CHAPTER 12 Chromosome Organization
Chromosomes are enormous DNA molecules that can be propagated stably through countless generations of dividing cells (Fig. 12-1). Genes are the reason for the existence of the chromosomes, but in higher eukaryotes, they actually make up only a small fraction of the chromosomal DNA, much of which does not encode proteins or other known functional RNAs. Cells package chromosomal DNA with roughly twice its weight of protein. This DNA-protein complex, called chromatin, is discussed in Chapter 13.
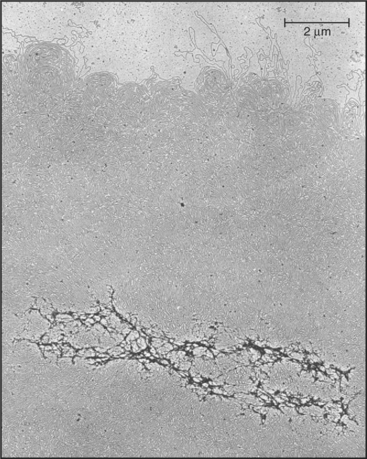
(From Paulson JR, Laemmli UK: The structure of histone-depleted chromosomes. Cell 12:817–828, 1977.)
In addition to the genes, only three classes of specialized DNA sequences are needed to make a fully functional chromosome: (1) a centromere, (2) two telomeres, and (3) an origin of DNA replication for approximately every 100,000 base pairs (bp). Centromeres regulate the partitioning of chromosomes during mitosis and meiosis. Telomeres protect the ends of the chromosomal DNA molecules and ensure their complete replication. DNA replication is discussed in Chapter 42. Chapter 15 considers the structure of genes. Box 12-1 lists a number of key terms presented in this chapter.
BOX 12-1 Key Terms
Chromosome Morphology and Nomenclature
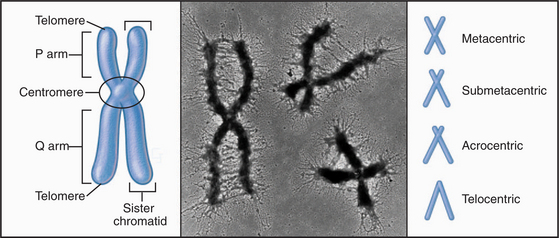
With few specialized exceptions, chromosomes from somatic cells of higher eukaryotes are visualized directly only during mitosis. Each mitotic chromosome consists of two sister chromatids that are held together at a waist-like constriction called the centromere. The portions of the chromosomes that are not in the centromere itself are called chromosome “arms” (Fig. 12-2).
One DNA Molecule per Chromosome
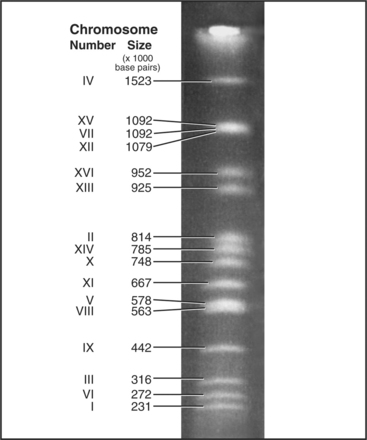
Each eukaryotic chromosome contains one DNA molecule that stretches between the telomeres at either end. Most prokaryotic and mitochondrial chromosomes are circular DNA molecules that lack telomeres, but naturally occurring eukaryotic nuclear chromosomes are generally linear DNA molecules with two telomeres. The clearest proof that each chromosome is composed of a single DNA molecule has been obtained for budding and fission yeasts, where intact chromosomal DNA molecules may be visualized by pulsed-field gel electrophoresis as a characteristic series of bands (Fig. 12-3). This technique can display the largest chromosome of fission yeast at 5,598,923 bp, but even the smallest human chromosome, which is about 40 million bp long, is too large to resolve in this way.
The Organization of Genes on Chromosomes
The first chromosome to be completely sequenced (in 1977) was that of the bacterial virus φx174 (Table 12-1). Starting in the 1990s much effort worldwide has been devoted to determining the complete sequences of the chromosomes of a wide variety of organisms (see Fig. 2-4). Sequencing efforts that have been completed to date have generated an enormous bank of data on the genetic composition of simple and complex organisms. For example, over 100 microbial genomes have been sequenced. One major goal of this effort—the sequence of the human genome—is now essentially complete.
Table 12-1 DNA CONTENT OF VARIOUS GENOMES
| Organism | Haploid Genome Size (bp) | Predicted Number of Protein-Coding Genes |
|---|---|---|
| φX174 (bacterial virus) | 5386 | 11 |
| Mycoplasma genitalium (pathogenic bacterium) | 580,070 | 480* |
| Rickettsia prowazekii (endoparasitic bacterium) | 1,111,523 | 834 |
| Escherichia coli (free-living bacterium) | 4,639,221 | 4288 |
| Bacillus subtilis (free-living bacterium) | 4,214,810 | 4100 |
| Saccharomyces cerevisiae (budding yeast) | 14,000,000 | 6604 |
| Schizosaccharomyces pombe (fission yeast) | 13,800,000 | 4824 |
| Caenorhabditis elegans (nematode worm) | 9.7 × 107 | 19,100 |
| Drosophila melanogaster (fruit fly) | 1.4 × 108 | 13,525 |
| Arabadopsis thaliana (plant) | 1.25 × 108 | 25,498 |
| Anopheles gambiae (malaria mosquito) | 2.78 × 108 | 14,000 |
| Oryza sativa japonica (rice) | 4.2 × 108 | 32,000–50,000 |
| Mus musculus (house mouse) | 2.6 × 109 | −30,000 |
| Rattus norvegicus (Brown Norway rat) | 2.75 × 109 | −21,000–46,000 |
| Xenopus laevis (South African clawed frog) | 3.1 × 109 | ? |
| Homo sapiens (human) | 3.1 × 109 | 20,000–25,000 |
| Triturus cristatus (salamander) | 2.2 × 1010 | ? |
Note: In most higher eukaryotes, with the exception of some plants, the huge tracts of repeated DNA sequences in and around centromeres are poor in genes and beyond the limits of present technology to sequence. Thus, when statistics are given on chromosome sizes in descriptions of genome sequencing projects, these portions are generally omitted. Where possible, the genome size figures given here reflect the entire genome (sequenced and unsequenced).
* It appears that only 265 to 350 of these genes are essential for life.
Complex genomes that have been sequenced thus far range in size from 580,000 bp for Mycoplasma genitalium, which causes urinary tract infections in humans to 2,863,476,365 bp for humans themselves. Numbers of protein-coding genes identified range from 480 in M. genitalium to 20,000 to 25,000 for humans (Table 12-1). However, because gene prediction algorithms are still being perfected, only rough estimates of gene number are available, even for completely sequenced genomes.
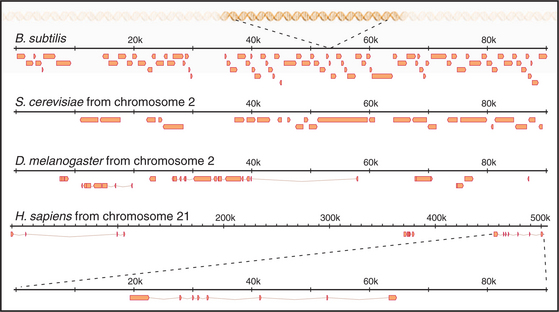
The first eukaryote whose genome was entirely sequenced was the budding yeast Saccharomyces cerevisiae. The 14 million bp yeast genome is subdivided into 16 chromosomes ranging in size from 230,000 bp to over 1 million bp (Fig. 12-3). This genome has a dramatic history. Ancestral budding yeast apparently had eight chromosomes but at one point underwent a duplication of the entire genome. This event was followed by numerous small deletions that resulted in the subsequent loss of most of the duplicated genes, with about 10% remaining. As a result, the modern budding yeast genome contains about 5700 predicted genes, many of which are paralogs (genes produced by duplication that have evolved to take on distinct functions; see Box 2-1). As a result, only about 1000 of these genes are indispensable for life. About 5% of yeast genes are segmented, containing regions that appear in mature RNA molecules (exons) and regions that are removed by splicing (introns) (discussed in detail in Chapter 16). Exons occupy approximately 75% of the budding yeast genome, with the remainder in regulatory regions, repeated DNAs, and introns (Fig. 12-4).
The next genome sequences to be completed were those of two very important “model” organisms that have been widely used by cell and developmental biologists: the nematode worm Caenorhabditis ele-gans and the fruit fly Drosophila melanogaster. These sequences revealed a number of important organizational differences from budding yeast. Although its ge-nome is eight times larger than that of budding yeast (97 million bp distributed in six chromosomes), the nematode has only about three times more genes. Surprisingly, the fly, despite its even larger genome and more complex body plan and life cycle, has about one third fewer genes than the worm. In fact, only about 27% of the C. elegans genome and 13% of the Drosophila genomic DNA code for proteins. Instead, the fly has much more noncoding repetitive DNA than the worm.
The “finished” sequence of the human genome, published in 2004, revealed an even lower density of genes. Humans have far fewer genes than had been predicted: about 20,000 to 25,000, in contrast to some earlier predictions of up to 100,000 (Table 12-1). Protein-coding regions occupy only about 1.2% of the chromosomes. In contrast, various repeated-sequence elements and pseudogenes appear to occupy about 50% of the genome, as is discussed in a later section. To put this all in perspective, every million bp of DNA sequenced yielded 483 genes in S. cerevisiae, 197 genes in C. elegans, 117 genes in D. melanogaster, and only 7 to 9 genes in humans. If the Escherichia coli chromosome were the size of chromosome 21, the smallest human chromosome at ˜40 × 106 bp, it would have nearly 37,000 genes—more than the entire human complement! In fact, chromosome 21 is predicted to have only 225 genes.
Transposons Make Up Much of the Human Genome
Even though humans no longer have active transposons, we still use at least two functional vestiges of these elements. It has been known for years that one of the ways in which the diversity of the immune system is generated is by cutting and pasting portions of the genes that encode the variable regions of the immunoglobulin chains (see Fig. 28-10). This process involves moving bits of DNA around, and it now appears that the enzymes that accomplish this process were originally encoded by ancient transposons. In addition, CENP-B (centromere protein B; see Fig. 13-23), an abundant protein that binds to the α-satellite DNA repeats in primate centromeres, is closely related to a transposase enzyme encoded by one family of transposons.
The best-known retrotransposons are LINES (long interspersed nuclear elements) and SINES (short interspersed nuclear elements). Reverse transcriptases encoded by LINES are responsible for movements of both LINES and SINES. The L1 class of LINES encodes two proteins, one of which has reverse transcriptase activity (Fig. 12-5). All DNA polymerases, including reverse transcriptases, work by elongating a preexisting stretch of double-stranded nucleic acid (see Chapter 42 for a discussion of the mechanism of DNA synthesis). L1 elements insert themselves into the chromosome by first nicking the chromosomal DNA, then using the newly created end as a primer for synthesis of a new DNA strand (Fig. 12-5). The template for this DNA synthesis by the reverse transcriptase is the LINE RNA, and the newly synthesized DNA is made as a direct extension of the chromosomal DNA molecule. Most LINES are only partial copies of the full-length element. Apparently, the reverse transcriptase is not very efficient (processive): It usually falls off before it completes copying the entire element.
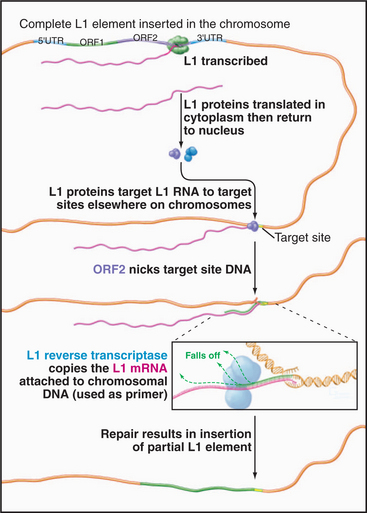
Figure 12-5 mechanism of transposition of an l1 element. The element is transcribed by RNA polymerase II (see Fig. 15-4). Proteins encoded by the element nick the chromosome, promote base pairing of the L1 transcript with the target site, and reverse transcribe the RNA into DNA. The L1 DNA is synthesized as an extension of the chromosome. The mechanism of final closing up of the nicks and gaps is not yet fully understood.
LINES and SINES plus other remnants of transposable elements account for up to 45% of the human genome. LINES, with a consensus sequence of 6 to 8 kb, make up about 20% of the genome. (A consensus sequence is the average arrived at by comparing a number of different sequenced DNA clones.) About 79% of human genes have at least one segment of L1 sequence inserted, typically in an intron. The Alu class of SINES, with a consensus sequence of about 300 bp, constitutes about 13% of the total DNA—almost a million copies scattered throughout the genome. Alu elements are derived from the 7SL RNA gene, which encodes the RNA component of signal recognition particle (see Fig. 20-5). They are actively transcribed by RNA polymerase III (see Fig. 15-10
Stay updated, free articles. Join our Telegram channel

Full access? Get Clinical Tree