In addition to the nuclear genome, a small but important part of the human genome resides in mitochondria in the cytoplasm (see Fig. 2-1). The mitochondrial chromosome, to be described later in this chapter, has a number of unusual features that distinguish it from the rest of the human genome.
Genes in the Human Genome
What is a gene? And how many genes do we have? These questions are more difficult to answer than it might seem.
The word gene, first introduced in 1908, has been used in many different contexts since the essential features of heritable “unit characters” were first outlined by Mendel over 150 years ago. To physicians (and indeed to Mendel and other early geneticists), a gene can be defined by its observable impact on an organism and on its statistically determined transmission from generation to generation. To medical geneticists, a gene is recognized clinically in the context of an observable variant that leads to a characteristic clinical disorder, and today we recognize approximately 5000 such conditions (see Chapter 7).
The Human Genome Project provided a more systematic basis for delineating human genes, relying on DNA sequence analysis rather than clinical acumen and family studies alone; indeed, this was one of the most compelling rationales for initiating the project in the late 1980s. However, even with the finished sequence product in 2003, it was apparent that our ability to recognize features of the sequence that point to the existence or identity of a gene was sorely lacking. Interpreting the human genome sequence and relating its variation to human biology in both health and disease is thus an ongoing challenge for biomedical research.
Although the ultimate catalogue of human genes remains an elusive target, we recognize two general types of gene, those whose product is a protein and those whose product is a functional RNA.
• The number of protein-coding genes—recognized by features in the genome that will be discussed in Chapter 3—is estimated to be somewhere between 20,000 and 25,000. In this book, we typically use approximately 20,000 as the number, and the reader should recognize that this is both imprecise and perhaps an underestimate.
• In addition, however, it has been clear for several decades that the ultimate product of some genes is not a protein at all but rather an RNA transcribed from the DNA sequence. There are many different types of such RNA genes (typically called noncoding genes to distinguish them from protein-coding genes), and it is currently estimated that there are at least another 20,000 to 25,000 noncoding RNA genes around the human genome.
Thus overall—and depending on what one means by the term—the total number of genes in the human genome is of the order of approximately 20,000 to 50,000. However, the reader will appreciate that this remains a moving target, subject to evolving definitions, increases in technological capabilities and analytical precision, advances in informatics and digital medicine, and more complete genome annotation.
DNA Structure: A Brief Review
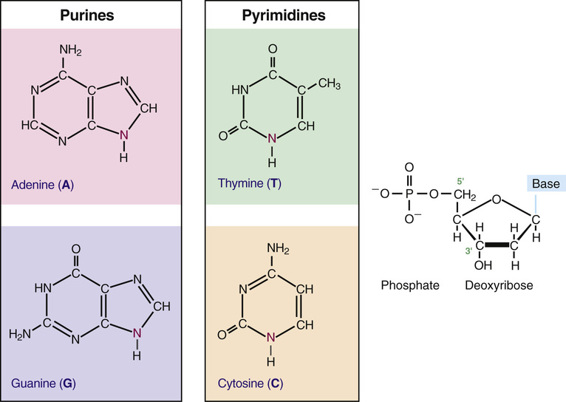
Before the organization of the human genome and its chromosomes is considered in detail, it is necessary to review the nature of the DNA that makes up the genome. DNA is a polymeric nucleic acid macromolecule composed of three types of units: a five-carbon sugar, deoxyribose; a nitrogen-containing base; and a phosphate group (Fig. 2-2). The bases are of two types, purines and pyrimidines. In DNA, there are two purine bases, adenine (A) and guanine (G), and two pyrimidine bases, thymine (T) and cytosine (C). Nucleotides, each composed of a base, a phosphate, and a sugar moiety, polymerize into long polynucleotide chains held together by 5′-3′ phosphodiester bonds formed between adjacent deoxyribose units (Fig. 2-3A). In the human genome, these polynucleotide chains exist in the form of a double helix (Fig. 2-3B) that can be hundreds of millions of nucleotides long in the case of the largest human chromosomes.
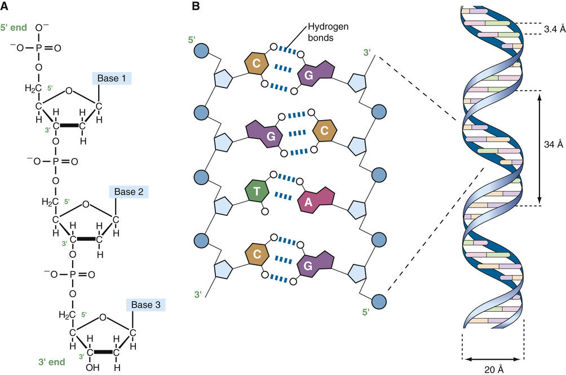
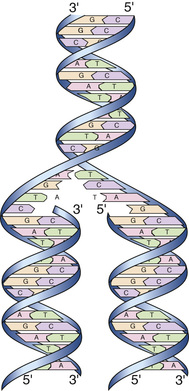
The anatomical structure of DNA carries the chemical information that allows the exact transmission of genetic information from one cell to its daughter cells and from one generation to the next. At the same time, the primary structure of DNA specifies the amino acid sequences of the polypeptide chains of proteins, as described in the next chapter. DNA has elegant features that give it these properties. The native state of DNA, as elucidated by James Watson and Francis Crick in 1953, is a double helix (see Fig. 2-3B). The helical structure resembles a right-handed spiral staircase in which its two polynucleotide chains run in opposite directions, held together by hydrogen bonds between pairs of bases: T of one chain paired with A of the other, and G with C. The specific nature of the genetic information encoded in the human genome lies in the sequence of C’s, A’s, G’s, and T’s on the two strands of the double helix along each of the chromosomes, both in the nucleus and in mitochondria (see Fig. 2-1). Because of the complementary nature of the two strands of DNA, knowledge of the sequence of nucleotide bases on one strand automatically allows one to determine the sequence of bases on the other strand. The double-stranded structure of DNA molecules allows them to replicate precisely by separation of the two strands, followed by synthesis of two new complementary strands, in accordance with the sequence of the original template strands (Fig. 2-4). Similarly, when necessary, the base complementarity allows efficient and correct repair of damaged DNA molecules.
Structure of Human Chromosomes
The composition of genes in the human genome, as well as the determinants of their expression, is specified in the DNA of the 46 human chromosomes in the nucleus plus the mitochondrial chromosome. Each human chromosome consists of a single, continuous DNA double helix; that is, each chromosome is one long, double-stranded DNA molecule, and the nuclear genome consists, therefore, of 46 linear DNA molecules, totaling more than 6 billion nucleotide pairs (see Fig. 2-1).
Chromosomes are not naked DNA double helices, however. Within each cell, the genome is packaged as chromatin, in which genomic DNA is complexed with several classes of specialized proteins. Except during cell division, chromatin is distributed throughout the nucleus and is relatively homogeneous in appearance under the microscope. When a cell divides, however, its genome condenses to appear as microscopically visible chromosomes. Chromosomes are thus visible as discrete structures only in dividing cells, although they retain their integrity between cell divisions.
The DNA molecule of a chromosome exists in chromatin as a complex with a family of basic chromosomal proteins called histones. This fundamental unit interacts with a heterogeneous group of nonhistone proteins, which are involved in establishing a proper spatial and functional environment to ensure normal chromosome behavior and appropriate gene expression.
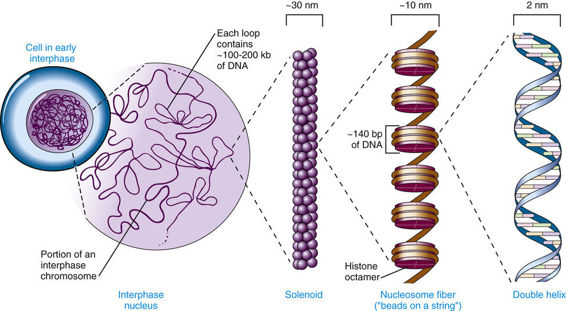
Five major types of histones play a critical role in the proper packaging of chromatin. Two copies each of the four core histones H2A, H2B, H3, and H4 constitute an octamer, around which a segment of DNA double helix winds, like thread around a spool (Fig. 2-5). Approximately 140 base pairs (bp) of DNA are associated with each histone core, making just under two turns around the octamer. After a short (20- to 60-bp) “spacer” segment of DNA, the next core DNA complex forms, and so on, giving chromatin the appearance of beads on a string. Each complex of DNA with core histones is called a nucleosome (see Fig. 2-5), which is the basic structural unit of chromatin, and each of the 46 human chromosomes contains several hundred thousand to well over a million nucleosomes. A fifth histone, H1, appears to bind to DNA at the edge of each nucleosome, in the internucleosomal spacer region. The amount of DNA associated with a core nucleosome, together with the spacer region, is approximately 200 bp.