes the side chains of aspartate 96 (Asp96), aspartate 85 (Asp85), glutamate 204 (Glu204), and the Schiff base. Local environments give the two aspartates remarkably different ionization constant (p Ka) values. Asp96 has a very high p Ka of about 10, so it can serve as a proton donor. Asp85 has a low p Ka of about 2, so it serves as a proton acceptor. Absorption of a photon changes the conformation of the retinal and the p Ka of the Schiff base. These four groups work together to transfer a single proton from the cytoplasm to the extracellular space.
The net result of this cycle is rapid vectorial transport of a proton from the cytoplasm out of the cell. Steps 4 to 6 are rate limiting, occurring at a rate of about 100 s−1. The other reactions are fast, provided that there is an adequate flux of light. Retinal not only captures energy by absorbing a photon but also acts as a switch that changes both the accessibility and affinities of the proton-binding groups in a sequential fashion.
In addition to bacteriorhodopsin, halobacterial plasma membranes contain two related proteins: halorhodopsin and sensory rhodopsin. Halorhodopsin absorbs light and pumps chloride into the cell. Interestingly, a single amino acid substitution can reverse the direction of pumping. Sensory rhodopsin couples light absorption by its bound retinal to phototaxis (swimming toward light) with a tightly coupled transducer protein. In the absence of this transducer, sensory rhodopsin transports protons out of the cell much like bacterial rhodopsin. The design of these seven-helix transporters is remarkably similar to that of the large family of seven-helix receptors, especially the photoreceptor proteins that vertebrates use for vision (see Fig. 24-2).
ATP-Driven Pumps
Three families of transport ATPases (Table 8-2) are essential for the physiology of all forms of life. F0F1-ATPases and P-type ATPases differ in structure, but both generate electrical and/or chemical gradients across membranes. ABC transporters not only produce ion gradients but also transport a much wider range of solutes across membranes. Chemical inhibitors have been useful in characterizing these pumps, and some are also used therapeutically (Table 8-3).
Table 8-3 TOOLS FOR STUDYING PUMPS
| Agent | Target |
|---|---|
| Cardiac glycosides* (e.g., ouabain, digitalis) | Na+K+ -ATPase |
| Omeprazole* | H+K+ -ATPase (parietal cell) |
| Oligomycin | F0F1-ATP synthase |
Free energy released by ATP hydrolysis puts a limit on the concentration gradient that these pumps can produce. If transport is electrically neutral (i.e., if it does not produce a membrane potential; see Fig. 10-17), the maximum gradient is about 1 million-fold. Such an extraordinary gradient is actually created by the P-type, electrically neutral H+K+−ATPase of gastric epithelial cells, which acidifies the stomach down to a pH of 1.
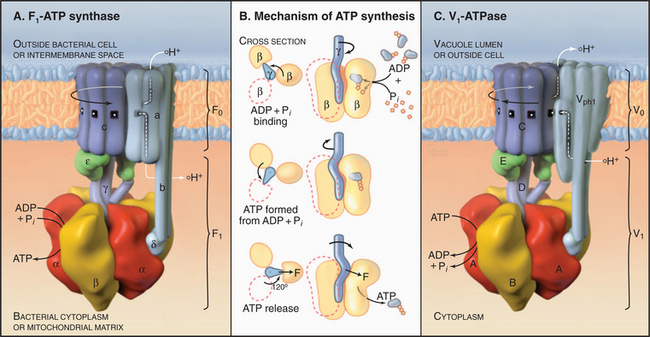
F0F1-ATPase Family
The two major subdivisions of this family are called F0F1-ATPases (or F-type ATPases) and V0V1-ATPases (or V-type ATPases) (Figs. 8-4 and 8-5). V0V1-ATPases, named for their location in the vacuolar system of eukaryotes, pump protons into organelles and out of Archaea. F0F1-ATPases of Bacteria, mitochondria, and chloroplasts generally run in the opposite direction, using proton gradients generated by other membrane proteins to drive ATP synthesis. However, purified F0F1-ATPases are freely reversible, using ATP hydrolysis to pump protons or alternatively proton gradients to synthesize ATP. Hence, these enzymes are called both ATP-synthases and F-type ATPases.
Phylogenetic analysis of the subunit polypeptides traces the origin of V-type ATPases to the precursor of all contemporary life forms (see Fig. 1-1
Stay updated, free articles. Join our Telegram channel

Full access? Get Clinical Tree