As introduced briefly in Chapter 2, the product of protein-coding genes is a protein whose structure ultimately determines its particular functions in the cell. But if there were a simple one-to-one correspondence between genes and proteins, we could have at most approximately 20,000 different proteins. This number seems insufficient to account for the vast array of functions that occur in human cells over the life span. The answer to this dilemma is found in two features of gene structure and function. First, many genes are capable of generating multiple different products, not just one (see Fig. 3-1). This process, discussed later in this chapter, is accomplished through the use of alternative coding segments in genes and through the subsequent biochemical modification of the encoded protein; these two features of complex genomes result in a substantial amplification of information content. Indeed, it has been estimated that in this way, these 20,000 human genes can encode many hundreds of thousands of different proteins, collectively referred to as the proteome. Second, individual proteins do not function by themselves. They form elaborate networks, involving many different proteins and regulatory RNAs that respond in a coordinated and integrated fashion to many different genetic, developmental, or environmental signals. The combinatorial nature of protein networks results in an even greater diversity of possible cellular functions.
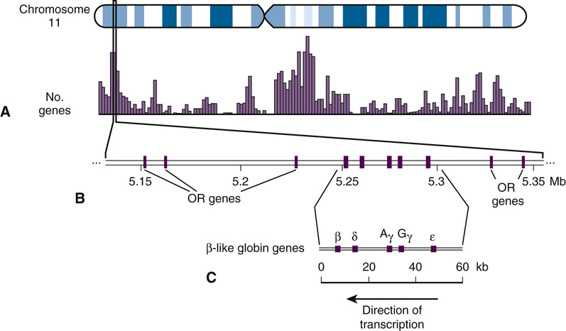
Genes are located throughout the genome but tend to cluster in particular regions on particular chromosomes and to be relatively sparse in other regions or on other chromosomes. For example, chromosome 11, an approximately 135 million-bp (megabase pairs [Mb]) chromosome, is relatively gene-rich with approximately 1300 protein-coding genes (see Fig. 2-7). These genes are not distributed randomly along the chromosome, and their localization is particularly enriched in two chromosomal regions with gene density as high as one gene every 10 kb (Fig. 3-2). Some of the genes belong to families of related genes, as we will describe more fully later in this chapter. Other regions are gene-poor, and there are several so-called gene deserts of a million base pairs or more without any known protein-coding genes. Two caveats here: first, the process of gene identification and genome annotation remains very much an ongoing challenge; despite the apparent robustness of recent estimates, it is virtually certain that there are some genes, including clinically relevant genes, that are currently undetected or that display characteristics that we do not currently recognize as being associated with genes. And second, as mentioned in Chapter 2, many genes are not protein-coding; their products are functional RNA molecules (noncoding RNAs or ncRNAs; see Fig. 3-1) that play a variety of roles in the cell, many of which are only just being uncovered.

Stay updated, free articles. Join our Telegram channel

Full access? Get Clinical Tree


