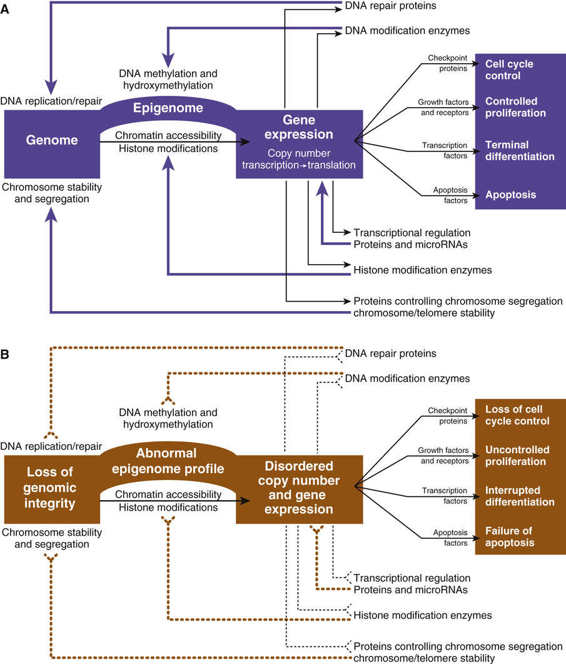
Figure 15-2 is a diagram outlining how mutations in specific regulators of growth and in global guardians of DNA and genome integrity perturb normal homeostasis (see Fig. 15-2A), leading to a vicious cycle causing loss of cell cycle control, uncontrolled proliferation, interrupted differentiation, and defects in apoptosis (see Fig. 15-2B).
Activated Oncogenes and Tumor Suppressor Genes
Both classes of driver genes—those with specific effects on cellular proliferation or survival and those with global effects on genome or DNA integrity (see Table 15-1)—can be further subdivided into one of two functional categories depending on how, if mutated, they drive oncogenesis.
The first category includes proto-oncogenes. These are normal genes that, when mutated in very particular ways, become driver genes through alterations that lead to excessive levels of activity. Once mutated in this way, driver genes of this type are referred to as activated oncogenes. Only a single mutation at one allele can be sufficient for activation, and the mutations that activate a proto-oncogene can range from highly specific point mutations causing dysregulation or hyperactivity of a protein, to chromosome translocations that drive overexpression of a gene, to gene amplification events that create an overabundance of the encoded mRNA and protein product (Fig. 15-3).
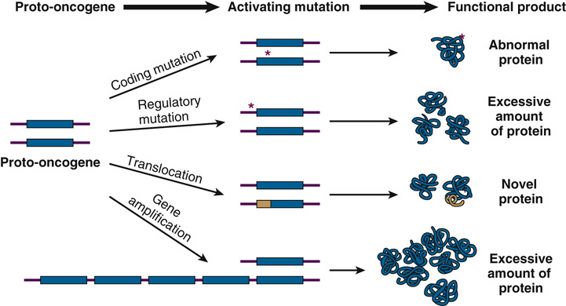
The second, and more common, category of driver genes includes tumor suppressor genes (TSGs), mutations in which cause a loss of expression of proteins necessary to control the development of cancers. To drive oncogenesis, loss of function of a TSG typically requires mutations at both alleles. There are many ways that a cell can lose the function of TSG alleles. Loss-of-function mechanisms can range from missense, nonsense, or frame-shift mutations to gene deletions or loss of a part or even an entire chromosome. Loss of function of TSGs can also result from epigenomic transcriptional silencing due to altered chromatin conformation or promoter methylation (see Chapter 3), or from translational silencing by miRNAs or disturbances in other components of the translational machinery (see Box).
Genetic Basis of Cancer
Regardless of whether a cancer occurs sporadically in an individual, as a result of somatic mutation, or repeatedly in many individuals in a family as a hereditary trait, cancer is a genetic disease.
• Genes in which mutations cause cancer are referred to as driver genes, and the cancer-causing mutations in these genes are driver mutations.
• Driver genes fall into two distinct categories: activated oncogenes and tumor suppressor genes (TSGs).
• An activated oncogene is a mutant allele of a proto-oncogene, a class of normal cellular protein-coding genes that promotes growth and survival of cells. Oncogenes facilitate malignant transformation by stimulating proliferation or inhibiting apoptosis. Oncogenes encode proteins such as the following:
• Proteins in signaling pathways for cell proliferation
• Transcription factors that control the expression of growth-promoting genes
• Inhibitors of programmed cell death machinery
• A TSG is a gene in which loss of function through mutation or epigenomic silencing directly removes normal regulatory controls on cell growth or leads indirectly to such losses through an increased mutation rate or aberrant gene expression. TSGs encode proteins involved in many aspects of cellular function, including maintenance of correct chromosome number and structure, DNA repair proteins, proteins involved in regulating the cell cycle, cellular proliferation, or contact inhibition, just to name a few examples.
• Tumor initiation can be caused by different types of genetic alterations. These include mutations such as the following:
• Activating or gain-of-function mutations, including gene amplification, point mutations, and promoter mutations, that turn one allele of a proto-oncogene into an oncogene
• Ectopic and heterochronic mutations (see Chapter 11) of proto-oncogenes
• Chromosome translocations that cause misexpression of genes or create chimeric genes encoding proteins with novel functional properties
• Loss of function of both alleles, or a dominant negative mutation of one allele, of TSGs
• Tumor progression occurs as a result of accumulating additional genetic damage, through mutations or epigenetic silencing, of driver genes that encode the machinery that repairs damaged DNA and maintains cytogenetic normality. A further consequence of genetic damage is altered expression of genes that promote vascularization and the spread of the tumor through local invasion and distant metastasis.
Stay updated, free articles. Join our Telegram channel

Full access? Get Clinical Tree


