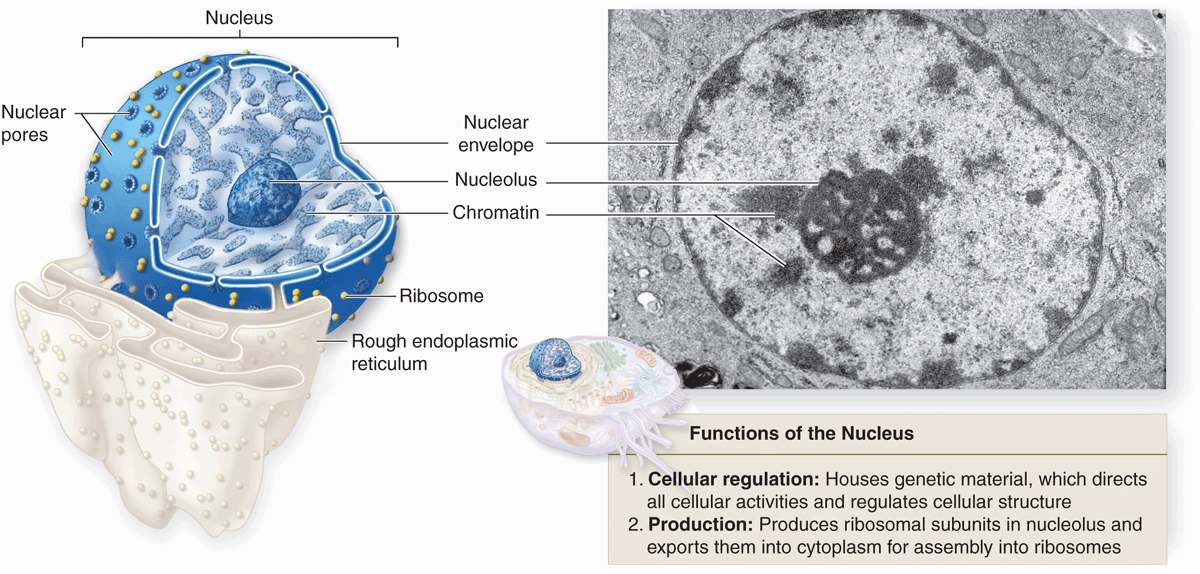
Containing the code for all of a cell’s enzymes and other proteins, the nucleus is the command center of the cell. The nucleus also contains the molecular machinery to replicate the DNA and to synthesize and process all types of RNA. During interphase pore complexes in the membrane enclosing the nucleus regulate macromolecular transfer between the nuclear and cytoplasmic compartments. Mature RNA molecules pass into the cytoplasm for their roles in protein synthesis, while proteins needed for nuclear activities are imported from the cytoplasm. Restricting protein synthesis to the cytoplasm helps ensure that newly made RNA molecules do not become involved in translation before processing is complete.
COMPONENTS OF THE NUCLEUS
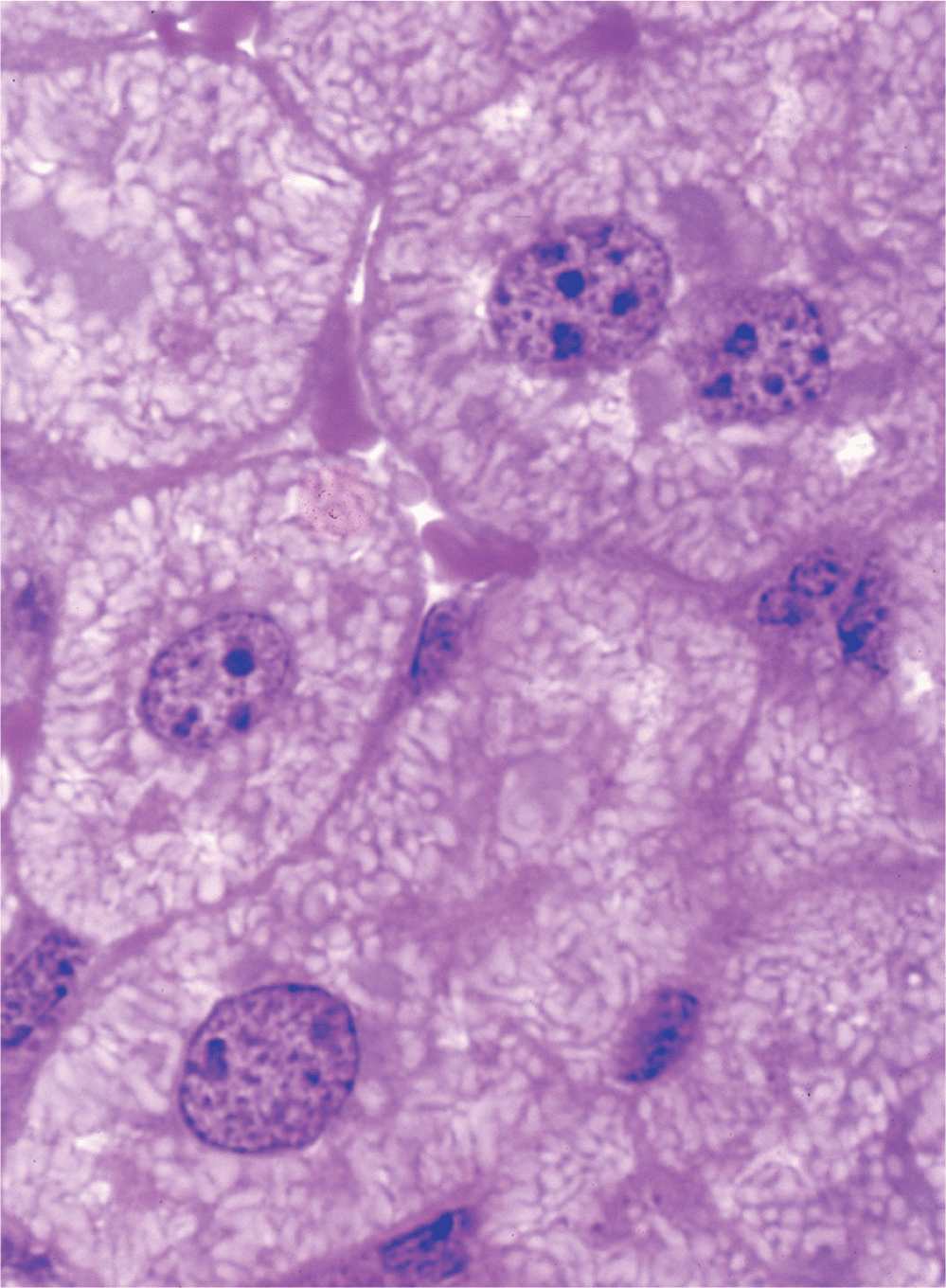
The nucleus frequently appears as a large rounded or oval structure, often near the center of the cell (Figure 3–1). Consisting of a nuclear envelope, a mass of DNA and associated proteins called chromatin, and a specialized subdomain called the nucleolus, the nucleus is typically the largest structure within a cell. In specific tissues the size and morphologic features of nuclei normally tend to be uniform.
FIGURE 3–1 Nuclei of large, active cells.
Nuclear Envelope
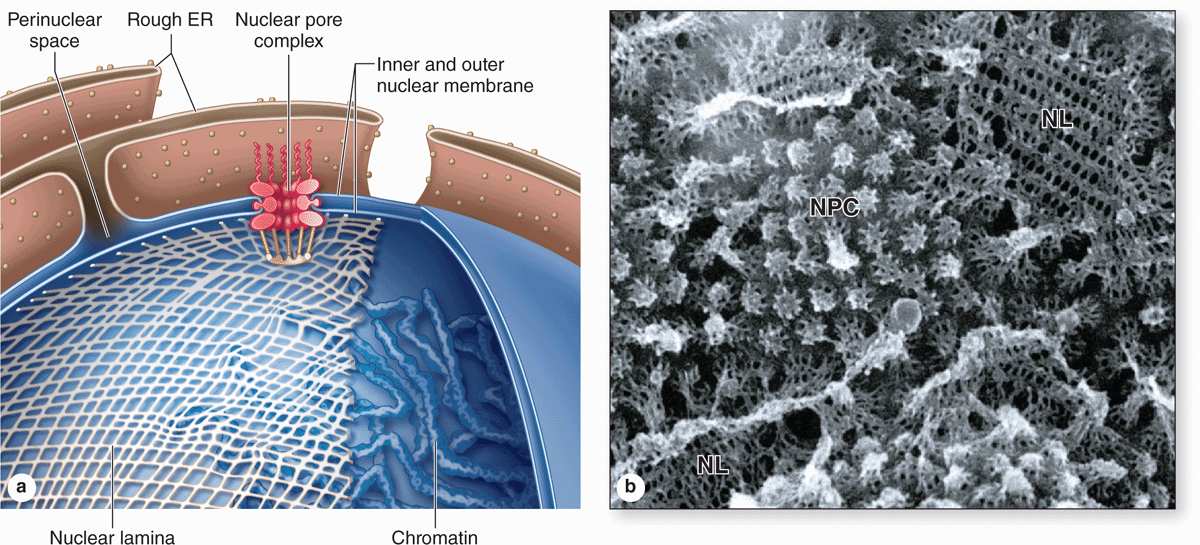
The nuclear envelope forms a selectively permeable barrier between the nuclear and cytoplasmic compartments. Electron microscopy reveals that the envelope has two concentric membranes separated by a narrow (30-50 nm)perinuclear space (Figure 3–2). This space and the outer nuclear membrane are continuous with the extensive cytoplasmic network of the rough endoplasmic reticulum (RER). Closely associated with the inner nuclear membrane is a highly organized meshwork of proteins called the nuclear lamina (Figure 3–4), which stabilizes the nuclear envelope. Major components of this layer are the class of intermediate filament proteins called lamins that bind to membrane proteins and associate with chromatin in nondividing cells.
FIGURE 3–2 Relationship of nuclear envelope to the rough ER (RER).
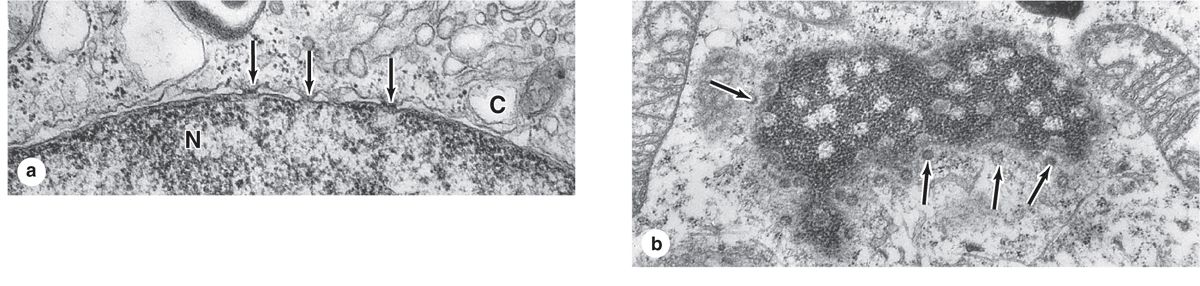
The inner and outer nuclear membranes are bridged at nuclear pore complexes (Figures 3–2 through 3–6). Various core proteins of a nuclear pore complex, called nucleoporins, display eightfold symmetry around the lumen. Although ions and small solutes pass through the channels by simple diffusion, the pore complexes regulate movement of macromolecules between the nucleus and cytoplasm. A growing cell has 3000-4000 such channels, each providing passage for up to 1000 macromolecules per second. Individual pores permit molecular transfer in both directions simultaneously. Macromolecules shipped out of the nucleus include ribosomal subunits and other RNAs associated with proteins, while inbound traffic consists of chromatin proteins, ribosomal proteins, transcription factors, and enzymes. Using mechanisms similar to that by which specific proteins are recognized and translocated across the RER membrane, proteins of complexes destined for the cytoplasm have specific nuclear export sequences and proteins to be imported have nuclear localization sequences. Such sequences bind specifically to transport proteins (importins, exportins, etc) that in turn interact with proteins of the pore complexes for transfer across the nuclear envelope. Energy for the transport is derived from guanosine 5’-triphosphate (GTP), with specific GTPases helping provide directionality to the transfer.
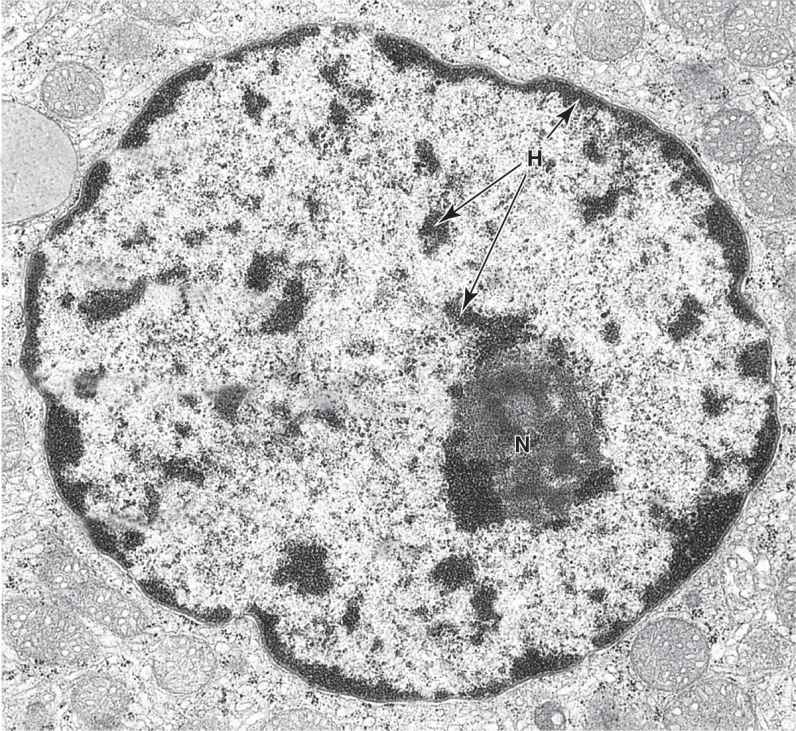
FIGURE 3–3 Ultrastructure of a nucleus.
FIGURE 3–4 The nuclear envelope, nuclear lamina, and nuclear pore complexes.
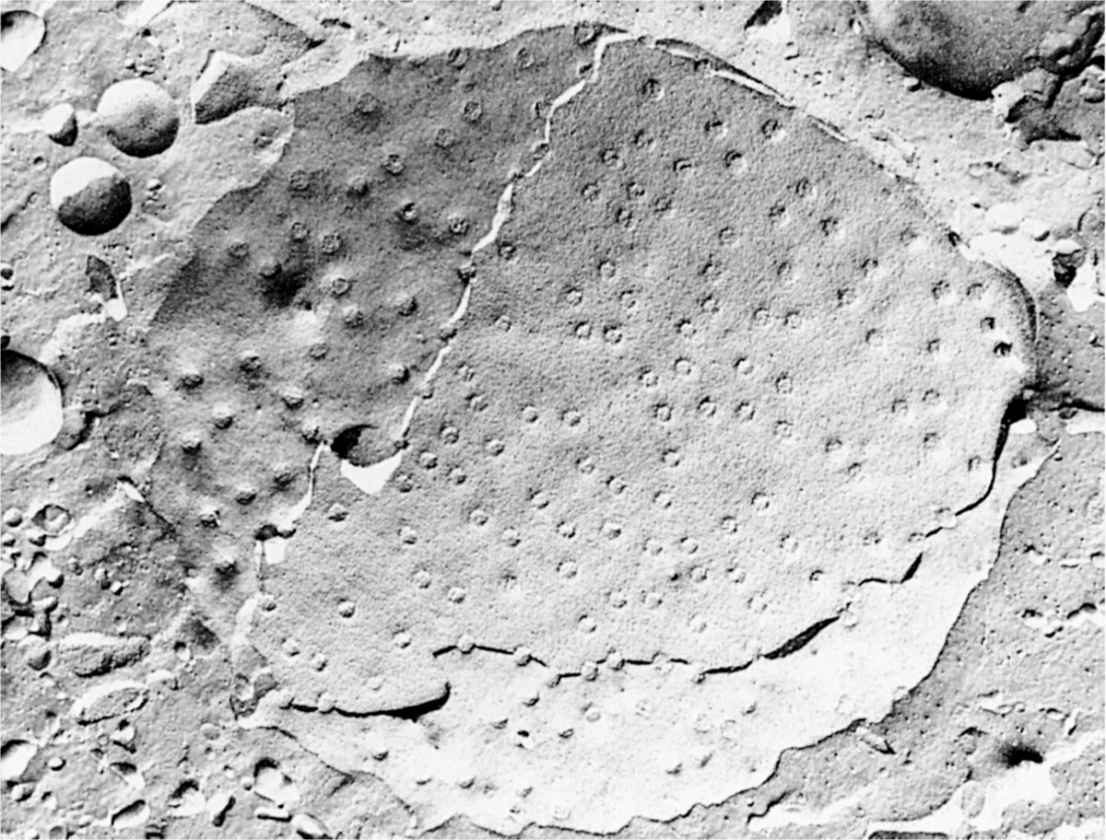
FIGURE 3–5 Nuclear pores.
FIGURE 3–6 Cryofracture of nuclear envelope showing nuclear pores.
Chromatin
In nondividing nuclei, chromatin consists of the DNA and its attendant proteins in a largely uncoiled state. Two types of chromatin can be distinguished with both the light and electron microscopes (Figures 3–2).Heterochromatin (Gr. heteros, other + chroma, color) appears as coarse, electron-dense material in the electron microscope and as intensely basophilic clumps in the light microscope.Euchromatin is visible as finely dispersed granular material in the electron microscope and as lightly stained basophilic areas in the light microscope. The dispersed euchromatin contains regions of the DNA undergoing active transcription and is more prominent in metabolically active cells.
The chromatin pattern of a nucleus is a guide to the cell’s activity. Generally cells with lightly stained nuclei are more active in protein synthesis than those with condensed, dark nuclei. In light-stained nuclei with much euchromatin and few heterochromatic clumps, more DNA surface is available for the transcription of RNA. In dark-stained nuclei rich in highly condensed heterochromatin, the tightly coiled DNA is less accessible for transcription.
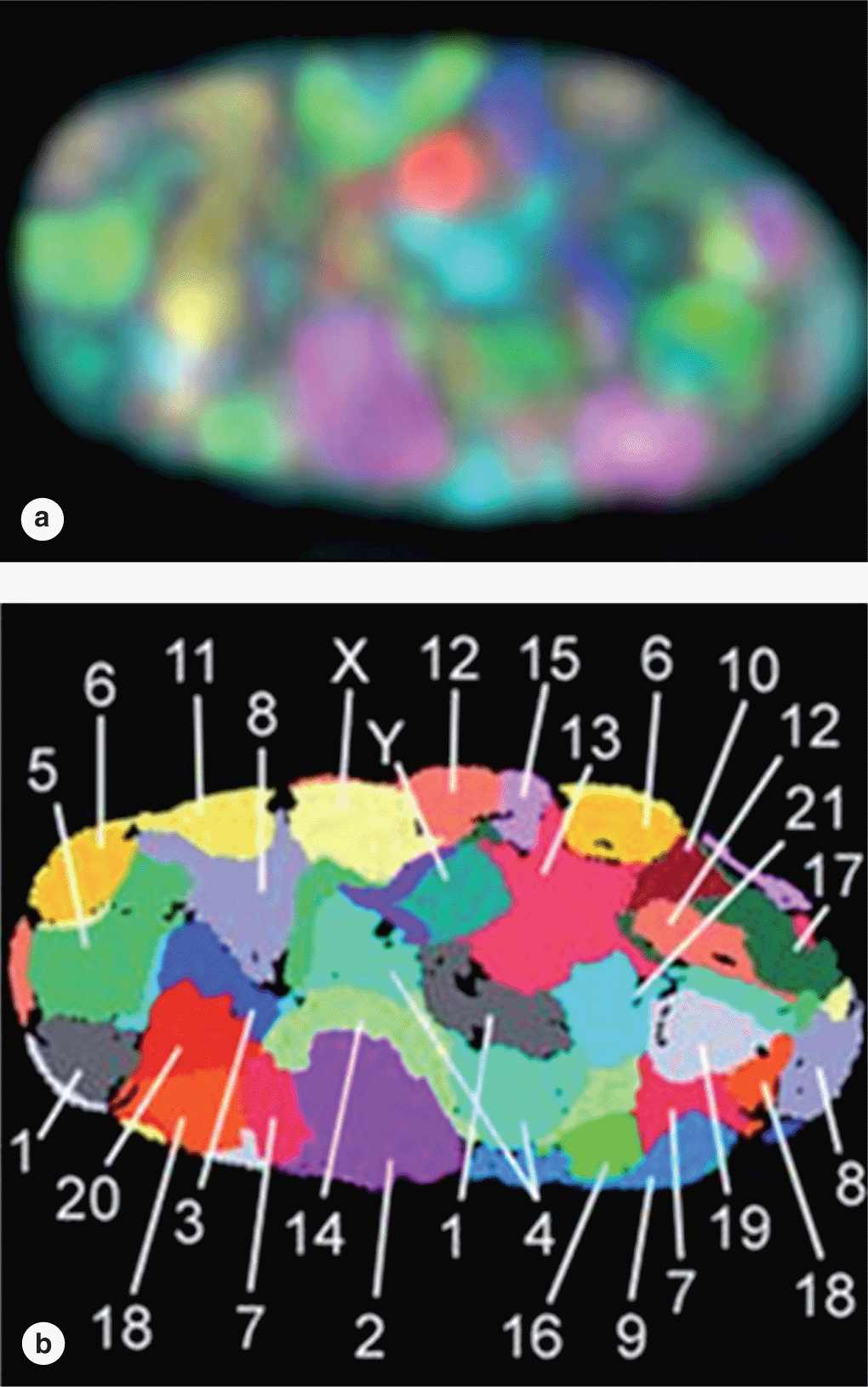
Although heterochromatin tends to be concentrated near the nuclear lamina, evidence for spatial organization of chromatin is not normally seen. Recent in situ hybridization studies of cultured human fibroblast nuclei, using a differently labeled fluorescent probes for sequences on each individual chromosome, have revealed that these structures occupy discrete chromosomal territories within dispersed chromatin (Figure 3–7). Such studies show further that chromosomal domains with few genes form a layer beneath the nuclear envelope, while domains with many active genes are located deeper in the nucleus.
FIGURE 3–7 Chromosome territories of a human fibroblast nucleus.
Routine microscopy of mammalian cell nuclei reveals a small, dense mass of heterochromatin present in females but not males. This “sex chromatin” (or Barr body) is one of the two X chromosomes present in females. This X chromosome represents facultative heterochromatin and remains tightly coiled between mitoses, while the other X chromosome is uncoiled, transcriptionally active, and not visible. The male cell has one X chromosome and one Y chromosome; like the other chromosomes, the interphase X chromosome is uncoiled and therefore no sex chromatin is visible in males.
Formation of facultative heterochromatin involves specific chemical modifications of chromatin proteins and usually occurs only within specific regions of a chromosome, with variability in cells of different tissues. Constitutive heterochromatin on the other hand is also specifically modified but is always located at the same sites, such as the centromere region of every chromosome.
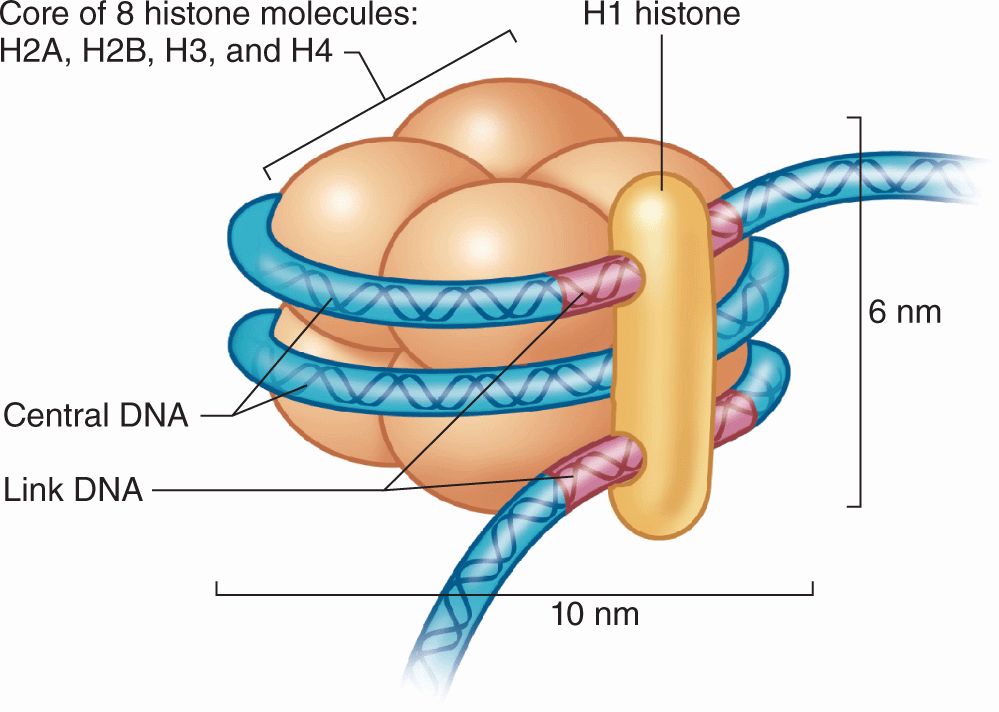
DNA in chromatin is extensively packaged by associating with basic proteins called histones and with various nonhistone proteins. The structural unit of DNA and histones is the nucleosome (Figure 3–8), which has a core of eight small histones (two copies each of histones H2A, H2B, H3, and H4), around which is wrapped DNA with about 150 base pairs. Each nucleosome also has a larger histone (H1) associated with both the wrapped DNA and the surface of the core. The series of nucleosomes in chromatin interacts with many nonhistone proteins having a wide variety of enzymatic functions.
FIGURE 3–8 Components of a nucleosome.
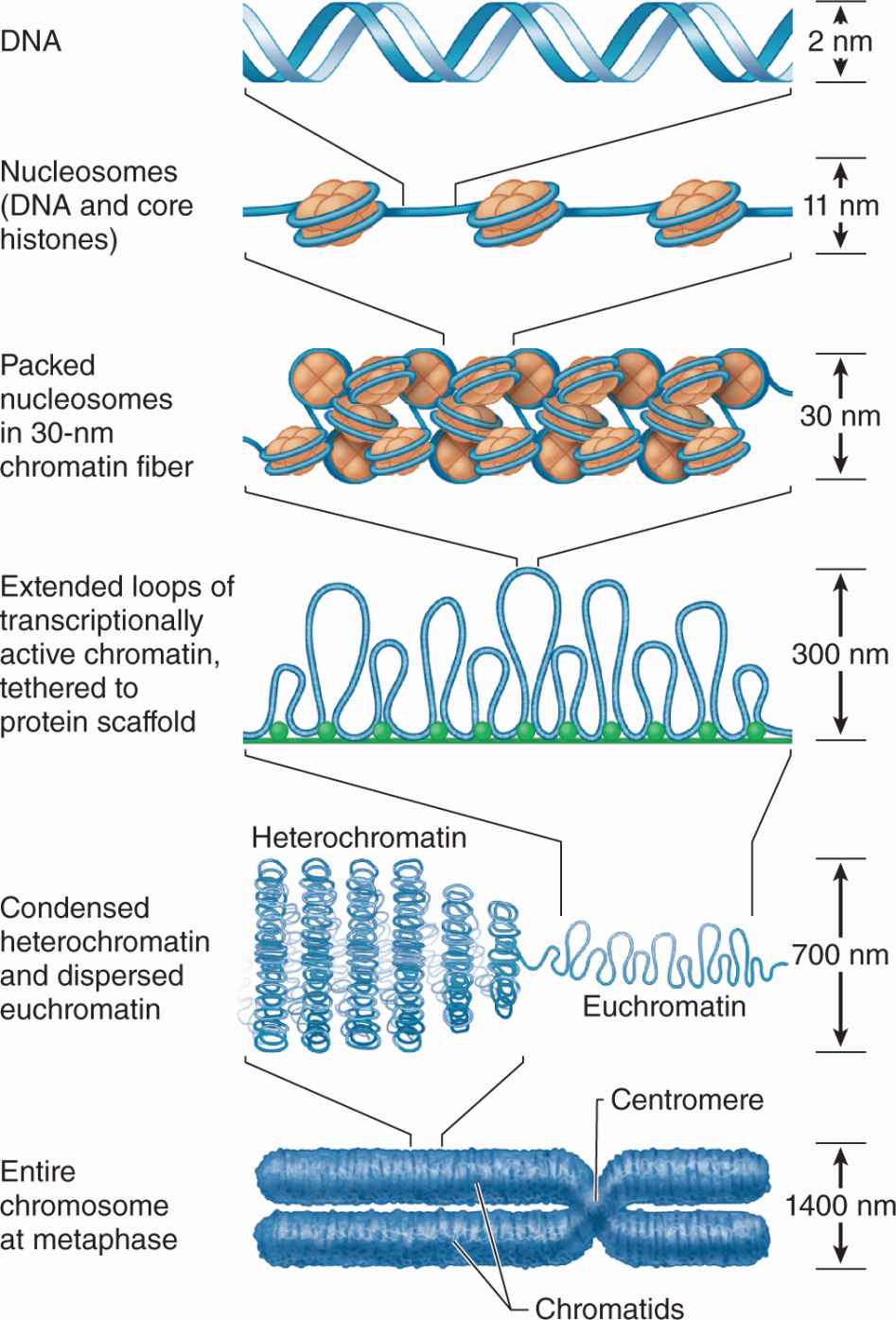
DNA bound to nucleosomes undergoes additional folding and packing to form the 30-nm fiber (Figure 3–9), although the folding mechanism and structures of this and the larger structures are not well understood. Higher orders of chromatin coiling include the formation of transcriptionally active DNA (euchromatin) into loops that are tethered to a central scaffold of proteins that include the condensins. Each long DNA double helix with its associated proteins is a chromatid; after DNA replication two chromatids held together by complexes of cohesin proteins make up each chromosome. Further packaging during the early phase of cell division causes chromosomes to become visible by light microscopy after staining (Figure 3–9).
FIGURE 3–9 From DNA to chromatin.
The X and Y sex chromosomes contain genes determining whether an individual will develop as a female or a male. In addition to the pair of sex chromosomes, cells contain pairs of autosomes. Each of these pairs of chromosomes contains one chromosome originally derived from the mother and one derived from the father. The members of each chromosomal pair are called homologous because, although from different parents, they contain forms (alleles) of the same genes. Cells of most tissues (somatic cells) are considered diploid because they contain pairs of chromosomes. Geneticists refer to diploid cells as 2n, where n is the number of unique chromosomes in a species, 23 in humans. Sperm cells and mature oocytes (germ cells) are haploid, with half the diploid number of chromosomes, each pair having been separated during meiosis (described below).
Microscopic analysis of chromosomes usually begins with cultured cells arrested in mitotic metaphase by colchicine or other compounds that disrupt microtubules. After processing and staining the cells, the condensed chromosomes of one nucleus are photographed by light microscopy and rearranged to produce a karyotype in which stained chromosomal regions (bands) can be analyzed (Figure 3–10).
FIGURE 3–10 Human karyotype.
Nucleolus
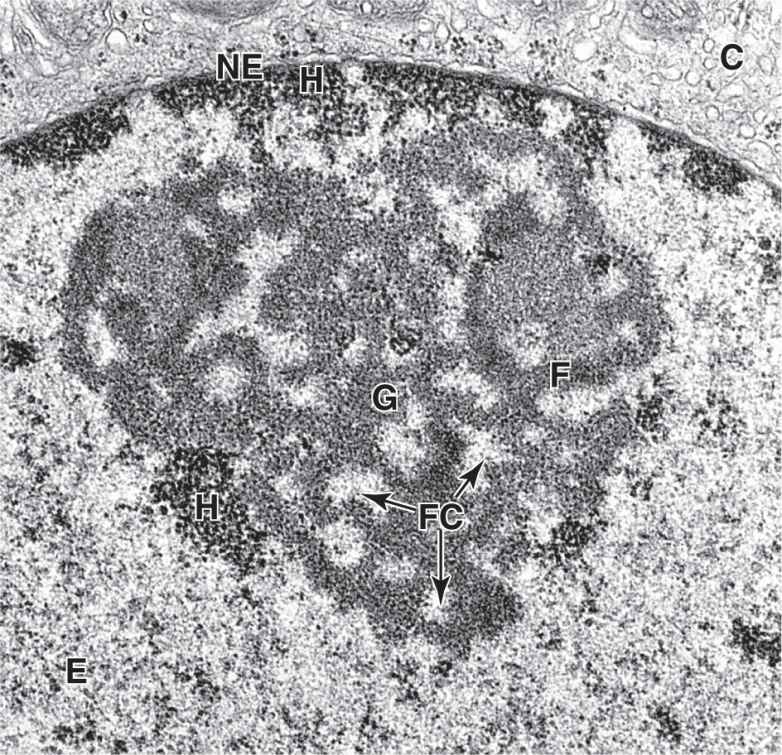
The nucleolus is a generally spherical, highly basophilic subdomain of nuclei in cells, actively making proteins (Figures 3–1 through 3–3). The intense basophilia of nucleoli is due not to heterochromatin but to the presence of densely concentrated ribosomal RNA (rRNA) that is transcribed, processed, and complexed into ribosomal subunits in nucleoli. Chromosomal regions with the genes for rRNA organize one or more nucleoli in cells requiring intense ribosome production for synthesis of proteins during growth or secretion. Ultrastructural analysis of an active nucleolus reveals fibrillar and granular subregions with different staining characteristics that reflect stages of rRNA maturation (Figure 3–11). Molecules of rRNA are processed in the nucleolus and very quickly associate with the ribosomal proteins imported from the cytoplasm via nuclear pore complexes. The newly organized small and large ribosomal subunits are then exported back to the cytoplasm through those same nuclear pores.
FIGURE 3–11 Regions within a nucleolus.

 MEDICAL APPLICATION
MEDICAL APPLICATION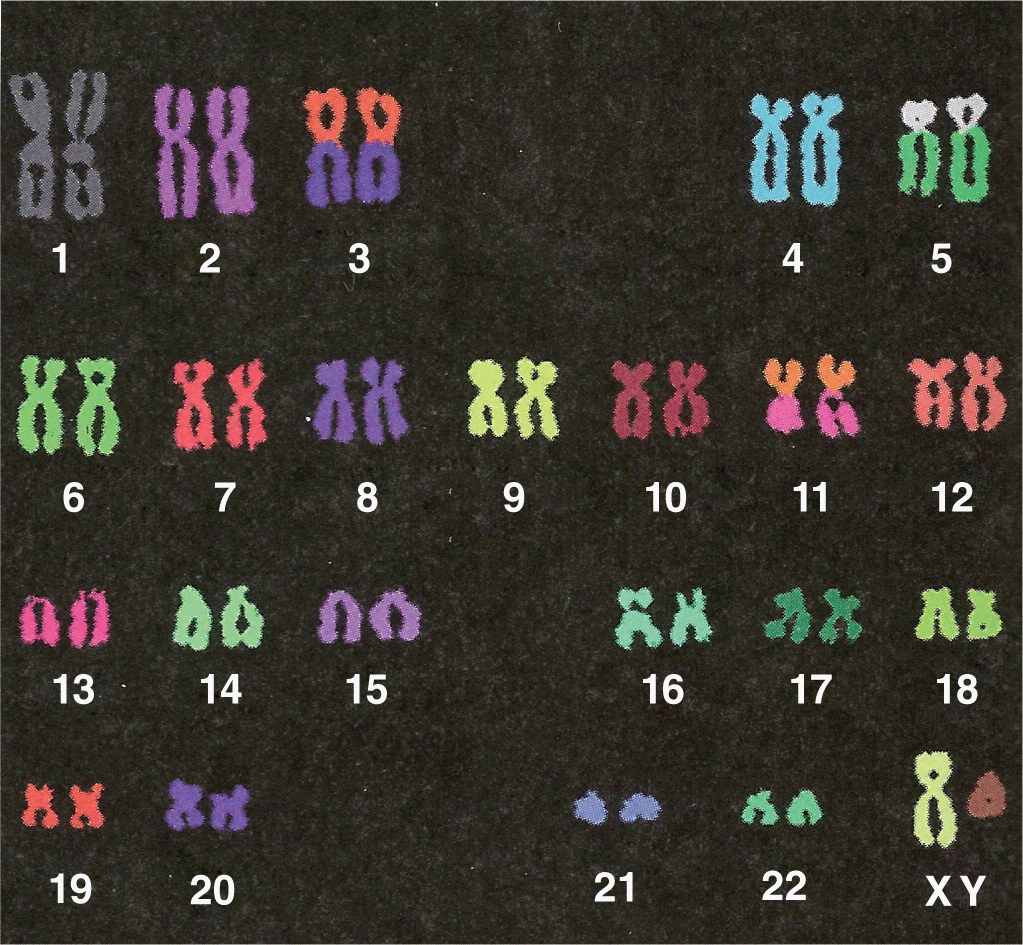
 MEDICAL APPLICATION
MEDICAL APPLICATION