Instabilities
1.
Physical
• Conformational stability
Thermal denaturation
Cold denaturation
Pressure-induced denaturation
Chemical denaturation
• Colloidal stability
• Surface/interfacial-induced denaturation
2.
Chemical
• Deamidation
• Asp isomerization
• Fragmentation
• Glycation
• Oxidation
• Disulfide scrambling
6.5 Use and Description of Biophysical Tools in Molecule Selection and Development
A typical early screen for a mAb might initially focus on a property, such as specificity or strength (affinity) of binding. Subsequent secondary screens and assays typically include specificity against related molecules to the target and binding to species homologues (to support preclinical safety studies). In this way a “screening cascade” of different assays is performed, reducing the number of potential antibody variants for subsequent development. As the number of individual variants is reduced, the amount and purity of sample is typically concomitantly increased. Ideally, screens for developability characteristics, such as solubility, expression, chemical stability, and reduced propensity for aggregation would be placed as early in the screening cascade as possible in order to maximize the potential sequence diversity available in a large population of variants. For example, laboratories have reported the use of modified phage-display selections to preferentially enrich for those variants that have increased thermal stability (Jespers et al. 2004; Famm et al. 2008). Taking this approach even further, antibody-fragment libraries have been designed to maximize stability, expression levels, and aggregation resistance (Prassler et al. 2011). High-throughput (HT) assays are ideally suited for performance early in the screening cascade, allowing biophysical properties to be preferentially selected for, alongside other properties, such as affinity and specificity.
There are a wide array of biophysical techniques and strategies available to determine protein instabilities (Table 6.2). Each of these techniques has inherent advantages and disadvantages with respect to its ability to detect a meaningful change in the structure of a protein and to translate this change into information that can be used to rank order, or exclude or include, a candidate in a preclinical development program. The sensitivity of the technique combined with the predictive nature of the analysis can help determine the suitability of a method for screening preclinical candidates. Often the techniques are employed in specific studies designed to elicit a behavior that can then be directly measured. The criteria used for identifying the ideal analytical methods and testing strategies are typically those associated with the phase of project development. For the screening of large-molecule libraries, methods that offer advantages in speed and throughput while using little material are preferred. At this point, the types of methodologies that should be employed become limited to those that are robust and sensitive towards specific behaviors that enable selection of a lead candidate for CMC development.
Table 6.2
List of most commonly used biophysical tools
Instabilities | Method | High-throughput amenable |
|---|---|---|
Conformational stability | Differential scanning calorimetry | Yes |
Fluorescence spectroscopy • Extrinsic • Intrinsic | Yes | |
Absorption spectroscopy | Yes | |
Circular dichroism | Yes | |
Fourier transform infrared spectroscopy | No | |
Raman spectroscopy | No | |
NMR | No | |
X-ray | No | |
Dynamic light scattering | Yes | |
Colloidal stability | Light scattering • SLS • DLS | Yes |
Membrane osmometer | No | |
Self-interaction chromatography | Yes | |
Analytical ultracentrifugation • Sedimentation velocity • Sedimentation equilibrium | No | |
Interfacial stability | Tensiometry | No |
Extensional rheology | No | |
Chemical stability | RPHPLC | No |
Bioanalyzer | Yes | |
LC/MS | No | |
Solution properties | ||
Viscosity | Rheometry | No |
DLS | Yes | |
Phase separation | Nephleometry | Yes |
Right angle light scattering | No | |
Absorption spectroscopy (transmission) | No | |
For the biophysical assessment of a large number of candidate molecules, HT approaches are preferable. In general, available techniques should support miniaturization, simple data visualization, and interpretation to help obtain more information in well-focused studies. The techniques that are most attractive are those that can robustly predict or measure specific molecular attributes that indicate if a candidate meets a defined developability criteria that are specific to its target indication or is within an acceptable developability design space either though mechanistic rules or phenomological correlation. Such a technique should allow for exploration of a larger number of instabilities or parameters more quickly with a lower cost. In general the methods should be flexible, miniature (consume minimal amounts of material), and employable in a HT mode such that a battery of different screens can be performed on the same sample simultaneously or in a highly parallel array.
6.6 Molecular Attributes and Instabilities that Should Be Screened Early
6.6.1 Aggregation
The propensity for a molecule to aggregate is one of the single most influential and challenging factors for formulation development. Candidates that are inherently resistant towards aggregation are considered superior. Nonnative aggregation is a multistep process and can be explained by the Lumry–Eyring framework. Each state has a different molecular configuration and energy, and each reaction step proceeds through an energy barrier (ΔG*, difference in energy or activation free energy). The step that has highest ΔG* is the rate-limiting step (Chi et al. 2003a). The first step in this process, which is a reversible process, gives rise to an aggregation-competent partially unfolded intermediate (I) state. In the second step, the partially unfolded intermediate self-associates to from irreversible higher-order aggregates (A). The final undesired A state can be controlled either by increasing conformational stability of the native (N) state (ΔG) which results in a decreased population of aggregation-competent partially unfolded intermediates (I state) and/or by increasing the colloidal stability of the partially unfolded I state such that self-association of this state is minimized (Chi et al. 2003b). During development, controlling aggregation is achieved by screening for stabilizing excipients and solution conditions to develop appropriate formulation controls. Alternatively, such stabilization can be achieved by either selecting or designing conformationally and/or colloidally stable candidates. Mutations that increase conformational stability increase the free energies of the unfolded and transition states relative to the native state. Increasing the free energy of transition state relative to native results in a larger ΔG*, which shifts the equilibrium towards native state. The net result is a decreased population of aggregation-competent partially unfolded state.
Both conformational and colloidal stability have an indirect but significant effect on surface- and interface-mediated properties. For example, adsorption of protein to various surfaces and interfaces during process, fill/finish, and storage can not only lead to significant reduction in yield but quite often can also induce structural perturbation in a protein resulting in reduced binding and activity as well as aggregation and particle formation (Sethuraman and Belfort 2005). Similarly shear stress on a protein during process and fill/finish operation along with mixing of the hydrophobic air–liquid interface with the bulk solution is known to induce structural perturbations and subsequent aggregations (Bekard et al. 2011; Thomas and Geer 2011; Biddlecombe et al. 2007; Bee et al. 2009). Typically, both surface- and interface-mediated aggregation and particle formation is controlled during development either by avoiding the surfaces that can have deleterious effect on a particular protein or by designing and developing appropriate formulation controls such that the adsorption process is avoided or approximately controlled. Alternatively, such surface and interface effects can be reduced by increasing the conformational stability of a protein (Karlsson et al. 2005; Norde 1986; Wendorf et al. 2004). In a study with human carbonic anhydrase mutants, Karlsson et al. (2005) showed that adsorption and desorption characteristics can be controlled by a protein-engineering approach. An early investment to either design or select molecules with better conformational stability can potentially reduce some of the process and fill/finish related surface- and interface-mediated effects. Such an approach (stability by design) will not only result in a wider process and formulation development window but can also result in development of a platform process and fill/finish conditions.
Thermal stability using differential scanning calorimetry (DSC) is a widely acceptable measure and predictor of a protein’s conformational stability and can be used to form a basis for screening candidate molecules under varying buffer conditions (Remmele et al. 1998). New techniques, strategies, and instrumentation are now available that provide thermal stability in a fraction of the time typically associated with DSC and use very low amounts of material (He et al. 2009; Goldberg et al. 2010). These techniques include spectroscopy-based techniques coupled with thermal ramping (reviewed in Chap. 2). These techniques can provide throughput on the order of 96–384 samples per hour and typically only require 10s of micrograms. Aggregation and particle formation can also be directly monitored by techniques such as dynamic light scattering (DLS), HIAC light obscuration, micro-flow imaging (MFI), and electrical charge-based methods (e.g., Coulter counter). Of these methods, DLS offers the greatest flexibility for early-stage use based on throughput (plate-based DLS) and material required. However, recent advances have resulted in improvements in particle detection including scanning ion occlusion spectroscopy (qNano), chip-based mass measurements (Archimedes), and nanoparticle tracking analysis (NanoSight). Each of these methods offers the possibility of increased sensitivity and throughput with a decrease in sample consumption, resulting in more screening possibilities.
6.6.2 Chemical Stability
As shown in Table 6.1, proteins have multiple liabilities that are prone to chemical degradation during manufacture and storage. The rate of degradation depends on both the solution condition and the structure of the molecule (Sinha et al. 2009; Vlasak and Ionescu 2011). Typically some of the most common liabilities like deamidation, nonenzymatic hydrolysis, and oxidation are highlighted very early and easily by in silico primary sequence liability analysis. Theoretically, molecules with the fewest chemical liabilities will allow one to develop a wider process and formulation design space. For molecules with identified liabilities, there will be an additional burden of determining the effect of the liabilities on product quality and designing appropriate process and formulation controls. Unlike physical stability studies, the slow rates of reaction associated with accessing chemical stability results in time-consuming forced degradation studies. For example, evaluation of the effect of deamidation on function and stability requires forced degradation studies at elevated temperature and pH, followed by liquid chromatography–mass spectrometry (LCMS) and subsequent selected ion monitoring (SIM). Similarly fragmentation propensity can be determined either using size-based separation techniques or side-chain chemistry-based analytical methods like reversed-phase high-performance liquid chromatography (RPHPLC). Even though HT analytics are available to follow fragmentation, the slow rate of reaction of forced degradation studies that can accurately predict nonenzymatic fragmentation under storage conditions typically requires time-consuming incubation studies in the range of 4–6 weeks.
6.6.3 Solution Properties at High Protein Concentrations
Other important developability criteria that are often neglected during early screening are the solution properties of a molecule at high concentration. Solution properties such as solubility, phase separation, and viscosity can often become the rate-limiting step during processing, filtration, drug delivery, etc., as solution properties of a molecule are governed by various attractive and repulsive nonspecific intermolecular interactions [reviewed in Saluja and Kalonia (2008) and Tadros (2011)]. Under concentrated solution conditions, the intermolecular distance becomes less than the molecule’s diameter. This reduction in intermolecular distance will not only increase the collision frequency but will also increase the duration of such an encounter (Scherer et al. 2010; Minton 2005). Thus at high concentrations, the nature of, and relative contributions from, various nonspecific interactions is dictated by both solution conditions and the surface properties of a molecule, including net charge, charge distribution, and surface hydrophobicity. From a development perspective, the consequences of such unfavorable interactions are a change in the protein’s solution behavior potentially resulting in issues such as poor solubility, phase separation, and/or increased viscosity (Saluja and Kalonia 2008; Chari et al. 2009). Typically, such unfavorable interactions can be controlled during development by modulating solution conditions like ionic strength and/or pH to develop appropriate formulation controls (Salinas et al. 2010; Yadav et al. 2012; Liu et al. 2005), but in some cases, depending on strength of intermolecular interactions, it may not be possible to achieve this control. Quite often the solution conditions might have an unintended opposite effect on various solution properties and overall stability (Salinas et al. 2010).
The rheological properties of a molecule under a given buffer system have typically been ascertained using a cone- and plate-type instrument to determine viscosity. The dimensions of the cone and plate instruments have been dramatically reduced over the years, resulting in a correlating decrease in the volume of material required. However, the time associated with these measurements has not fallen to the same degree. New techniques such as viscosity–DLS (see He et al., Chap. 2, this volume) and chip-based measurements provide an attractive alternative to the traditional cone and plate approach. Alternatively, solution properties can also be predicted by accurate measurements of net protein–protein interactions through colloidal stability (Yadav et al. 2012; George and Wilson 1994; Mehta et al. 2012). The osmotic second virial coefficient (B 22), which is a thermodynamic parameter, is often used to directly quantify overall protein–protein interactions (Chi et al. 2003a, b; Valente et al. 2005; Payne et al. 2006). A positive B 22 value indicates an overall dominance of repulsive interactions, where as a negative B 22 value indicates an overall attractive intermolecular interactions. The B 22 of a colloidally active system can be accurately measured by various techniques like DLS (Yadav et al. 2011), static light scattering (SLS), analytical ultracentrifugation (AUC), osmometry, and self-interaction chromatography (SIC) (Le Brun et al. 2010).
One of the main reasons for limited screening of these undesirable solution properties during early development is the limited availability of HT techniques. Most of the available techniques mentioned above are low throughput in nature and also require large amounts of material. But recently significant progress has been made in this area, and several HT methods are available to either directly measure some of the solution properties (reviewed inChap. 2) or accurately predict solution properties (Johnson et al. 2009; Bajaj et al. 2007; Sule et al. 2011; Tessier et al. 2008). Availability of such HT techniques allows one to screen and select candidate molecules with desired surface properties or interaction potential, which is a molecule that has a B 22 value close to 0 under the desired solution conditions. Applying an approach of selecting or engineering a molecule with desired surface properties, or interaction potential, will enable the formulation scientist to focus mostly on aggregation or other chemical degradations, thus widening the formulation development design space.
One consequence of such an approach is that there is a potential for conflict between the different parameters that are being simultaneously screened. For example, if antibody affinity is found to correlate with an unfavorable biophysical characteristic, such as aggregation propensity, careful consideration would need to be given as to which characteristic would be most economic to engineer/mitigate against the aggregation propensity. Examples of candidate therapeutic antibodies that have been extensively reengineered to improve biophysical properties and as such improve developability have begun to be described (Pepinsky et al. 2010).
6.7 Case Studies
6.7.1 Case Study 1
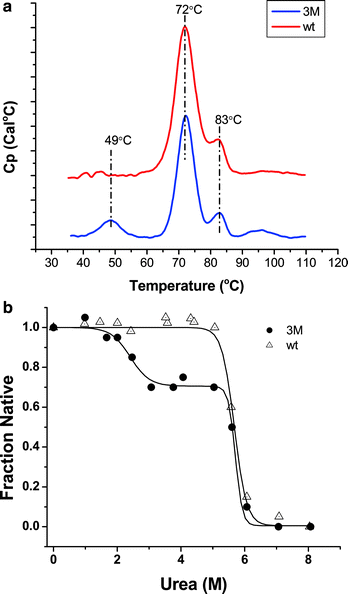
As stated in Sect. 6.2, for molecules with a precedented mechanism of action, the focus is on enhanced safety, higher affinity to target, and increased half-life. The efficacy and potency of a molecule can be fine-tuned by efficient mediation of effector functions (Anderson et al. 1997; Weng and Levy 2003). This case study demonstrates how engineering of Fc variants to modulate these activities can have an unintentional effect on a molecule’s developability and how this unintentional effect can be screened for and detected early by applying appropriate biophysical tools. In this case a triple mutant S239D/A330L/I332E (resulting in mAbA-3M) was introduced into the CH2 domain of a human mAb (mAbA). Compared to its wild type, these mutations resulted in an approximately 10–100 fold increase in mAbA-3M binding to its receptor, but at a price of reduced stability (Fig. 6.1) (Oganesyan et al. 2008; Anandakumar et al. 2008). Subsequent developability-accelerated stability studies indicated a significant increase in the aggregation rate at 40 °C (Fig. 6.2). Based on various biophysical studies like DSC and denaturant unfolding, it was speculated that the reduced conformational stability was due to minor structural perturbations in the region around the CH2 domain. By solving the structure of the mutant, Oganesyan et al. determined that the mutations resulted in opening of the cleft in the CH2 domain of the Fc fragment (Oganesyan et al. 2008). They concluded that the enhanced interactions with human Fcγ(gamma)RIIIA could be due to enhanced Fc openness as well as the introduction of hydrophobic and electrostatic interactions at the corresponding interface (Oganesyan et al. 2008). Even though developability studies flagged the mAbA-3M as having a problematic aggregation potential, it was selected as the lead candidate because of its increased potency. Investigators were able to move mAbA-3M forward because appropriate formulation controls and resources were allotted based on the information obtained during developability studies.